The AlphaFold revolution in Structural Biology
From distance matrices to diffusion
In just six years, DeepMind transformed protein structure prediction from a frustrating computational puzzle into a virtually solved problem—and then extended that solution to the entire molecular machinery of life. AlphaFold’s evolution from a distance-predicting neural network (AF1) to an end-to-end structure generator (AF2) to a unified diffusion model for all biomolecules (AF3) represents one of the most consequential achievements in computational biology. This technical deep-dive traces that architectural journey, explaining how each innovation built upon the last to achieve what the Nobel Committee called “one of the most important scientific breakthroughs made by mankind in the 21st century.”
The breakthrough was not incremental. AlphaFold2 achieved median backbone accuracy of 0.96 Å—comparable to experimental structures—while AlphaFold3 expanded prediction to protein-nucleic acid complexes, small molecule binding, and covalent modifications within a single unified framework. Understanding how these systems work illuminates not just protein structure prediction but the broader potential of deep learning in molecular biology.
The foundation: AlphaFold1’s two-stage approach
AlphaFold1 entered the CASP13 competition in December 2018 with a conceptually simple but powerful insight: rather than predicting binary contacts between residues, predict full probability distributions over inter-residue distances, then use these distributions as a protein-specific statistical potential for structure optimization.
1. Distance distribution prediction
The core of AlphaFold1 was a deep dilated residual convolutional network with 220 residual blocks and 128 channels. Each block contained batch normalization layers, 1×1 projection layers, and critically, dilated convolutions—inspired by WaveNet—that provided non-contiguous receptive fields spanning the entire protein sequence without excessive computational cost.
The network output probability distributions over 64 distance bins ranging from 2 Å to 22 Å for each residue pair $(i, j)$, predicting distances between $C_{\beta}$ atoms ($C_{\alpha}$ for glycine). This granular distance information proved far more useful than the binary contact predictions used by previous methods like PSICOV and EVfold.
Input features were derived primarily from multiple sequence alignments (MSAs):
- Position-specific scoring matrices (PSSMs)
- Hidden Markov Model profiles from HHblits
- Co-evolutionary coupling matrices
- One-hot sequence encodings
The co-evolutionary principle underlying this approach—that residues in spatial contact tend to show correlated mutations over evolutionary timescales—was not new. What was new was learning these patterns end-to-end through deep neural networks rather than explicit Potts models or direct coupling analysis (DCA).
2. From distributions to structures
The second stage converted predicted distance distributions into 3D coordinates through gradient descent optimization. The probability distribution $P(d_{ij})$ was transformed into a potential of mean force:
\[\mathrm{V_{distance}} = -\log[\mathrm{P(d_{ij} | sequence, MSA) / P(d_{ij} | length)}]\]The denominator represents a learned reference state—a key innovation over traditional knowledge-based potentials. This reference potential was trained on the same data but without sequence or MSA input, effectively normalizing for length-dependent biases.
The distance potential was combined with physics-based terms from Rosetta (van der Waals repulsion, backbone torsion preferences, steric clash terms), then minimized using L-BFGS gradient descent with multiple noisy restarts. No fragment assembly, no Markov chain Monte Carlo sampling—just direct optimization of a differentiable energy function.
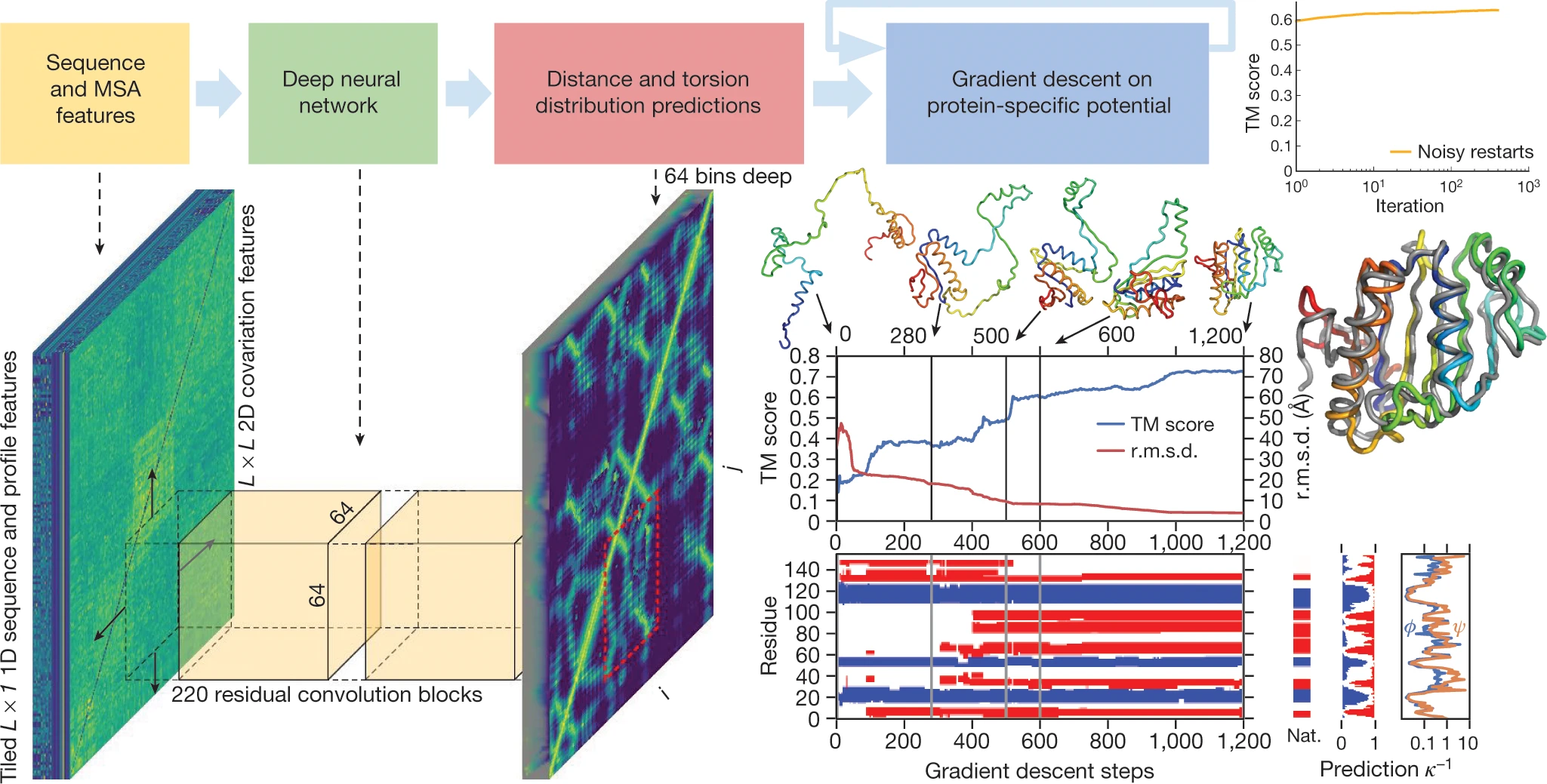
3. CASP13 performance
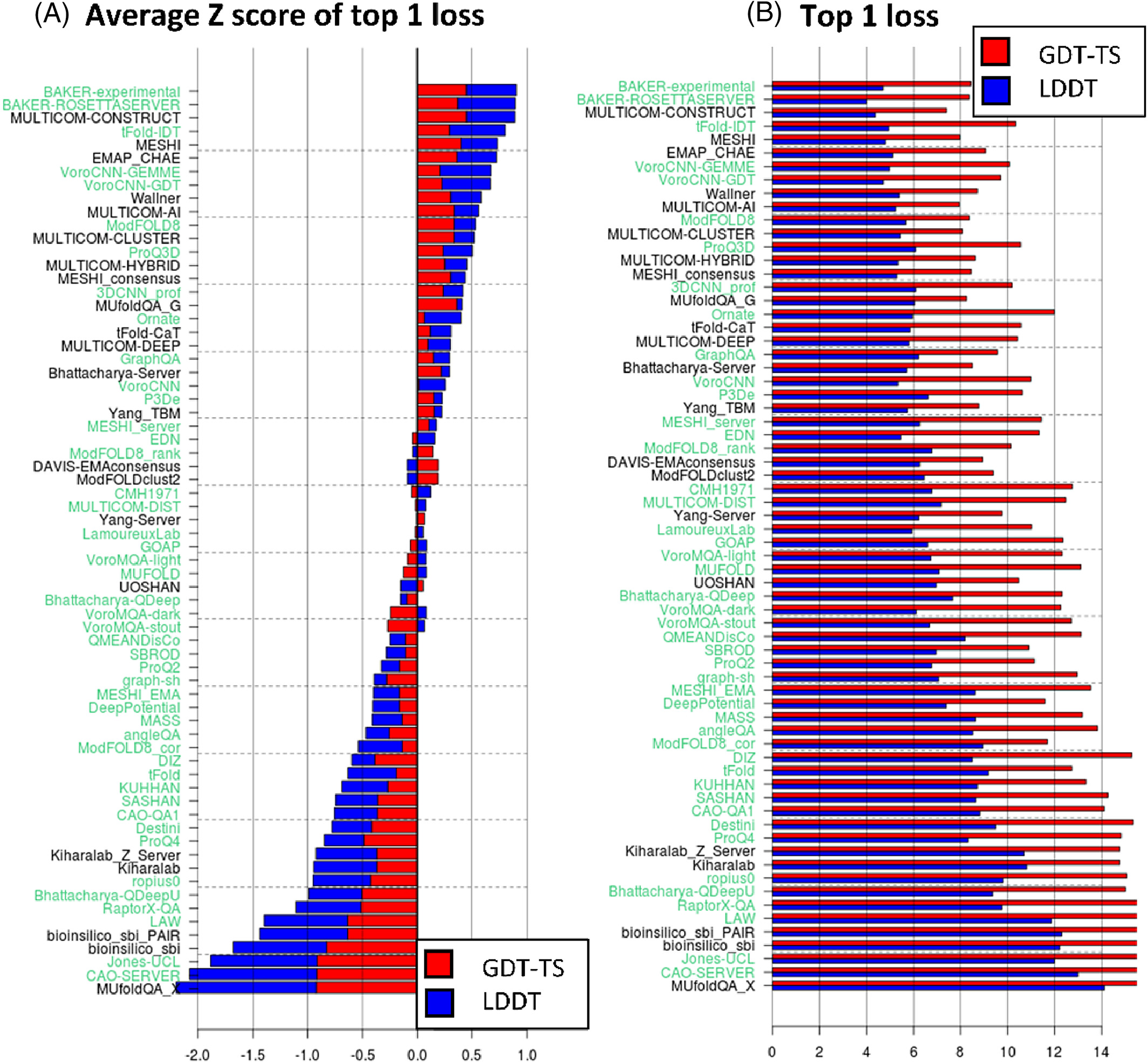
AlphaFold1 achieved first place in the free modeling (FM) category with unprecedented margin: summed Z-score of 52.8 versus 36.6 for the second-place Zhang group. The system made 25 of the 43 best predictions in the FM category, achieving median GDT_TS of 58.9 (versus 52.5 for competitors).
Perhaps more striking: 24 of 43 FM domains achieved TM-score ≥ 0.7, compared to just 14 for the next best method. The improvement represented roughly “two CASPs worth” of progress in a single competition—a harbinger of what was to come.
AlphaFold2: End-to-end differentiable structure prediction
AlphaFold2, presented at CASP14 in November 2020 and published in Nature in July 2021 (Jumper et al.), represented a fundamental architectural rethinking. Rather than predicting distances then assembling structures, AF2 predicted 3D atomic coordinates directly through an end-to-end differentiable pipeline. The result was a 2.7× improvement in summed Z-scores over the next best method and median backbone accuracy competitive with experimental techniques.
1. The Evoformer: A neural network for structural reasoning
The Evoformer forms the “trunk” of AlphaFold2, processing two complementary representations through 48 blocks with non-shared weights:
MSA representation (N_seq × N_res × 256 channels): Encodes evolutionary information from multiple sequence alignments. Rows represent homologous sequences; columns represent residue positions. Co-evolutionary signals—correlated mutations indicating structural proximity—are captured through attention mechanisms operating across both dimensions.
Pair representation (N_res × N_res × 128 channels): Encodes pairwise relationships between residues, representing edges in a graph where nodes are amino acids. This representation directly captures distance and orientation information critical for structure generation.
The key insight is that these representations communicate continuously. MSA information improves pairwise understanding through outer product operations, while pairwise information guides MSA interpretation through attention biases. This bidirectional communication enables the network to reason jointly about evolutionary patterns and structural constraints.
2. Row-wise and column-wise attention
The Evoformer processes the MSA representation through factorized attention:
Row-wise gated self-attention operates horizontally across each sequence, identifying which amino acids within a sequence relate to each other. Critically, the pair representation contributes bias terms to the attention logits—projections from the pairwise embeddings are added directly to attention weights, allowing structural information to guide sequence interpretation.
Column-wise gated self-attention operates vertically at each position, identifying which sequences in the MSA are most informative. This propagates structural information between homologous sequences, effectively learning which evolutionary relationships are most relevant for structure prediction.
The factorization—separating row and column attention rather than computing full N_seq × N_res × N_res attention—reduces memory requirements dramatically while preserving expressivity.
3. Triangle operations enforce geometric consistency
For a pairwise representation to correspond to a physically realizable 3D structure, distances must satisfy the triangle inequality: $d(i,j) ≤ d(i,k) + d(k,j)$. The Evoformer enforces this constraint through four complementary operations:
Triangular multiplicative updates (outgoing and incoming) update edge representations using products of edges sharing a common vertex. The outgoing update modifies edge $z_{ij}$ using all pairs $z_{ik}$ and $z_{kj}$ (triangles sharing starting node $i$), while the incoming update uses $z_{ki}$ and $z_{kj}$ (triangles sharing ending node $j$). These operations use elementwise multiplication rather than dot products, achieving O(N_res²) rather than O(N_res³) complexity.
Triangular self-attention (starting and ending node) computes attention over edges sharing a common vertex, with bias terms from the third edge in each triangle. This learns geometric and chemical constraints while ensuring balanced attention across residue positions.
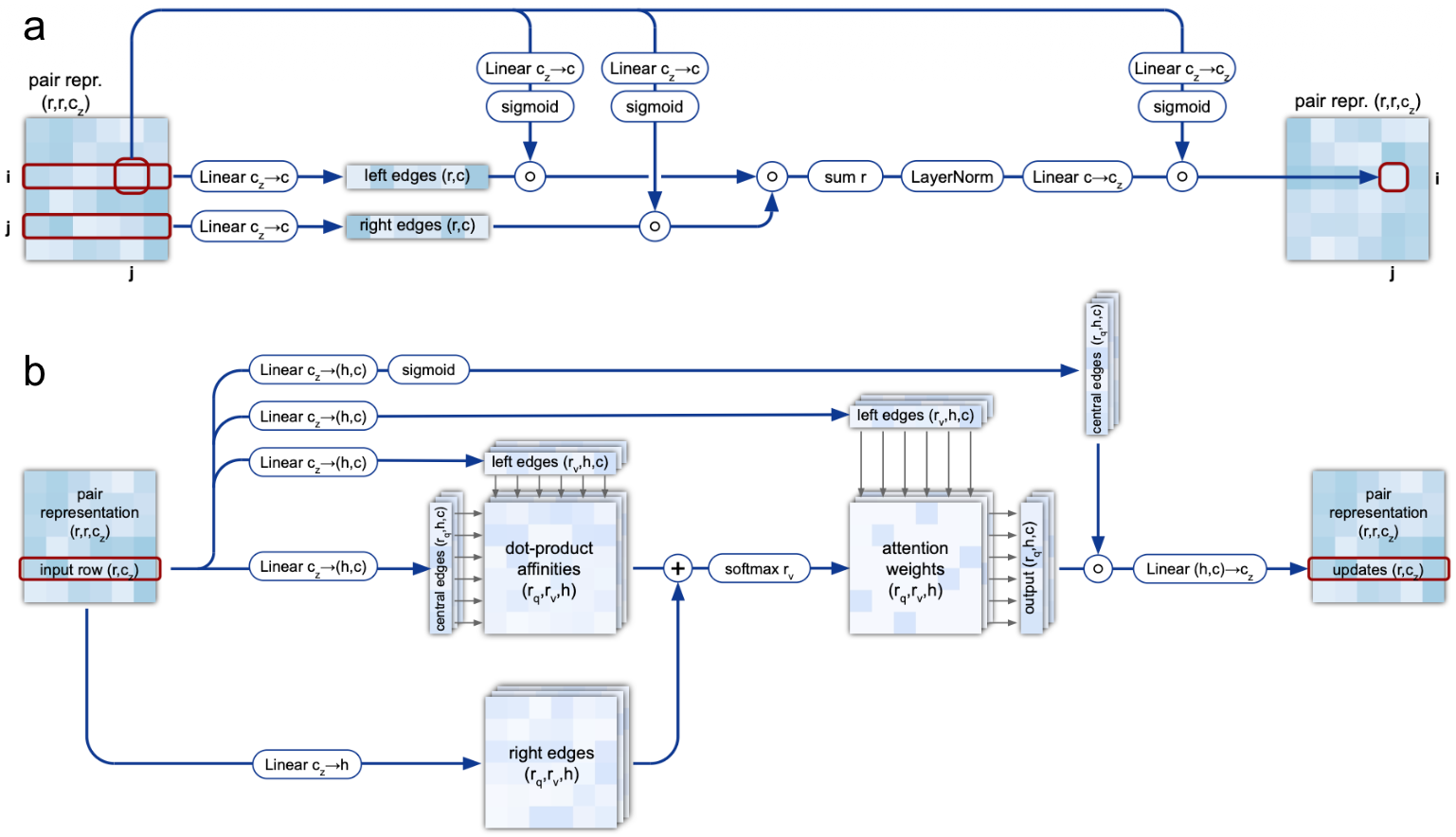
4. The outer product mean bridges representations
Information flows from MSA to pair representation through the outer product mean operation. For each residue pair $(i, j)$, the network computes outer products of MSA embeddings at positions $i$ and $j$, then averages over all sequences in the alignment. This captures covariation signals—when positions $i$ and $j$ show correlated mutations across evolution, the outer product will reflect this coupling.
Unlike earlier methods that computed covariation statistics once during preprocessing, the Evoformer applies outer product mean within every block, enabling iterative refinement as MSA embeddings themselves improve.
5. Structure Module: From representations to coordinates
The Structure Module converts abstract Evoformer representations into explicit 3D coordinates through 8 blocks with shared weights. Its central innovation is Invariant Point Attention (IPA), which computes geometry-aware attention while maintaining SE(3) invariance.
Invariant Point Attention combines three information sources:
- Standard attention on single representation (queries, keys, values)
- Pair representation contributing bias to attention logits
- 3D point attention—the novel component
For 3D point attention, the network projects the single representation into query, key, and value points in each residue’s local coordinate frame. These points are transformed to a global frame using backbone frames $T_i$, then attention weights incorporate squared Euclidean distances between query and key points:
\[w_{ij} = \mathrm{softmax}(q_i · k_j + b_{ij} + \gamma · \sum||T_i(q_{\mathrm{point}}) - T_j(k_{\mathrm{point}})||^2)\]Because the L2-norm distance computation uses coordinates in the global frame, applying any rigid transformation to all frames identically cancels out. The network learns representations that are truly invariant to the choice of global coordinate system—essential since protein structure doesn’t depend on external reference frames.
The Structure Module represents each residue as an independent backbone frame (rotation matrix + translation vector) defining the $\mathrm{N-C_{\alpha}-C}$ geometry. All frames initialize to identity/origin, then update iteratively through IPA. Side-chain torsion angles ($\chi_1$, $\chi_2$, etc.) are predicted by small per-residue networks operating on final activations, represented as unit vectors on the circle (sine/cosine pairs) to handle angular periodicity.
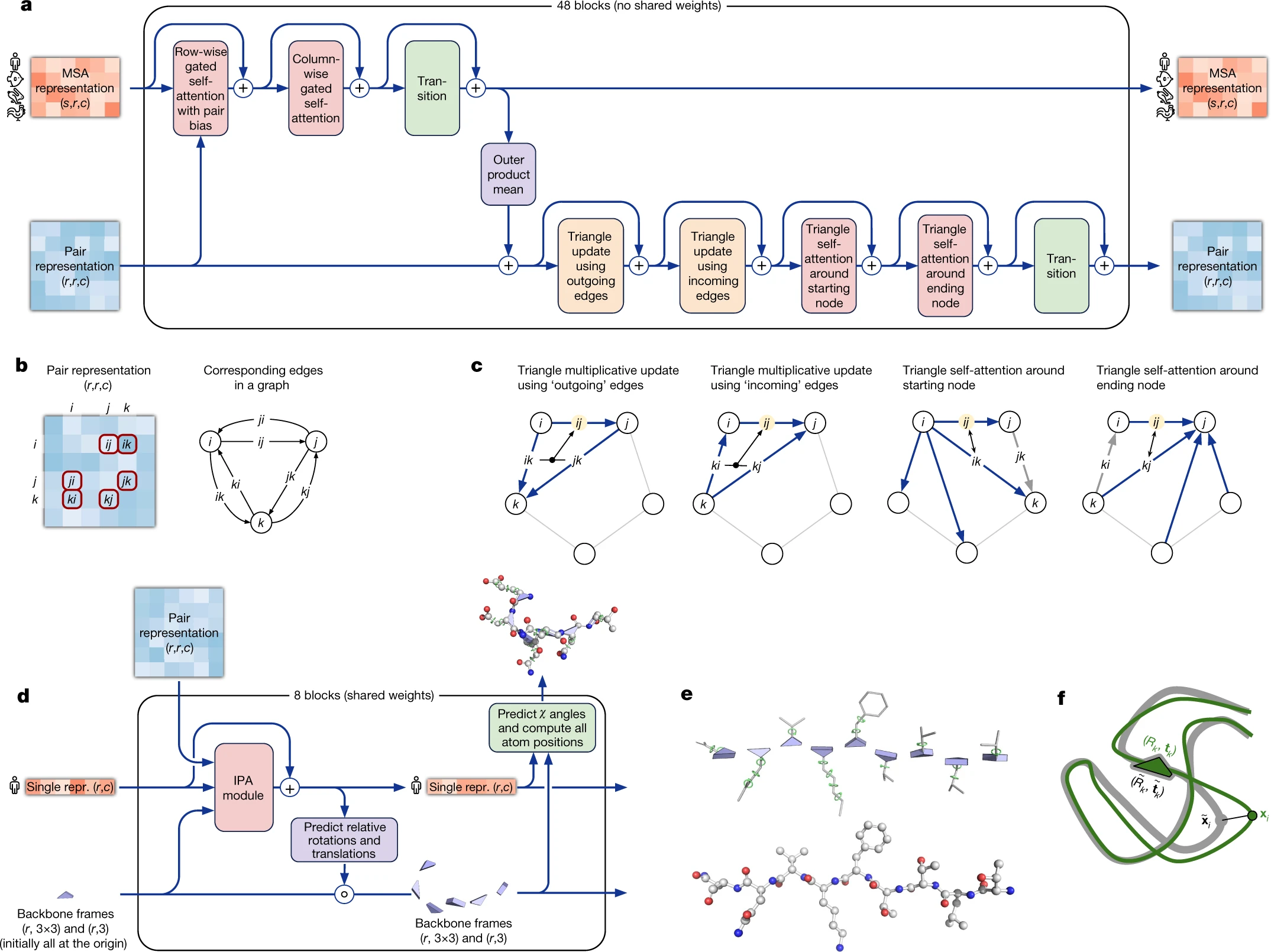
6. Training innovations: FAPE loss and self-distillation
Frame Aligned Point Error (FAPE) loss compares predicted and true atom positions under multiple different alignments. For each alignment frame, the predicted structure is superposed onto the true structure, then distances computed for all atoms. This multi-frame approach captures both local accuracy (when using local alignment frames) and global accuracy (when using global frames).
Critically, FAPE is chirality-sensitive—it is not invariant to reflections, preventing the network from producing mirror-image structures with correct distances but wrong handedness.
Self-distillation proved essential for achieving top accuracy. The training pipeline:
- Train initial model on ~100,000 PDB structures
- Use trained model to predict structures for ~350,000 diverse Uniclust30 sequences
- Filter to high-confidence predictions (pLDDT > 0.5)
- Retrain from scratch on mixture: 25% PDB, 75% predicted structures
This effectively leverages vast unlabeled sequence data, contributing ~1.5 GDT improvement in ablation studies.
7. Recycling enables iterative refinement
AlphaFold2’s recycling mechanism feeds outputs back as inputs for multiple passes (typically 3 iterations). Each iteration receives the previous backbone coordinates and pair representation, enabling progressive refinement. Gradients are stopped between recycling iterations during training, with loss computed after a random number of iterations.
Recycling functions as adaptive computation: easy targets converge quickly while difficult targets benefit from extended search. This architectural choice enables far deeper effective computation without proportional parameter increases.
8. CASP14: Unprecedented accuracy
AlphaFold2’s CASP14 performance shocked the structural biology community:
- Median GDT_TS: 92.4 (out of 100)
- Median Cα RMSD: 0.96 Å (next best: 2.8 Å)
- Median all-atom RMSD: 1.5 Å (next best: 3.5 Å)
- Summed Z-scores: 244.0 versus 90.8 for second place
For reference, a carbon atom has diameter ~1.4 Å. AlphaFold2 was predicting protein structures to within approximately one atom’s width—accuracy that had never been achieved computationally and often took experimental labs years of crystallography effort.
The system made the best prediction for 88 of 97 targets, including challenging cases like the 2,180-residue T1044 RNA polymerase and T1064 (SARS-CoV-2 ORF8), predicted with GDT_TS 87 despite being a free modeling target with no available templates.
AlphaFold3: Diffusion models and unified biomolecular prediction
AlphaFold3 (Abramson et al., Nature 2024) extends structure prediction beyond single proteins to encompass the full complexity of biomolecular interactions: protein-protein complexes, protein-nucleic acid assemblies, small molecule binding, ion coordination, and covalent modifications—all within a single unified framework.
1. From direct prediction to generative modeling
The most fundamental architectural change in AF3 is the shift from direct coordinate prediction to diffusion-based generation. Rather than outputting a single structure, AF3 is a conditional diffusion model that generates a distribution of structures, sampling multiple conformations and ranking by confidence.
The diffusion module operates directly on raw atomic coordinates rather than residue frames and torsion angles. This seemingly simple change eliminates substantial complexity:
- No torsion-based parameterization
- No stereochemical violation losses
- No amino-acid-specific frames
- Full compatibility with arbitrary molecules (ligands, nucleic acids, ions)
During training, atomic coordinates are corrupted with noise at various scales. The network learns to predict true coordinates from noised versions, with different noise levels teaching different structural features: low noise improves local stereochemistry while high noise captures global structure.
At inference, structures are initialized from random noise (sampled from a Gaussian distribution) and iteratively denoised through 200 steps. By default, AF3 generates 5 diffusion samples per model seed, ranking by confidence metrics to select the best prediction.
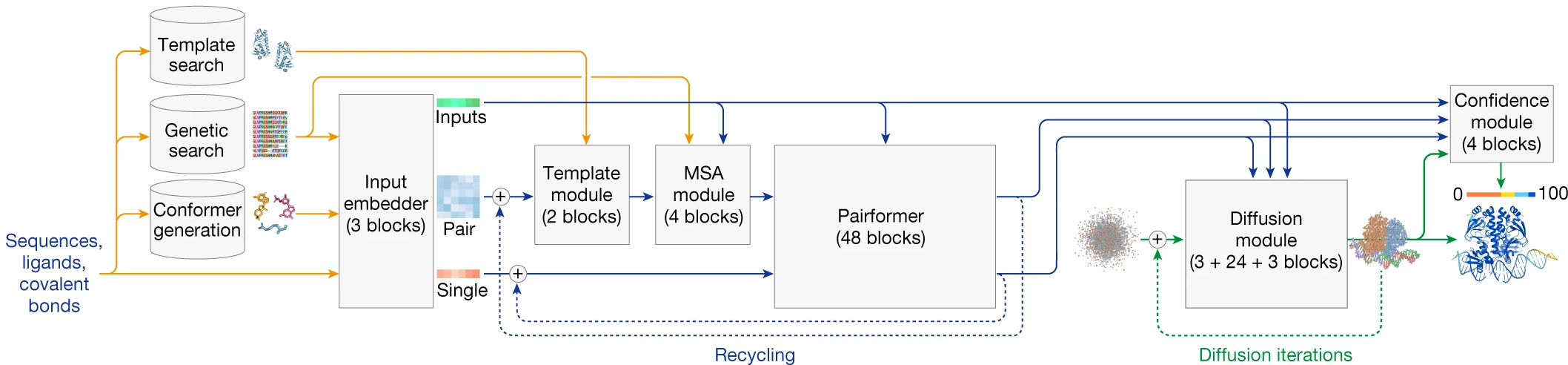
2. Pairformer: A streamlined Evoformer
AF3 replaces the Evoformer with a Pairformer that de-emphasizes MSA processing in favor of focusing computational power on pairwise relationships:
| Feature | AF2 Evoformer | AF3 Pairformer |
|---|---|---|
| Blocks | 48 | 48 |
| Sequence-level | MSA representation (row + column attention) | Single representation (row-wise only) |
| MSA processing | Throughout all blocks | Separate 4-block MSA Module |
| MSA→Pair | Outer product mean every block | Only in MSA Module |
The separate MSA Module processes alignments through just 4 blocks using pair-weighted averaging rather than full key-query attention, then passes only the first row (single sequence representation) to the Pairformer. Triangle multiplicative updates and triangle self-attention are retained from AF2, maintaining geometric reasoning capabilities.
This restructuring reflects an empirical finding: the elaborate MSA processing in the Evoformer may have been overkill. A simpler MSA treatment combined with stronger pairwise reasoning achieves comparable or better accuracy with a more unified architecture.
3. Unified tokenization for all molecules
AF3 handles diverse molecular types through a unified tokenization strategy:
| Molecule Type | Tokenization | Token Center |
|---|---|---|
| Standard amino acids | One token per residue | Cα |
| Standard nucleotides | One token per residue | C1’ |
| Modified residues | Per-atom | Each heavy atom |
| Ligands | Per-atom | Each heavy atom |
| Ions | Per-atom | Single atom |
All tokens share the same embedding space, enabling the network to reason about protein-nucleic acid interfaces, ligand binding pockets, and ion coordination using identical attention mechanisms. The maximum token length is 5,000, accommodating large complexes including ribosomal structures.
4. Eliminating equivariance constraints
Surprisingly, AF3 abandons the SE(3) equivariance that was central to AF2’s Structure Module. The authors found that operating directly on raw coordinates without equivariance constraints works well when using diffusion-based training—the network learns appropriate invariances from data rather than having them architecturally enforced.
This simplification enables a two-level attention scheme: atom-level attention operates on individual atomic positions while token-level attention aggregates across residues or molecules. The combination achieves expressive structure generation without the complexity of Invariant Point Attention.
5. Cross-distillation addresses hallucination
Diffusion models can generate plausible-looking structures in regions where the true structure is disordered. To combat this hallucination tendency, AF3 employs cross-distillation: training data is enriched with predictions from AlphaFold-Multimer v2.3, which produces extended loops in disordered regions rather than compact hallucinated structures.
This teaches AF3 to generate appropriately uncertain predictions for disordered regions—marked with low confidence scores and extended conformations rather than confidently incorrect compact structures.
6. Performance across molecular interaction types
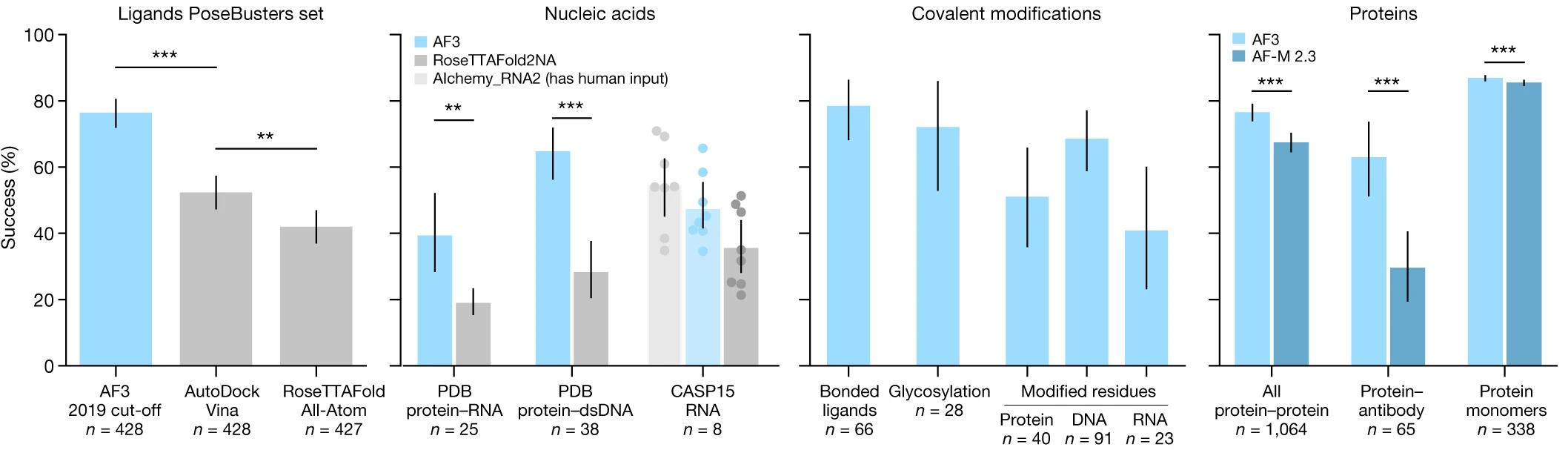
AF3 demonstrates improved accuracy across nearly all biomolecular interaction types:
Protein-ligand binding (PoseBusters benchmark): 76.4% success rate (pocket-aligned RMSD < 2 Å), significantly outperforming AutoDock Vina (P = 2.27 × 10⁻¹³) and RoseTTAFold All-Atom (P = 4.45 × 10⁻²⁵). This represents the first time a deep learning method has substantially exceeded traditional docking approaches without requiring pre-defined binding pockets.
Protein-nucleic acid complexes: Significantly outperforms RoseTTAFold2NA on both RNA (P = 1.6 × 10⁻⁷) and DNA (P = 5.2 × 10⁻¹²) complexes. Successfully handles structures with thousands of residues, including 7,663-residue ribosomal complexes.
Protein-protein interfaces: Improved DockQ scores versus AlphaFold-Multimer v2.3, with particularly marked improvement on antibody-antigen interfaces (P = 6.5 × 10⁻⁵) when using high seed counts (up to 1,000).
Covalent modifications: Accurately predicts glycosylation (~46% success for single-residue glycans), phosphorylation, and other post-translational modifications.
Architectural evolution summarized
The progression from AF1 to AF3 reflects increasingly sophisticated approaches to the same fundamental challenge:
| Aspect | AlphaFold1 | AlphaFold2 | AlphaFold3 |
|---|---|---|---|
| Core approach | Distance prediction + optimization | End-to-end coordinate prediction | Diffusion-based generation |
| Output | Distance distributions | Single structure | Structure distribution |
| Main architecture | Dilated ResNet (220 blocks) | Evoformer (48 blocks) + Structure Module (8 blocks) | MSA Module (4 blocks) + Pairformer (48 blocks) + Diffusion Module |
| Coordinate generation | L-BFGS gradient descent | IPA with SE(3) equivariance | Direct diffusion on raw coordinates |
| Primary loss | Cross-entropy on distance bins | FAPE (Frame Aligned Point Error) | Weighted MSE + smooth LDDT |
| Molecule types | Single-chain proteins | Proteins + multimers | Proteins, nucleic acids, ligands, ions |
| CASP performance | CASP13: GDT 58.9 (FM median) | CASP14: GDT 92.4 (overall median) | State-of-the-art across complex types |
Impact on structural biology and beyond
1. The AlphaFold Database transformed accessibility
The AlphaFold Protein Structure Database, launched in partnership with EMBL-EBI in July 2021, initially contained ~350,000 structures including the complete human proteome. By July 2022, expansion to over 214 million structures covered nearly all catalogued proteins—more structures than experimental methods had determined in 50 years of structural biology.
Over 3 million users from 190+ countries have accessed the database. The AlphaFold2 paper has been cited over 20,000 times, making it one of the most-cited publications in scientific history.
2. Experimental structural biology adapted rather than disappeared
Rather than rendering experimental structural biology obsolete, AlphaFold has transformed how experiments are conducted. For X-ray crystallography, AlphaFold models serve as molecular replacement templates, eliminating the need for selenomethionine phasing in most cases. 30 of 32 CASP14 models successfully served as search models for molecular replacement.
For cryo-EM, AlphaFold predictions guide model building into density maps. Studies demonstrate that 22 of 25 AlphaFold2 models successfully refined against high-resolution cryo-EM maps (up to ~4 Å resolution) with over 90% alpha-carbon accuracy.
The relationship is complementary: AlphaFold provides starting models, experiments provide validation and capture conformational states, dynamics, and interactions that prediction cannot access.
3. Drug discovery applications emerge
AlphaFold has enabled structure-based drug discovery for targets previously lacking experimental structures:
CDK20 inhibitor discovery: First demonstration of AlphaFold-guided hit identification achieved a CDK20 inhibitor (Kd = 566.7 nM, IC50 = 208.7 nM) within 30 days from target selection, synthesizing only 7 compounds—a dramatic acceleration over traditional timelines.
Malaria vaccine development: Oxford University researchers used AlphaFold to determine the structure of Pfs48/45 protein, critical for transmission-blocking malaria vaccines. Years of X-ray crystallography had yielded only low-resolution images; AlphaFold provided the “really sharp view” needed to advance vaccine design to human clinical trials.
Neglected tropical diseases: The Drugs for Neglected Diseases Initiative is using AlphaFold to develop medicines for Chagas disease and leishmaniasis, with over 20 new chemical entities now in their portfolio.
4. Spawning a new ecosystem of methods
AlphaFold’s success catalyzed an ecosystem of related methods:
RoseTTAFold (David Baker lab): Three-track neural network achieving competitive accuracy with AF2, later extended to RFdiffusion for protein backbone generation and RFdiffusion3 for all-atom design.
ESMFold (Meta AI): 15-billion parameter language model achieving 60× faster prediction than AF2 for short sequences, requiring no MSA computation. Demonstrates that evolutionary information can be learned implicitly from massive sequence datasets.
OpenFold: Fully open-source, trainable PyTorch reproduction of AlphaFold2 under Apache License 2.0, enabling academic research on architectural variations. OpenFold3-preview now replicates AF3 capabilities.
ColabFold: Free accessible platform using MMseqs2 for 40-60× faster MSA generation, democratizing access through Google Colab notebooks.
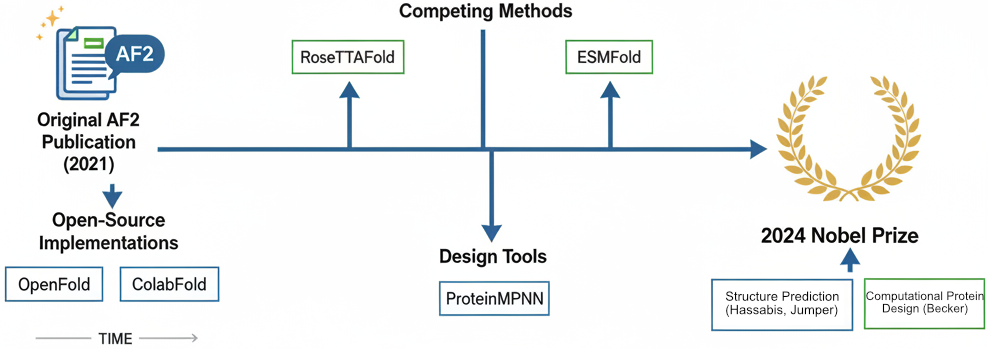
5. Recognition: The Nobel Prize
In October 2024, Demis Hassabis and John Jumper received the Nobel Prize in Chemistry “for protein structure prediction,” sharing the award with David Baker “for computational protein design.” The Royal Swedish Academy described AlphaFold as solving “a 50-year-old problem”—the protein folding challenge that had defined computational biology since Anfinsen’s work in the 1960s.
This represented the first Nobel Prize awarded primarily for an AI-based scientific tool, signaling recognition that machine learning had achieved a transformative breakthrough in natural science.
Limitations and open challenges
Despite revolutionary accuracy, significant limitations remain:
Single conformations only: AlphaFold predicts one structure per input, unable to capture conformational ensembles or dynamics essential for understanding allostery, cryptic binding sites, and functional mechanisms.
Mutation effects: Point mutations often yield predictions nearly identical to wild-type, despite potentially dramatic effects on stability, function, or disease phenotype.
Intrinsically disordered regions: The ~30% of the human proteome that lacks stable 3D structure appears as low-confidence “unrealistic swirls.” While pLDDT < 50 correctly flags these regions, their functional importance in signaling and phase separation remains inaccessible to structure prediction.
Conformational changes upon binding: AlphaFold cannot predict how ligand or partner binding induces conformational change—a fundamental limitation for understanding induced-fit binding and allosteric regulation.
These limitations define the frontier for future development. Methods addressing conformational ensembles, dynamics, and mutation effects remain active research areas, building on the foundation AlphaFold has established.
Conclusion
The evolution from AlphaFold1 to AlphaFold3 demonstrates how architectural innovations compound. AF1’s insight that full distance distributions outperform binary contacts led to AF2’s recognition that end-to-end differentiable structure prediction eliminates the information bottleneck of intermediate representations. AF3’s adoption of diffusion models and unified molecular tokenization extends this logic, replacing specialized components with general-purpose generative modeling.
The technical lessons extend beyond protein structure. Triangle operations enforcing geometric consistency, invariant point attention respecting physical symmetries, recycling mechanisms enabling iterative refinement, and self-distillation leveraging unlabeled data represent architectural patterns applicable across scientific machine learning.
For structural biology specifically, AlphaFold has shifted the field from “can we determine this structure?” to “what biological questions can we now answer?” With 214 million predicted structures freely available and AF3 extending prediction to molecular complexes, the bottleneck has moved from structure determination to experimental validation and functional interpretation—a transformation that will shape biological research for decades.
Key references
AlphaFold1
- Senior, A.W. et al. “Improved protein structure prediction using potentials from deep learning.” Nature 577, 706–710 (2020). DOI: 10.1038/s41586-019-1923-7
AlphaFold2
- Jumper, J. et al. “Highly accurate protein structure prediction with AlphaFold.” Nature 596, 583–589 (2021). DOI: 10.1038/s41586-021-03819-2
AlphaFold3
- Abramson, J. et al. “Accurate structure prediction of biomolecular interactions with AlphaFold 3.” Nature 630, 493–500 (2024). DOI: 10.1038/s41586-024-07487-w
CASP Assessments
- Kryshtafovych, A. et al. “Critical assessment of methods of protein structure prediction (CASP)—Round XIII.” Proteins 87, 1011–1020 (2019).
- CASP14 assessment papers, Proteins (2021)
Impact and Database
- Tunyasuvunakool, K. et al. “Highly accurate protein structure prediction for the human proteome.” Nature 596, 590–596 (2021).
- AlphaFold Database: https://alphafold.ebi.ac.uk/
Enjoy Reading This Article?
Here are some more articles you might like to read next: