Inside AlphaFold2
a technical deep dive into the architecture
DeepMind’s AlphaFold2 transformed structural biology by achieving near-experimental accuracy in protein structure prediction, fundamentally solving a 50-year grand challenge. At CASP14, AlphaFold2 achieved a median backbone RMSD of 0.96 Å—roughly the width of a carbon atom—while the next-best method managed only 2.8 Å. This post dissects the architectural innovations that made this breakthrough possible: the Evoformer’s bidirectional attention mechanisms, novel triangular updates for geometric consistency, the SE(3)-equivariant Structure Module, and the elegant recycling mechanism that enables iterative refinement. Understanding these components is essential for anyone working with or building upon modern protein structure prediction systems.
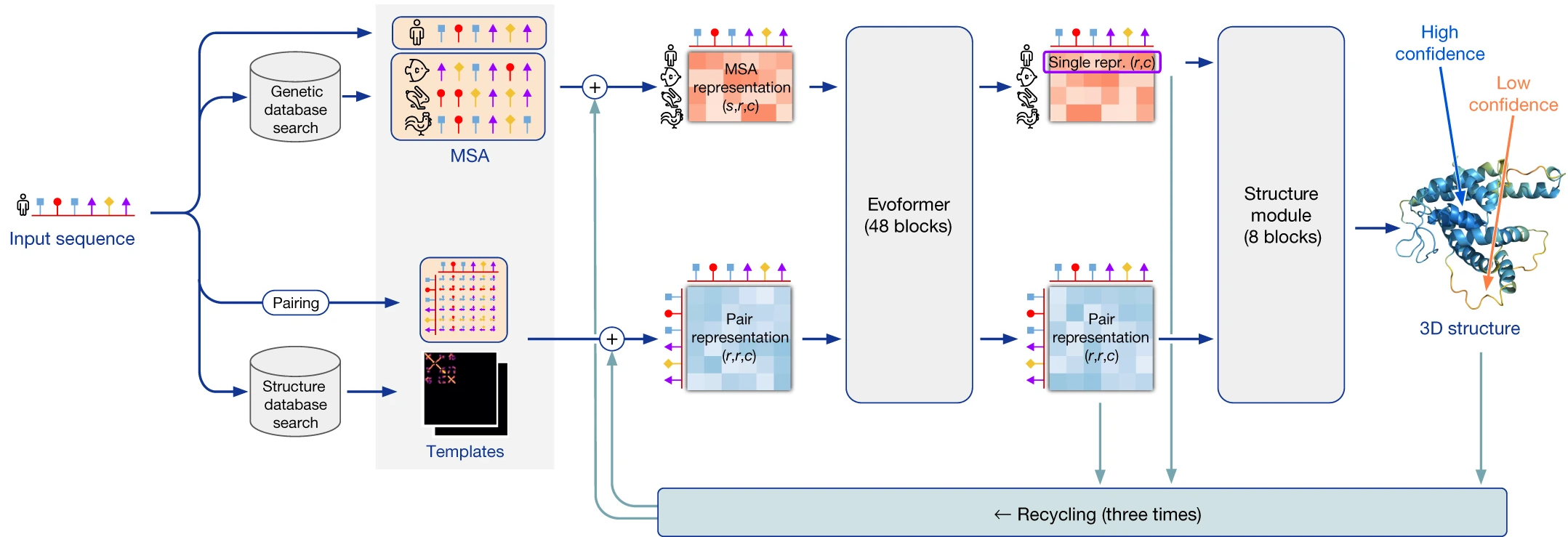
From sequence to structure: the complete AlphaFold2 pipeline
AlphaFold2 predicts all-atom 3D coordinates directly from a protein’s amino acid sequence through three major stages: input embedding, representation learning via the Evoformer, and coordinate generation through the Structure Module. Unlike its predecessor AlphaFold1, which predicted distance distributions and then optimized coordinates separately, AlphaFold2 is end-to-end differentiable, allowing gradients to flow from the final 3D loss back through the entire network.
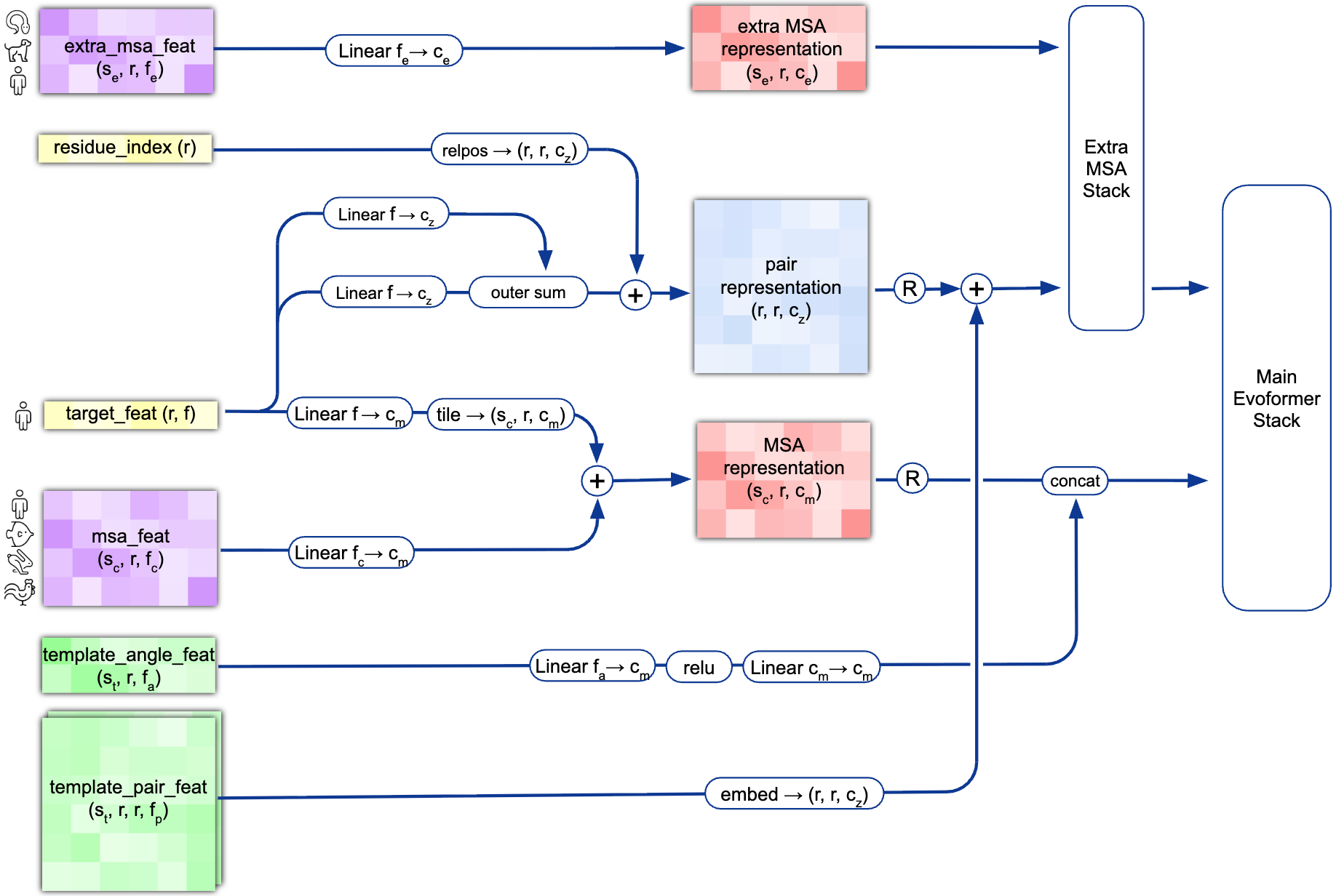
The pipeline begins with extensive MSA construction. AlphaFold2 queries multiple sequence databases using JackHMMer (against UniRef90 and MGnify) and HHBlits (against Uniclust30 and BFD), gathering up to 15,000+ homologous sequences. These sequences are clustered to create $N_{clust}$ (~512) cluster centers with associated profiles, while additional unclustered sequences (~5,120) are processed separately through the Extra MSA Stack. Template structures, when available, are identified via HHSearch against PDB70 and processed through dedicated template embedding modules.
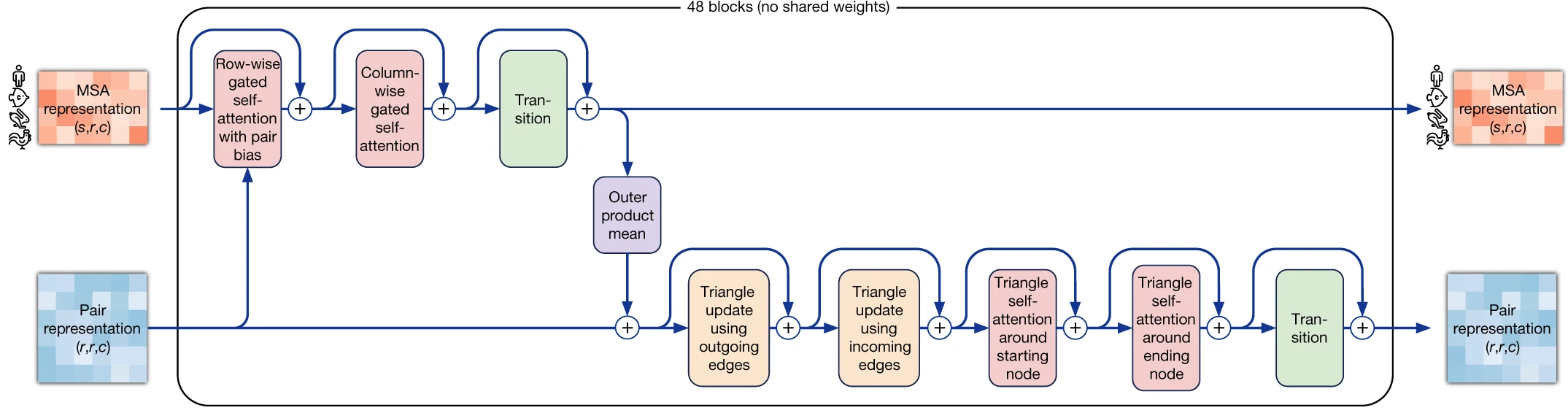
The two core representations that flow through the network are the MSA representation (shape: $N_{seq} \times N_{res} \times 256$) encoding evolutionary relationships across homologous sequences, and the pair representation (shape: $N_{res} \times N_{res} \times 128$) encoding residue-residue relationships that implicitly capture distance and geometric constraints. Information flows bidirectionally between these representations through 48 Evoformer blocks, with the pair representation biasing MSA attention and the MSA representation updating pair features through outer product operations.
The Evoformer: where evolutionary and structural information merge
The Evoformer constitutes the representational core of AlphaFold2, comprising 48 blocks with untied weights that jointly refine both MSA and pair representations. Each block orchestrates a carefully designed sequence of attention operations and updates that enable the network to reason simultaneously about coevolution patterns and geometric constraints.
1. Row-wise attention captures coevolutionary signals
Within each MSA sequence, row-wise gated self-attention identifies which residue positions correlate with each other—the signature of coevolution. The key innovation here is pair bias injection: attention affinities are augmented by learned projections from the current pair representation:
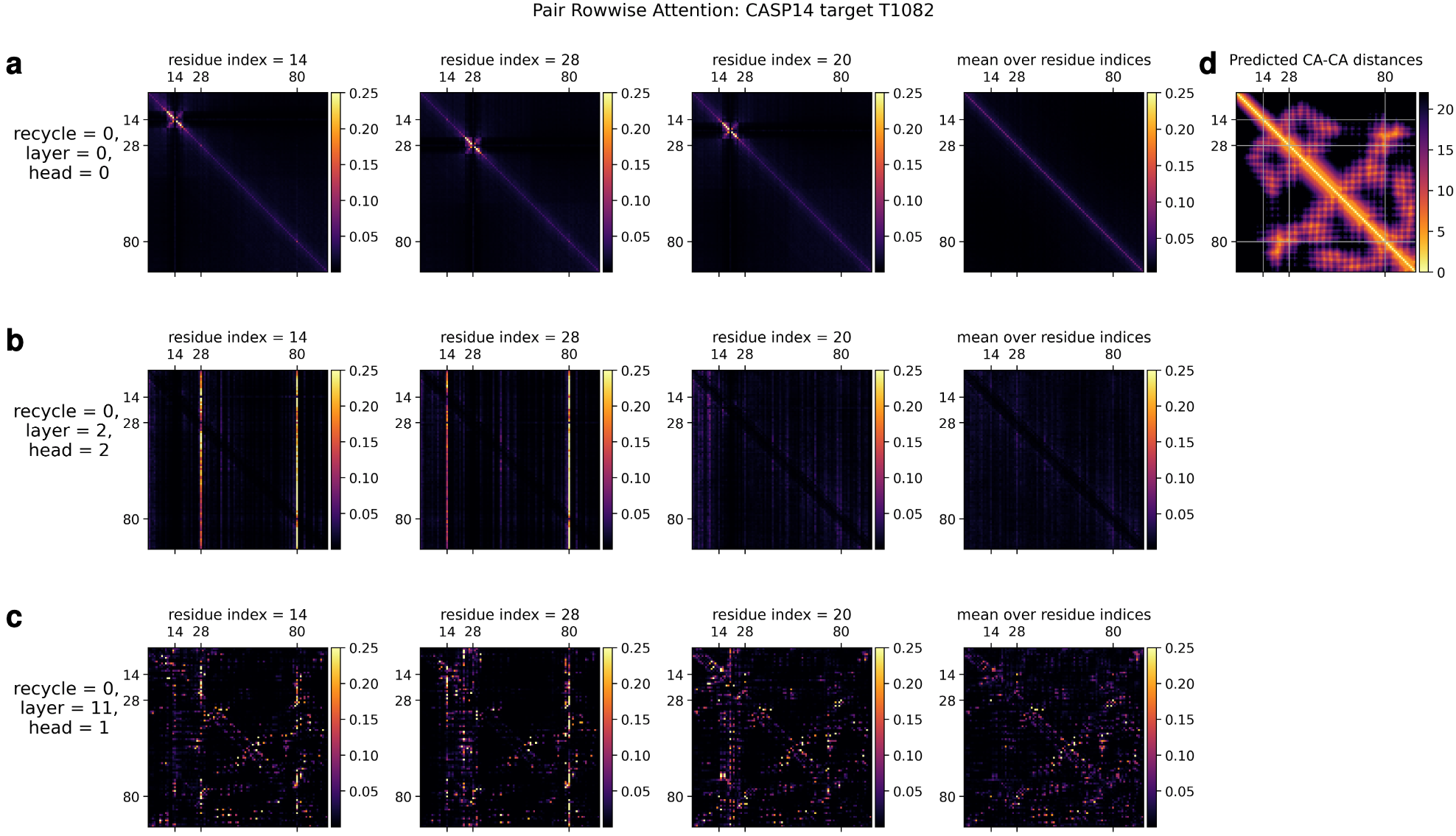
\[a_{ij} = \mathrm{softmax}(\frac{a_i^T j_j}{\sqrt(c + b_{ij})})\]where $b_{ij}$ is derived from the pair representation $z_{ij}$. This creates a powerful feedback loop: the pair representation’s hypothesis about which residues are spatially proximal directly influences how the network interprets coevolutionary patterns. The gating mechanism (sigmoid activation multiplied element-wise with outputs) allows the network to selectively suppress uninformative attention patterns. Empirically, attention maps reveal that the network learns to detect $\alpha$-helix hydrogen bonding patterns (attention radius ~4 residues) and cysteine-cysteine interactions indicative of disulfide bonds.
2. Column-wise attention aggregates across homologs
Column-wise attention operates orthogonally, enabling information exchange at each sequence position across all homologous sequences. This captures evolutionary conservation patterns—highly conserved positions suggest structural or functional importance. Visualization studies show that through recycling iterations, column attention increasingly focuses on the query sequence (first MSA row), effectively learning to weight sequences by their informativeness for the target structure.
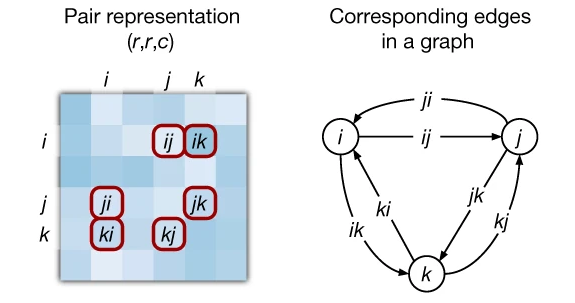
3. Triangular updates enforce geometric consistency
The most architecturally novel components of the Evoformer are the triangular multiplicative updates and triangular self-attention operations on the pair representation. These implement a soft version of the triangle inequality constraint: for any three residues $i, j, k$, the pairwise relationships should be geometrically consistent.
The triangular multiplicative update (outgoing edges) updates edge ($i,j$) by aggregating information from all edges sharing starting node $i$:
\[z_{ij} \leftarrow z_{ij} + g_{ij} \odot \mathrm{Linear}(\mathrm{LayerNorm}(\sum_k a_{ik} \odot b_{kj} ))\]where $a$ and $b$ are gated projections and the sum runs over all intermediate nodes $k$. The Hadamard product (⊙) provides an efficient $O(N^2 \mathrm{c})$ alternative to full $O(N^3)$ attention while capturing similar geometric information. The incoming edges variant operates symmetrically, considering edges sharing ending node $j$.
Triangular self-attention takes this further by adding explicit attention over triangle configurations. When edge ($i,j$) attends to edges ($i,k$), the bias from the third edge ($j,k$) is added to attention affinities—directly encoding the triangle inequality informationally rather than through hard constraints. This design allows the network to learn soft geometric constraints without requiring explicit distance supervision during training.
4. Outer product mean: the MSA-to-pair bridge
The outer product mean operation transfers coevolutionary signal from MSA to pair representations:
\[z*{ij} \mathrel{+}= \mathrm{Linear}(\mathrm{Flatten}(\frac{1}{N*{seq}} \sum*s a*{si} \otimes b\_{sj}))\]This computes the mean outer product between projected MSA columns $i$ and $j$ across all sequences, naturally capturing pairwise covariation—if position $i$ mutates, does position $j$ co-vary? This operation, while used in prior coevolution methods, is deeply integrated into the Evoformer’s iterative refinement loop.
The Structure Module: from abstract features to 3D coordinates
After 48 Evoformer blocks, the Structure Module converts abstract representations into explicit atomic coordinates. This module introduces the elegant residue gas representation: each residue is treated as an independent rigid body (the backbone N-C$\alpha$-C triangle) floating freely in 3D space, parameterized by rotation $R_i \in \mathrm{SO}(3)$ and translation $t_i \in \mathbb{R}^3$. Initially, all residues are placed at the origin with identity rotation—a “black hole” configuration that the network gradually expands into the folded structure.
The Structure Module comprises 8 layers with shared weights, each applying Invariant Point Attention (IPA), transition MLPs, and backbone frame updates. Crucially, gradients into frame orientations are stopped between iterations (Algorithm 20, line 20), preventing “lever effects” from chained rotation compositions and improving training stability.
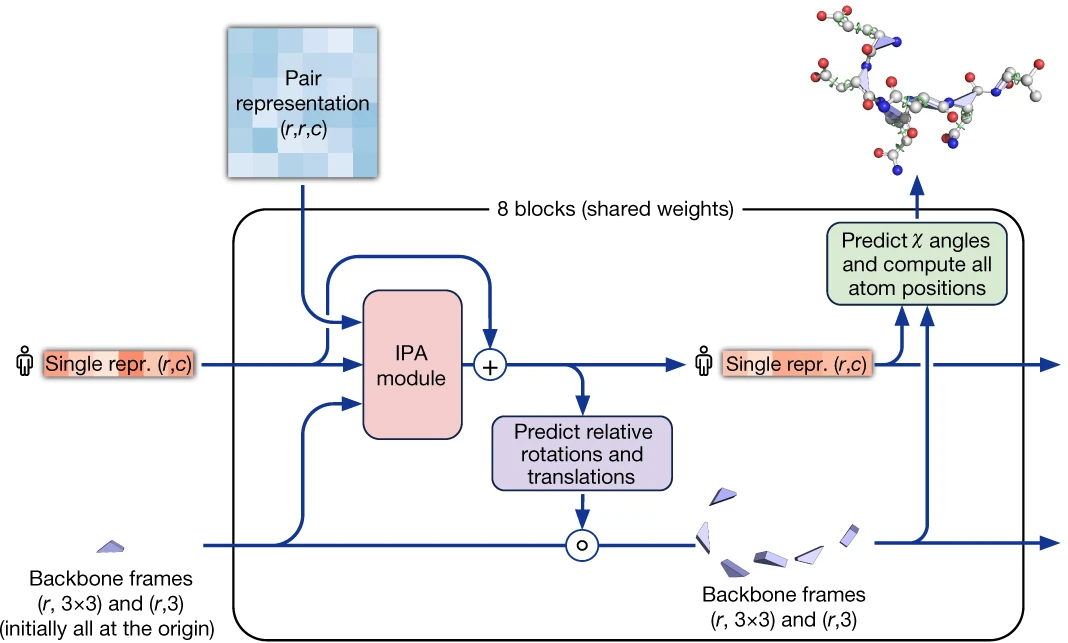
1. Invariant Point Attention achieves SE(3)-equivariance
IPA is the geometric heart of the Structure Module, enabling 3D reasoning while maintaining invariance to global rotations and translations. The attention weight computation combines three terms:
\[a*{ij} = \mathrm{softmax}\_j \left(w_L \left(\frac{q_i \cdot k_j}{\sqrt{c}} +b*{ij} - \frac{\gamma \cdot w_c}{2}\sum_p ||T_i \circ \vec{q}\_i - T_j \circ \vec{k}\_j ||^2 \right) \right)\]The first term is standard scalar attention on the single representation. The second term ($b_{ij}$) injects bias from the pair representation. The third term—the critical geometric component—computes squared distances between query and key points after transformation from local to global coordinates. Points $\vec{q}$ and $\vec{k}$ are generated in each residue’s local frame, transformed to global coordinates via frame compositions $T_i$ and $T_j$, and their squared distances computed.
This squared distance term is SE(3)-invariant: applying any global rigid transformation $T_{\mathrm{global}}$ to all frames leaves the distances unchanged because rotations preserve norms and translations cancel in the difference. The negative sign causes nearby points to receive higher attention weights, implementing a soft spatial locality bias.
The output aggregates three value types: scalar values (standard attention), pair values (from $z_{ij}$), and point values transformed back to local coordinates. The norm of point outputs is also included—an invariant scalar providing magnitude information.
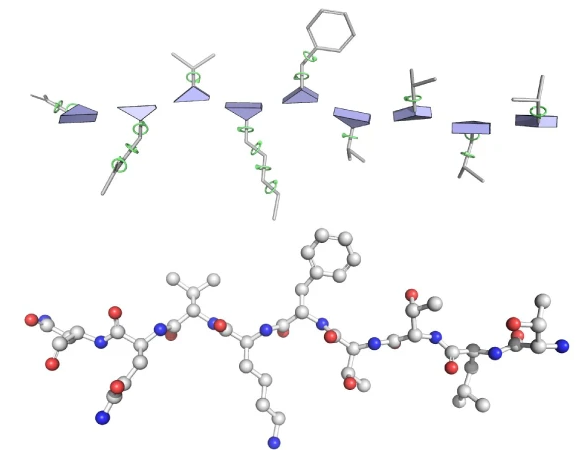
2. From frames to atoms: torsion angles and coordinate generation
Side chains are predicted via 7 torsion angles per residue: three backbone angles (ω, φ, ψ) and four sidechain χ angles. A shallow 2-layer ResNet predicts each angle as ($\sin \theta$, $\cos \theta$) pairs, avoiding discontinuity at $\pm \pi$. Final atomic coordinates emerge by composing backbone frames with torsion-induced rotations applied to ideal residue geometries (fixed bond lengths and angles from amino acid chemistry).
The pLDDT confidence score (0-100 scale) is predicted from the final single representation via a classification head over 50 bins spanning lDDT values. Interpretation guidelines: pLDDT > 90 indicates high confidence with reliable backbone and sidechains; 70-90 suggests generally correct backbone; 50-70 indicates low confidence possibly representing disorder; < 50 should be interpreted as likely disordered.
The Predicted Aligned Error (PAE) matrix predicts position error at residue x when aligning on residue y, enabling inter-domain confidence assessment. Low PAE between domains indicates confident domain packing; high PAE suggests uncertain relative positioning.
Recycling: iterative refinement without backpropagation
The recycling mechanism allows AlphaFold2 to iteratively refine predictions by feeding outputs back as inputs, effectively quadrupling network depth without quadrupling backpropagation cost. The default configuration uses 3 recycling iterations (4 total forward passes), with gradients stopped between iterations.
Three outputs are recycled: the first row of the MSA representation (projected and added to initial MSA features), the pair representation (layer-normalized and projected), and the predicted backbone structure (converted to a binned distogram and projected to pair features). This structure-to-pair recycling is particularly powerful: it allows the network to use its current geometric hypothesis to refine coevolutionary interpretation.
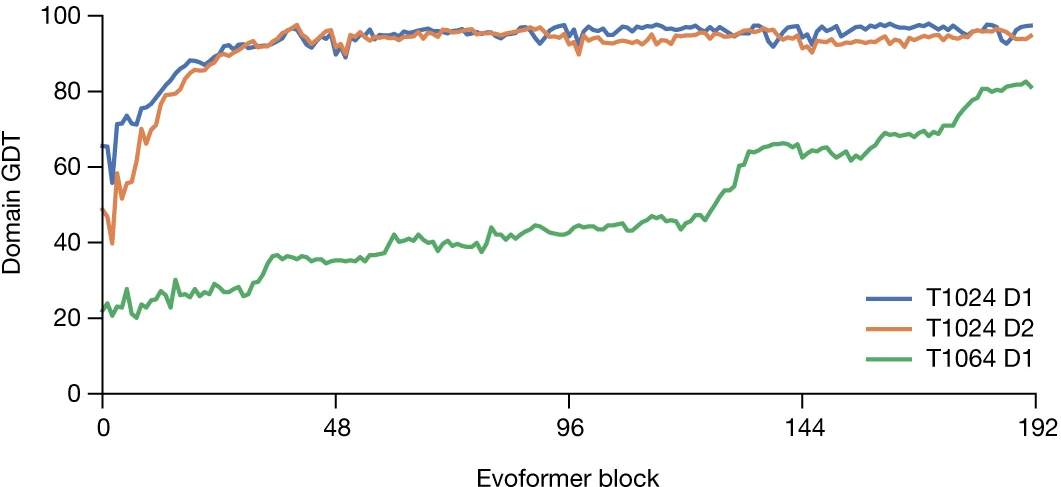
Ablation studies reveal that recycling is more critical than IPA for overall performance—removing recycling causes larger accuracy drops than removing IPA when both are individually ablated. However, without recycling, IPA becomes essential. The combined ablation (no IPA and no recycling) causes severe performance degradation, demonstrating their complementary roles in geometric reasoning.
During training, a random number of recycling iterations is sampled per example, with loss computed only on the sampled iteration. This encourages the model to produce reasonable structures early while still benefiting from refinement. Different MSA samples used each iteration provide data augmentation.
Training methodology: multi-loss supervision and self-distillation
AlphaFold2’s training combines multiple loss functions supervising different aspects of the prediction. The primary structural loss is Frame Aligned Point Error (FAPE), computed by aligning predicted atoms to ground truth under each backbone frame independently:
\[\mathrm{FAPE} = \frac{1}{N_{\mathrm{frames}}N_{\mathrm{atoms}}}\sum_k \sum_i \mathrm{clamp}(||(R_k^{\mathrm{pred}})^{-1}(\vec{x}_i^{\mathrm{pred}} - \vec{t}_k^{\mathrm{pred}}) - (R_{k}^{\mathrm{true}})^{-1}(\vec{x}_{i}^{\mathrm{true}} - \vec{t}_{k}^{\mathrm{true}})||, \pu{10 Å})\]where:
- $\vec{x}_k^{\mathrm{pred}}$ and $\vec{x}_k^{\mathrm{true}}$ : the coordinates of atom $k$ in the global coordinate system for the predicted and true structures, respectively.
- $R_k$, $\vec{t}_k $ : the rotation matrix $R_k$ and translation vector $\vec{t}_k$ that define the rigid body transformation for the $i$-th residue’s local backbone frame (N, C$\alpha$, C atoms).
- The term \(R_k^{-1}(\vec{x}_i - \vec{t}_k)\) transforms a global coordinate $\vec{x}_{i}$ into the local coordinate system of frame $k$.
In simpler terms:
- Establish local frames: For every residue $k$, define a local 3D coordinate system (frame) based on its backbone atoms (N, C$\alpha$, C). Do this for both the predicted and true structures.
- Transform points: For a given pair of residues $i$ and $k$, transform the coordinates of atom $k$ from the global system into the local frame of residue $i$ for both the predicted and true structures.
- Calculate distance: Find the squared Euclidean distance between these two locally transformed points. This distance is invariant to the global orientation of the molecule.
- Clamp: Cap this squared distance at a predefined maximum value (e.g., $10^{2}=\pu{100 Å}$).
- Average: Sum these clamped errors over all pairs of frames and all chosen atoms, and take the average.
FAPE is SE(3)-invariant (independent of global positioning) but not reflection-invariant, correctly penalizing wrong chirality. Clamping at 10Å emphasizes local accuracy while de-emphasizing catastrophic outliers.
Additional losses include:
- Distogram loss: Cross-entropy over discretized inter-residue distances, supervising the pair representation
- Masked MSA loss: BERT-style masked language modeling encouraging the network to learn phylogenetic relationships
- Torsion angle loss: Binned cross-entropy over predicted backbone and sidechain dihedral angles
- Structural violation loss (fine-tuning only): Penalizes unrealistic bond lengths, angles, and steric clashes
- pLDDT and pTM losses: Classification losses for confidence prediction
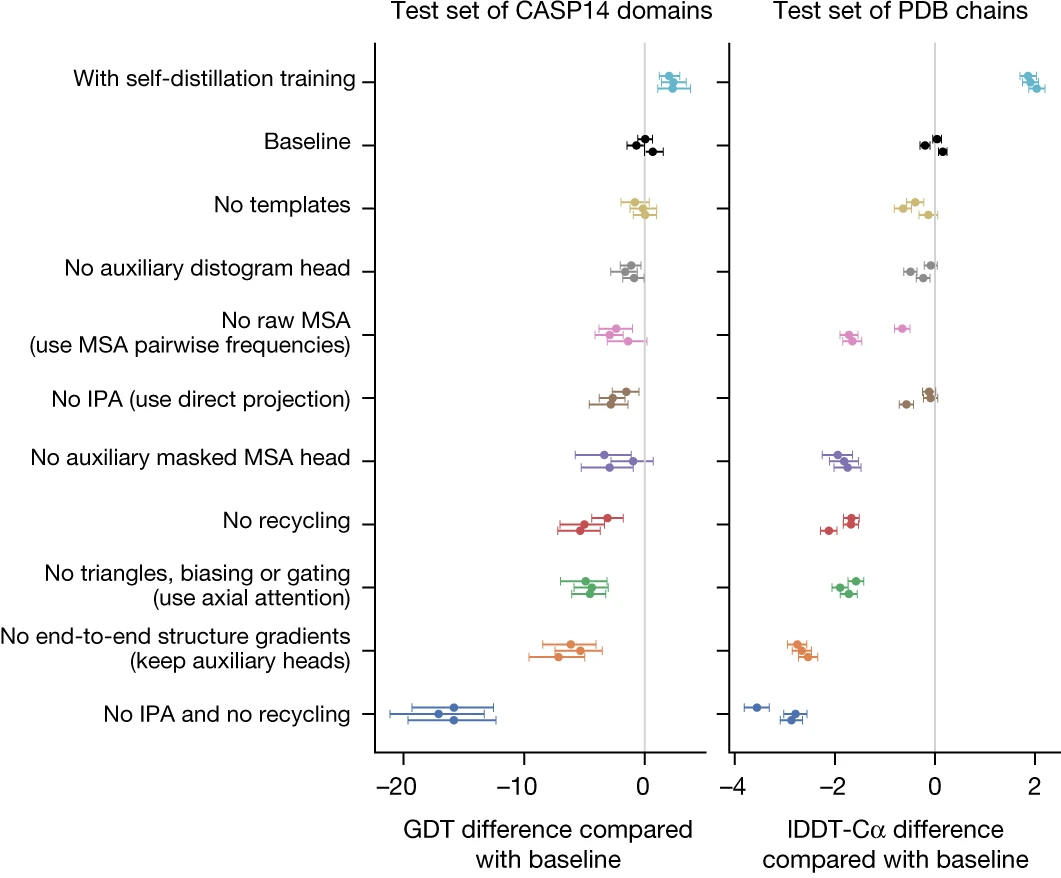
Training proceeds in stages. Initial training uses PDB structures (downloaded August 2019) with 256-residue crops and batch size 128 across 128 TPU v3 cores. Self-distillation then generates ~355,000 predictions on Uniclust30 sequences, filtered to high-confidence predictions (pLDDT > 0.5). The model is retrained from scratch on a mixture of 75% self-distilled structures and 25% PDB structures—this augmentation proves critical, improving accuracy by ~2.7 lDDT-C$\alpha$ points. Fine-tuning extends crops to 384 residues, adds violation losses, and enables pTM prediction.
What made AlphaFold2 succeed: key innovations over prior methods
Several architectural decisions distinguish AlphaFold2 from AlphaFold1 and other CASP14 competitors:
End-to-end differentiability replaced the two-stage approach (predict distances, then optimize coordinates) with direct coordinate prediction. This allows the network to learn structural constraints implicitly rather than through handcrafted optimization, eliminating pipeline brittleness.
Attention-based architecture replaced the convolutional neural networks of AlphaFold1. Transformers naturally capture long-range dependencies without the limited receptive fields that hampered convolution-based contact prediction. The axial attention decomposition (row-wise + column-wise) provides O(N²) scaling rather than O(N⁴) for naive pair attention.
Triangular operations encode geometric constraints informationally rather than literally. Rather than explicitly enforcing triangle inequalities, the network learns to propagate geometrically consistent information through triangle-aware attention patterns.
Residue gas representation enables parallel local refinement. By treating each residue as an independent rigid body initially unconstrained by peptide bond geometry, the network avoids complex loop closure problems during intermediate iterations.
Decoupled learning and state representations, as noted by Mohammed AlQuraishi’s analysis, is perhaps the deepest insight: the 2D pair representation facilitates learning long-range interactions, while the 1D single representation ultimately encodes the structure. The final structure emerges from {$s_i$}, not {$z_{ij}$}, avoiding awkward 2D→3D projection.
Critical implementation parameters
For practitioners implementing or fine-tuning AlphaFold2-derived models, key parameters include:
| Parameter | Value | Description |
|---|---|---|
| Evoformer blocks | 48 | Untied weights per block |
| Structure module layers | 8 | Shared weights |
| MSA representation ($c_m$) | 256 | Channel dimension |
| Pair representation ($c_z$) | 128 | Channel dimension |
| Single representation ($c_s$) | 384 | For structure module |
| Recycling iterations | 3 | Default (plus initial) |
| MSA cluster sequences | ~512 | $N_{clust}$ |
| Extra MSA sequences | ~5,120 | $N_{extra_seq}$ |
| Templates | ≤4 | Maximum used |
| IPA heads | 12 | Per IPA layer |
| Training crop size | 256→384 | Initial→fine-tuning |
Conclusion
AlphaFold2 represents a masterful integration of evolutionary information (via MSA processing), geometric reasoning (via triangular operations and IPA), and iterative refinement (via recycling). The architecture demonstrates that end-to-end learning can discover complex physical constraints without explicit physics-based potentials—the network learns hydrogen bonding patterns, disulfide geometry, and domain packing from data alone.
For computational structural biologists, several insights emerge for future development: the importance of bidirectional information flow between sequence and structural representations, the power of encoding geometric constraints informationally rather than literally, and the effectiveness of iterative refinement through recycling. These principles now underpin AlphaFold3, AlphaFold-Multimer, and numerous derivative methods like ESMFold and OpenFold, continuing to reshape how we approach protein structure prediction.
Primary Reference: Jumper, J., Evans, R., Pritzel, A. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021). https://doi.org/10.1038/s41586-021-03819-2
Supplementary Methods: The 70+ page Supplementary Information contains complete algorithmic specifications (Algorithms 1-32) essential for implementation.
Open Implementations:
- DeepMind AlphaFold: https://github.com/google-deepmind/alphafold
- OpenFold (independent reimplementation): https://github.com/aqlaboratory/openfold
Enjoy Reading This Article?
Here are some more articles you might like to read next: