Understanding AlphaFold3
a quick look at the architecture
AlphaFold3 represents a fundamental architectural reimagining of structure prediction, moving from protein-specific machinery to a unified framework that predicts structures for proteins, nucleic acids, small molecules, ions, and post-translational modifications within a single model. The key innovations—replacing the Evoformer with a simplified Pairformer, substituting the equivariant Structure Module with a diffusion-based generator, and adopting token-based representations—enable 76.4% accuracy on protein-ligand benchmarks and significantly outperform previous methods across virtually all biomolecular interaction categories. This post provides a comprehensive technical walkthrough of the architecture described in Abramson et al., Nature 2024.
The architectural philosophy behind AlphaFold3
AlphaFold3’s design philosophy diverges sharply from its predecessor. Where AlphaFold2 built protein-specific inductive biases directly into the architecture—residue frames, torsion angle parameterization, SE(3)-equivariant Invariant Point Attention—AlphaFold3 strips these away in favor of a more general approach. The core insight is that many of these constraints, while theoretically elegant, added complexity without proportional accuracy gains and fundamentally limited the system to protein-only prediction.
The architecture consists of three main sections executed sequentially. First, input preparation converts sequences and molecular structures into numerical tensors while retrieving evolutionary information through MSA search and structural templates. Second, the representation learning trunk iteratively refines single-residue and pairwise representations through the Template Module, MSA Module, and 48 Pairformer blocks. Third, the diffusion module generates three-dimensional coordinates through iterative denoising, replacing AlphaFold2’s deterministic Structure Module entirely.
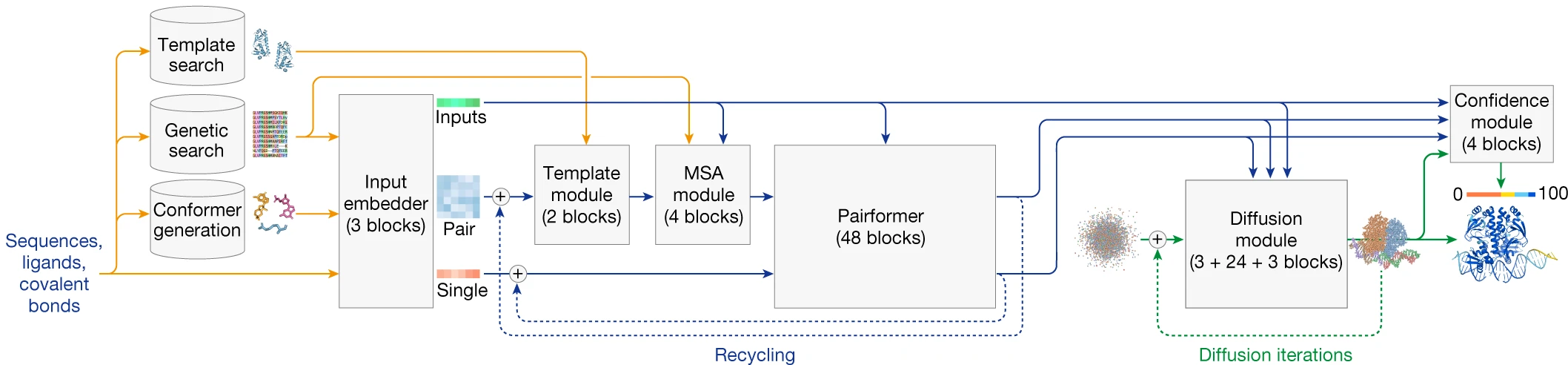
Token-based input representation enables molecular generality
The shift from residue-based to token-based representation is foundational to AlphaFold3’s expanded capabilities. Standard amino acids and nucleotides receive one token per residue, with C$\alpha$ atoms serving as token centers for proteins and C1’ atoms for nucleic acids. Non-standard residues, ligands, ions, and modifications are tokenized at the atomic level—one token per heavy atom—enabling the same architecture to process arbitrary molecular graphs.
This hybrid scheme creates two parallel representation levels. Token-level tensors capture the coarse structure: the single representation s with dimensions (N_tokens × 384) and the pair representation z with dimensions (N_tokens × N_tokens × 128). Atom-level tensors handle fine-grained detail: q for per-atom features (N_atoms × 128) and p for atom-pair relationships (N_atoms × N_atoms × 16). The maximum input length is 5,000 tokens, enabling prediction of large complexes while maintaining computational tractability.
Input feature generation
The input pipeline begins with standard genetic database searches. Jackhmmer queries UniRef90 and MGnify for protein MSAs, HHBlits searches BFD and UniRef30, and nhmmer handles nucleotide databases including Rfam and RNAcentral for RNA sequences. Template structures are retrieved via hmmsearch against PDB70. For ligands, RDKit’s ETKDGv3 algorithm generates reference conformers from SMILES notation, providing initial three-dimensional coordinates.
The Input Embedder transforms these raw features into the initial representations. Atom-level features—including atomic number, charge, and reference conformer positions—are embedded and processed through three Atom Transformer blocks. These blocks use sparse attention patterns where each atom attends to 32 nearby atoms from a neighborhood of 128, managing computational cost while capturing local chemical environment. The atom-level representations are then aggregated to token-level through mean pooling over atoms belonging to each token.
The Pairformer: a simplified successor to Evoformer
The Pairformer represents a substantial architectural simplification over AlphaFold2’s Evoformer while maintaining the same depth of 48 blocks. The critical difference lies in what the module processes: while Evoformer maintained coupled MSA and pair representations throughout its depth, Pairformer operates exclusively on single (s) and pair (z) representations, having already distilled MSA information through a separate preprocessing module.
1. MSA Module preprocessing
Before the Pairformer, a dedicated MSA Module consisting of only 4 blocks extracts co-evolutionary signal. This module uses a simplified attention mechanism—pair-weighted averaging rather than full query-key attention—and applies the Outer Product Mean operation to transfer information from the MSA into the pair representation. Only the first row of the MSA representation (corresponding to the query sequence) propagates forward as the initial single representation.
This architectural choice reflects an empirical observation: MSA information, while valuable, can be efficiently compressed early in the pipeline. The dramatic reduction from 48 interlocked Evoformer blocks to 4 dedicated MSA blocks indicates that co-evolutionary signal, once captured, does not require continuous refinement alongside structural representations.
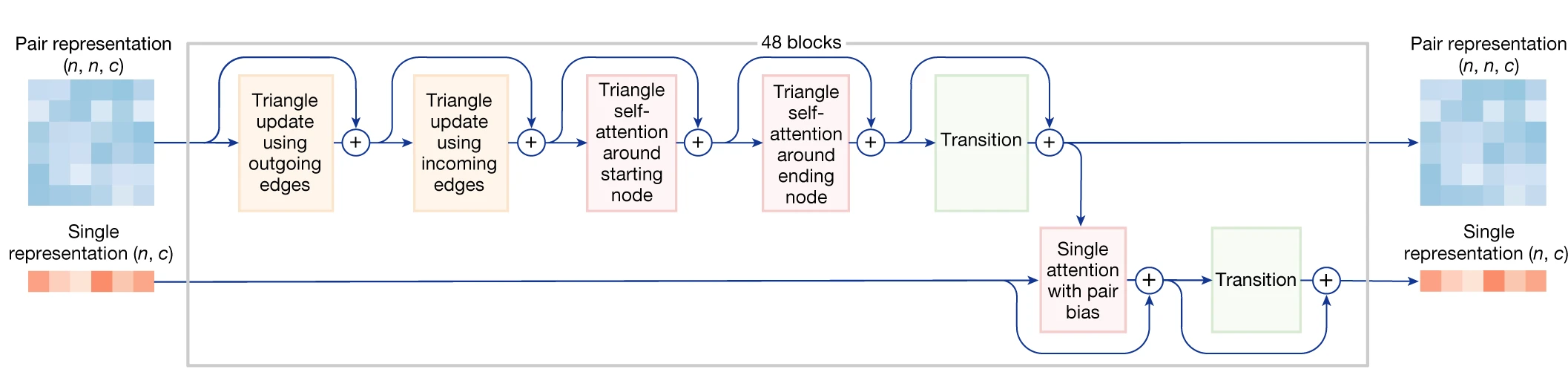
2. Pairformer block architecture
Each Pairformer block contains six sequential operations that update the pair and single representations:
Triangle multiplicative updates enforce geometric consistency by aggregating information along triangle edges. The outgoing update computes interactions between positions i and j by considering all intermediate positions k, effectively asking “what do positions i and j both know about every other position k?” The incoming update performs the complementary aggregation along columns. These operations enforce the triangle inequality principle: if residue A is close to B, and B is close to C, the A-C relationship is constrained.
Triangle attention augments standard self-attention with pair representation biases. Starting-node attention operates row-wise with attention scores biased by z_jk, while ending-node attention operates column-wise with bias from z_kj. The pair representation thus directly modulates which positions attend to each other.
Single attention with pair bias updates the single representation by attending across all tokens, with attention weights biased by the pair representation. This is the primary mechanism for information flow from pairwise relationships back to per-token features.
All transition layers use SwiGLU activations rather than AlphaFold2’s ReLU, following modern transformer best practices. Extensive gating mechanisms control information flow, inspired by LSTM architectures.
3. Key differences from Evoformer
| Component | Evoformer (AF2) | Pairformer (AF3) |
|---|---|---|
| MSA track in main trunk | Yes (coupled processing) | No (separate 4-block module) |
| Outer Product Mean | Every block | Only in MSA Module |
| Column-wise MSA attention | Yes | Not applicable |
| Activation function | ReLU | SwiGLU |
| Information flow | Bidirectional MSA↔pair | Primarily pair→single |
The simplification is significant: removing the MSA track from the main trunk eliminates the computationally expensive column-wise attention across evolutionary sequences, enabling the architecture to scale to non-protein molecules that lack evolutionary information entirely.
Diffusion-based structure generation replaces the Structure Module
The most dramatic architectural departure from AlphaFold2 is the replacement of the deterministic, equivariant Structure Module with a conditional diffusion model operating directly on raw atomic coordinates. This shift eliminates the need for residue frames, torsion angle parameterization, and explicit SE(3) equivariance—all of which fundamentally limited AlphaFold2 to protein-like molecules with well-defined backbone geometries.
1. The case against equivariance
AlphaFold2’s Invariant Point Attention (IPA) was engineered to be SE(3)-equivariant: rotations and translations of the input produced correspondingly rotated and translated outputs without affecting the underlying computation. This elegant constraint, however, came at architectural cost and restricted the representation to residue-specific frames.
AlphaFold3 takes a different approach: rather than building equivariance into the architecture, it learns rotational and translational invariance through data augmentation. During training, coordinates are randomly rotated and translated before each diffusion step. The network learns that structure quality is independent of global orientation, achieving the same invariance empirically rather than architecturally. This seemingly simple change enables direct coordinate prediction without the frame-based parameterization that limited generality.
2. The diffusion process
AlphaFold3 uses the EDM (Elucidating Diffusion Models) framework from Karras et al. 2022. Training samples noise levels σ directly from σ_data · exp(-1.2 + 1.5 · N(0,1)), while inference uses 200 predetermined denoising steps from high to low noise.
The diffusion process is inherently multi-scale. At high noise levels, only large-scale global structure matters—fine atomic details are overwhelmed by noise. As noise decreases, the model progressively refines local stereochemistry. This property elegantly addresses the multi-scale nature of structure prediction: global fold determination precedes local geometry optimization without explicit architectural separation.
3. Inference procedure
Structure generation proceeds as follows:
- Sample initial coordinates from a Gaussian distribution centered at the origin
- For each of 200 timesteps:
- Center coordinates at origin and apply random rotation (SE(3) augmentation)
- Add small random translation
- Perturb coordinates with noise appropriate to current timestep
- Pass through Diffusion Module to predict denoised coordinates
- Update coordinates using EDM update rule
- Return final atomic coordinates
The Diffusion Module itself uses a two-level attention architecture managing the token/atom duality. Atom-level processing uses sparse sequence-local attention (32 atoms attending to 128 neighbors), while token-level processing uses full self-attention. The architecture flows: atom attention → aggregate to tokens → token attention → broadcast to atoms → atom attention → predict coordinate updates.
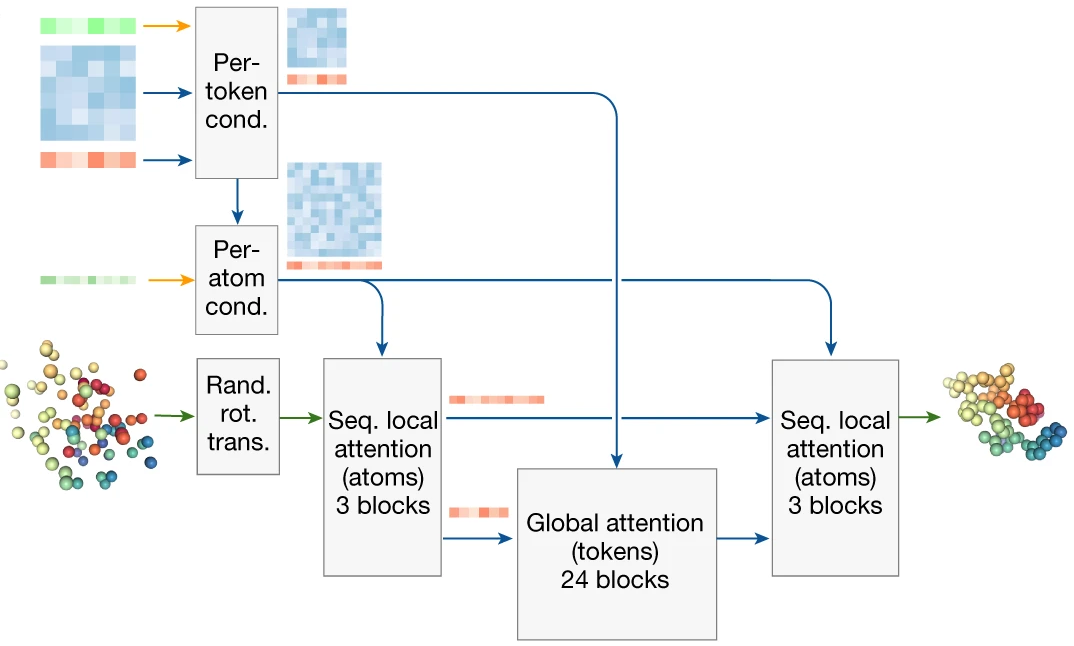
4. Conditioning the diffusion process
The Diffusion Module receives rich conditioning from the trunk:
- z_trunk: Pair representation from Pairformer (N × N × 128)
- s_trunk: Single representation from Pairformer (N × 384)
- s_inputs: Original input single representation
- Reference conformer: Initial coordinates from RDKit for ligands
- Fourier-embedded timestep: Current position in diffusion schedule
The timestep embedding is critical—it tells the network the current noise level, enabling appropriate behavior across the multi-scale denoising trajectory. Conditioning tensors are processed through transition blocks with adaptive layer normalization, where conditioning generates the normalization parameters dynamically.
Training methodology and the hallucination problem
Training AlphaFold3 required solving a novel challenge: diffusion models naturally “hallucinate” plausible-looking structures in regions of genuine disorder. Where AlphaFold2 produced characteristic extended “spaghetti” loops in low-confidence regions—a useful signal of uncertainty—early AlphaFold3 versions generated compact, plausible-looking but incorrect structures.
1. Cross-distillation for disorder handling
The solution is cross-distillation: training AlphaFold3 on structures predicted by AlphaFold-Multimer v2.3 in addition to experimental structures. Since AlphaFold2 produces extended conformations in disordered regions, AlphaFold3 learns to mimic this behavior, producing appropriate structural signatures of uncertainty rather than confident-looking hallucinations.
This cross-distillation applies only to protein chains—nucleic acids and ligands are removed from distillation structures since AlphaFold2 cannot process them. Structures with atomic clashes after re-adding removed molecules are excluded from training.
2. Loss functions and training efficiency
The primary loss is weighted aligned MSE on atom positions, with upweighting for DNA, RNA, and ligand atoms:
\[L*{\mathrm{diffusion}} = \mathrm{weight}(\sigma) \times (L*{\mathrm{MSE}} + \alpha*{\mathrm{bond}} \times L*{\mathrm{bond}}) + L\_{\mathrm{smooth \, LDDT}}\]The weighting factor (t² + σ_data²)/(t + σ_data)² balances loss contributions across noise levels. A smooth LDDT loss with sigmoid-based differentiable thresholding (at 4Å, 2Å, 1Å, 0.5Å) encourages accurate local geometry. Bond distance losses are added during fine-tuning stages.
Notably absent: explicit stereochemical violation penalties. The diffusion framework implicitly learns valid geometries through the multi-scale denoising process without requiring physics-based constraints in the loss function.
Training exploits the asymmetry between trunk and diffusion costs. The trunk (Pairformer) is computationally expensive; the Diffusion Module is relatively cheap. Therefore, each trunk forward pass generates 48 augmented diffusion samples (random rotations, translations, independent noise), all trained in parallel. This batch expansion enables efficient diffusion training without proportional trunk computation.
3. Training stages
Training proceeds through multiple stages with increasing crop sizes:
- Initial training: 384 tokens, 256 inputs × 48 diffusion samples = 12,288 samples per batch
- Fine-tuning stage 1: 640 tokens
- Fine-tuning stage 2: 768 tokens, 32 diffusion samples per input
Three cropping strategies ensure diverse training examples: contiguous (sequential residues), spatial (spatially nearby residues), and spatial interface (focused on interaction surfaces).
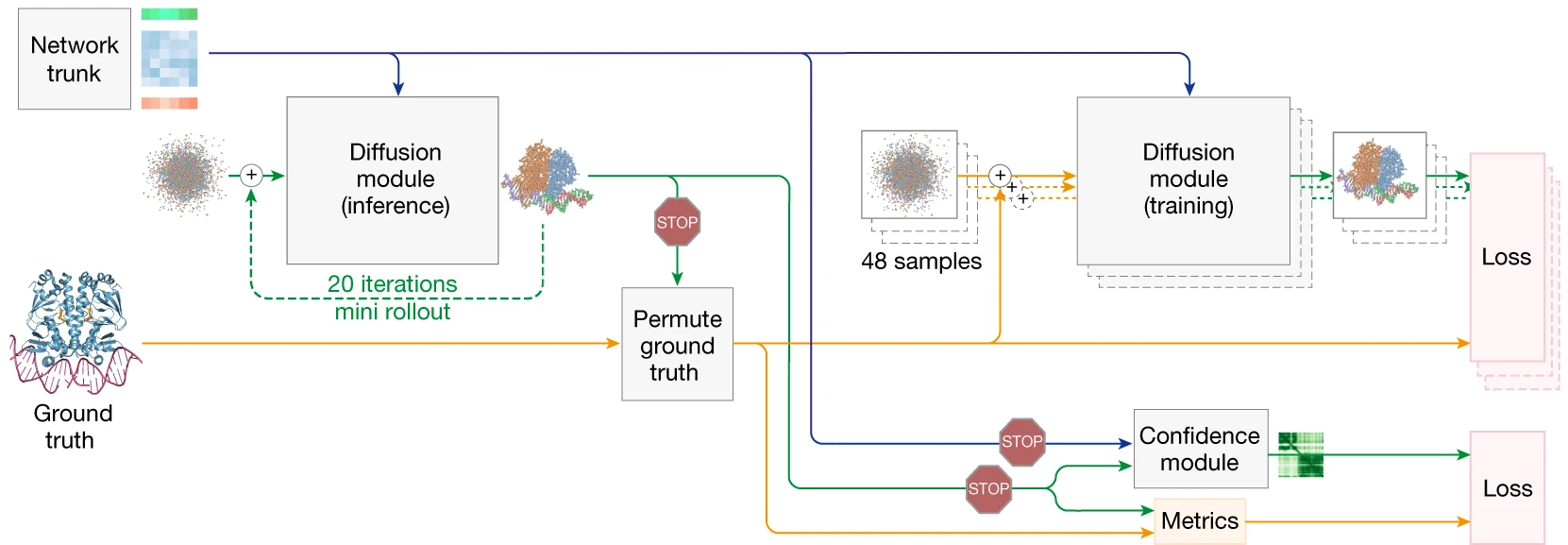
Confidence prediction mechanisms
AlphaFold3 produces multiple confidence metrics, refined through a specialized Confidence Module that processes diffusion-generated coordinates through a mini-Pairformer before prediction.
1. Confidence metrics
pLDDT (predicted Local Distance Difference Test) provides per-atom confidence scores from 0-100, indicating expected local structural accuracy. The confidence head receives single representations enriched with coordinate information from a diffusion “rollout” during training.
PAE (Predicted Aligned Error) estimates the positional error of residue j when the structure is aligned on residue i, stored as an asymmetric N×N matrix. This metric is particularly valuable for assessing domain relationships and interface quality.
pDE (Predicted Distance Error) is new to AlphaFold3, predicting absolute distance errors between atom pairs rather than alignment-dependent errors. The head outputs probabilities across 64 distance bins.
ipTM (interface predicted TM-score) aggregates pairwise confidence for cross-chain interfaces, serving as the primary ranking metric for multi-chain predictions.
2. Confidence training through mini-rollout
During training, the Diffusion Module performs a “mini-rollout” using larger step sizes to generate predicted coordinates quickly. These coordinates update the representations through a mini-Pairformer (2 blocks), and confidence heads predict from these enriched features. Crucially, gradients are detached—confidence training does not backpropagate through the main diffusion model, preventing the confidence head from biasing structure prediction.
Expanded capabilities beyond proteins
AlphaFold3’s unified architecture enables prediction across molecular types that were completely inaccessible to AlphaFold2:
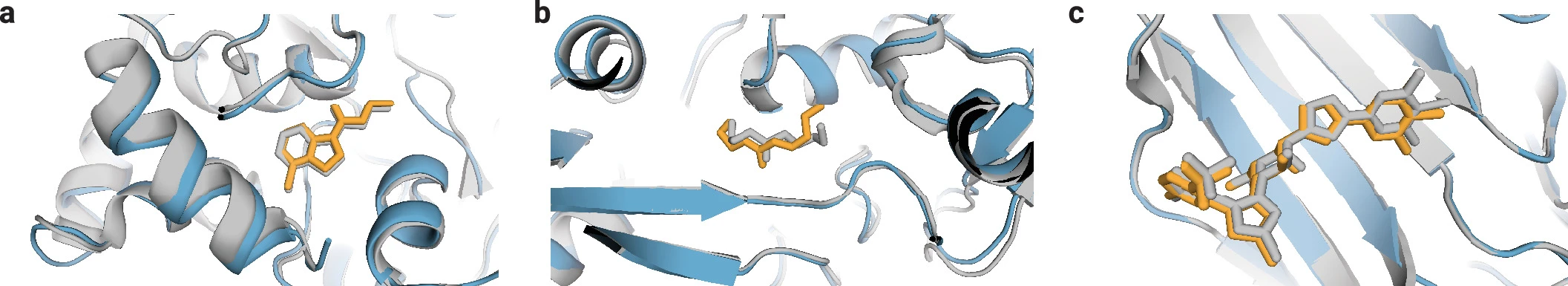
1. Protein-ligand complexes
Small molecules are represented atom-by-atom with reference conformers from RDKit. On the PoseBusters benchmark, AlphaFold3 achieves 76.4% success rate (RMSD < 2Å and passing validity checks)—approximately 50% improvement over the best traditional docking methods operating without protein structure information. This makes AlphaFold3 the first AI system to surpass physics-based tools for biomolecular structure prediction.
Limitations remain: chirality violations occur in approximately 4.4% of ligand predictions, and accuracy degrades for ligands requiring large protein conformational changes (>5Å backbone RMSD).
2. Protein-nucleic acid complexes
DNA and RNA chains use nucleotide-level tokenization with MSA support for RNA sequences. AlphaFold3 significantly outperforms RoseTTAFold2NA on both protein-DNA (p = 5.2×10⁻¹²) and protein-RNA (p = 1.6×10⁻⁷) benchmarks. Large RNA structures may require generating hundreds to thousands of samples to obtain clash-free models.
3. Post-translational modifications and covalent ligands
Modifications are specified via Chemical Component Dictionary (CCD) codes at defined positions. Performance varies by modification type:
| Modification type | Success rate |
|---|---|
| Covalent ligands | ~79% |
| Modified protein residues | ~51% |
| Single glycans | ~46% |
| Multi-residue glycans | ~42% |
| Modified RNA residues | ~41% |
4. Ions and cofactors
Metal ions and common cofactors (ATP, ADP, NAD, FAD, heme, etc.) are supported via CCD codes, enabling prediction of catalytic sites and metal coordination geometries.

Architectural specifications at a glance
| Component | Specification |
|---|---|
| Pairformer blocks | 48 |
| MSA Module blocks | 4 |
| Template Embedder blocks | 2 (mini-Pairformer) |
| Input Embedder (Atom Transformer) blocks | 3 |
| Single representation channels (c_s) | 384 |
| Pair representation channels (c_z) | 128 |
| MSA representation channels (c_m) | 64 |
| Atom-level single channels (c_atom) | 128 |
| Atom-level pair channels (c_atompair) | 16 |
| Maximum tokens | 5,000 |
| Diffusion steps (inference) | 200 |
| Recycling iterations | 4 |
| Default samples | 25 (5 seeds × 5 diffusion samples) |
Inference time scales with complexity: approximately 22 seconds for a 1,024-token complex on 16 A100 GPUs, extending to over 5 minutes for 5,120-token complexes.
Conclusion: architectural simplification enables generalization
AlphaFold3’s most surprising lesson may be that removing complexity—equivariance constraints, MSA coupling, frame-based representations—enables greater capability. The Pairformer’s streamlined design, the diffusion module’s coordinate-level generativity, and the unified token representation collectively create an architecture that handles biomolecular diversity without molecular-specific engineering.
Three innovations stand out as particularly consequential. First, replacing IPA with diffusion-based generation eliminates the architectural constraints that limited AlphaFold2 to proteins, enabling direct all-atom coordinate prediction for arbitrary molecules. Second, cross-distillation solves the hallucination problem inherent to generative models, preserving meaningful confidence signals in disordered regions. Third, the token-based representation provides a natural abstraction boundary—one token per residue for polymers, one token per atom for small molecules—enabling efficient computation at appropriate granularity.
The practical impact is substantial: 50% improvement over docking methods for protein-ligand prediction, significant gains for protein-nucleic acid complexes, and entirely new capabilities for modified residues and covalent interactions. For computational structural biologists, AlphaFold3 represents not just an accuracy improvement but a paradigm shift toward unified biomolecular modeling within a single learned architecture.
References
Abramson, J., Adler, J., Dunger, J., Evans, R., Green, T., Pritzel, A., Ronneberger, O., Willmore, L., et al. “Accurate structure prediction of biomolecular interactions with AlphaFold 3.” Nature 630, 493–500 (2024). https://doi.org/10.1038/s41586-024-07487-w
Jumper, J., Evans, R., Pritzel, A., et al. “Highly accurate protein structure prediction with AlphaFold.” Nature 596, 583–589 (2021).
Karras, T., Aittala, M., Aila, T., & Laine, S. “Elucidating the Design Space of Diffusion-Based Generative Models.” NeurIPS (2022).
The Illustrated AlphaFold by Elana Simon and Jake Silber for an ML audience.
Enjoy Reading This Article?
Here are some more articles you might like to read next: