When Transformers learn to speak protein
a deep dive into BERTology meets biology
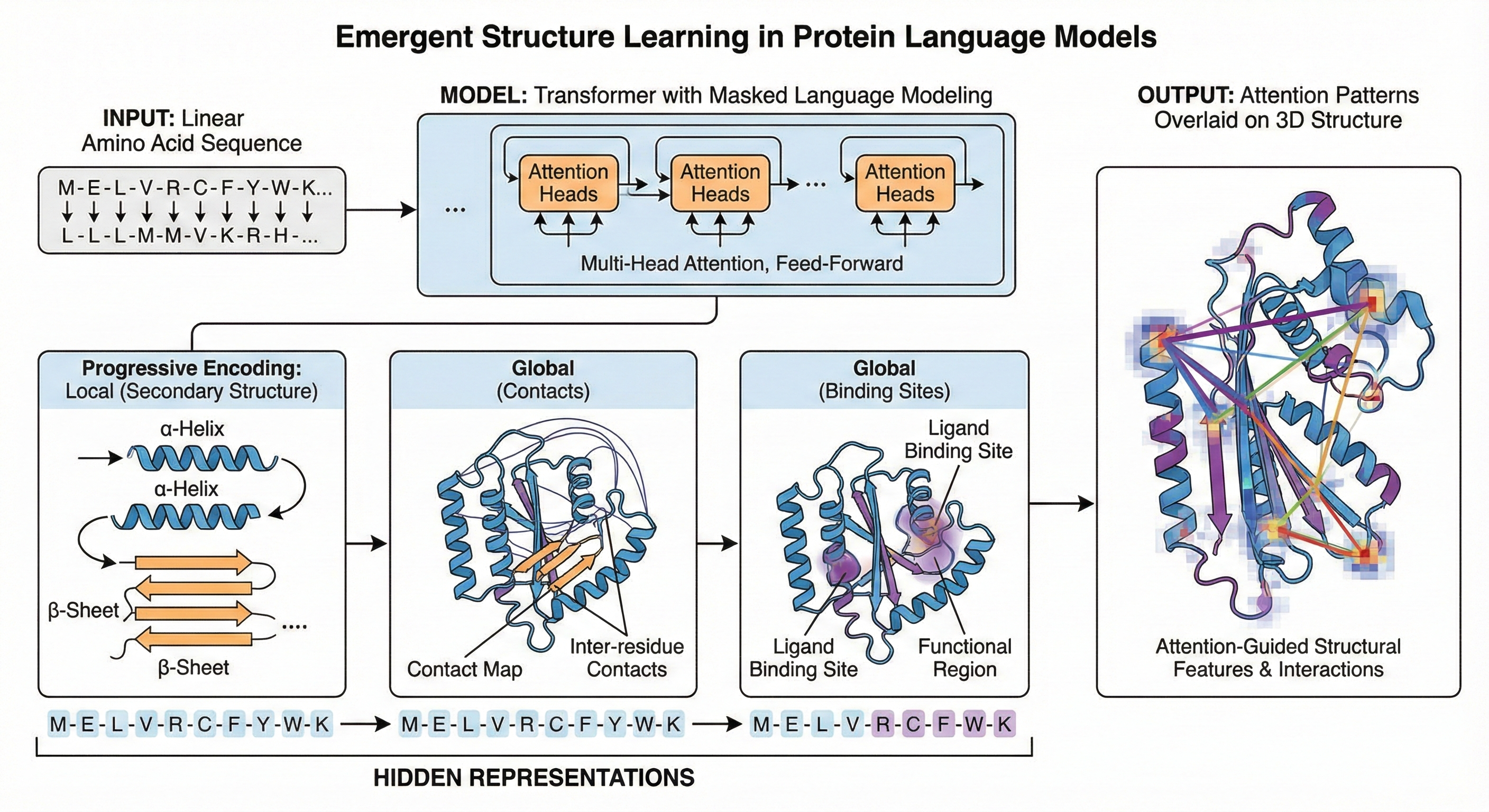
Protein language models have learned something remarkable: they can “see” 3D structure without ever being shown it. Vig et al.’s influential 2021 paper demonstrates that Transformer attention mechanisms, trained solely through masked language modeling on amino acid sequences, spontaneously develop awareness of protein folding patterns, binding sites, and functional regions. This finding bridges two rapidly evolving fields—Natural Language Processing (NLP) interpretability and computational biology—while revealing that the statistical patterns encoded in evolution contain sufficient information for neural networks to recover fundamental principles of protein biochemistry.
The implications extend beyond academic curiosity. Understanding what these models learn, and how they learn it, illuminates both the promise and limitations of AI-driven protein engineering. This note dissects the methods, mathematics, and findings that established attention analysis as a tool for interpreting biological sequence models.
The fundamental insight: proteins as language
Proteins are nature’s molecular machines—long chains of amino acids that fold into precise 3D structures to catalyze reactions, transmit signals, and build cellular architecture. For decades, computational biologists sought to predict protein structure from sequence, a challenge so difficult it spawned the CASP competition and eventually AlphaFold’s breakthrough. But before AlphaFold, researchers asked a different question: could unsupervised learning on sequences alone capture meaningful biological information?
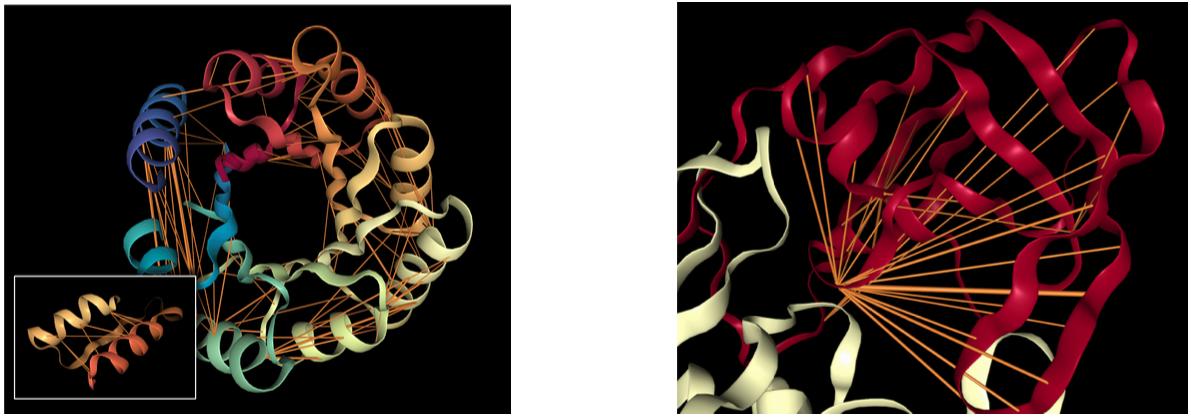
Figure 1: Attention mechanisms in protein Transformers capture structural and functional properties without explicit supervision. (a) Head 12-4 in TapeBert identifies long-range contacts in a de novo designed TIM-barrel protein (PDB: 5BVL). Orange lines represent attention weights (width proportional to magnitude, values $\lt 0.1$ hidden). Note how residues 117D and 157I receive strong mutual attention despite being 40 positions apart in sequence but spatially adjacent in the characteristic β-α-β barrel architecture. (b) Head 7-1 targets binding sites in HIV-1 protease (PDB: 7HVP), with primary attention focused on position 27G—the precise binding pocket for protease inhibitor drugs. This head was selected based on correlation with experimentally annotated binding sites, yet the model learned this pattern purely from masked language modeling without functional annotations. Visualizations created using NGL Viewer. Source: Vig et al. 2022, Figure 1.
The answer came from an unexpected direction. When BERT and other Transformers revolutionized NLP in 2018-2019, researchers recognized that protein sequences—strings of 20 amino acid “letters” rather than words—could be modeled identically. The Tasks Assessing Protein Embeddings (TAPE) benchmark (Rao et al., 2019) demonstrated that Transformers pretrained on the Pfam database of 31 million protein domains learned representations useful for downstream tasks like secondary structure prediction and fluorescence prediction. ProtTrans (Elnaggar et al., 2021) scaled this approach dramatically, training models on up to 2.1 billion sequences from the BFD database.
What is the TAPE benchmark?
The Tasks Assessing Protein Embeddings (TAPE) is a standardized benchmark introduced by Rao et al. (2019) to evaluate how well machine learning models learn protein representations (embeddings) from sequence data. It was designed to provide a “GLUE-like” evaluation suite for the protein domain, addressing the lack of standardized metrics in the field.
The benchmark consists of five biologically relevant tasks categorized into three major areas:
1. Structure Prediction:
- Secondary Structure Prediction: Predicts whether each amino acid residue in a sequence belongs to a helix, sheet, or coil.
- Contact Prediction: Predicts which pairs of amino acids are spatially close (within 8 Å) in the protein's 3D folded structure.
2. Evolutionary Understanding:
- Remote Homology Detection: A classification task to identify the structural "fold" of a protein from its sequence, even when it has very low similarity to known proteins.
3. Protein Engineering:
- Fluorescence Prediction: A regression task to predict the log-fluorescence intensity of Green Fluorescent Protein (GFP) variants. It tests a model's ability to generalize to unseen combinations of mutations.
- Stability Prediction: Measures a model's ability to predict the biochemical stability of a protein sequence.
Key Features
- Pretraining Dataset: TAPE provides a large corpus of unlabeled protein sequences (from Pfam) for self-supervised pretraining.
- Biologically Relevant Splits: Unlike random splits, TAPE uses curated training and test sets (e.g., holding out entire protein families) to ensure models learn true biological generalization rather than just memorizing similar sequences.
- Standardized Baseline: The paper benchmarked several architectures, including Transformers, LSTMs, and ResNets, finding that self-supervised pretraining significantly improves performance across most tasks.
But what exactly do these models learn? The “Bertology Meets Biology” paper tackles this question by adapting the rich toolkit of NLP interpretability—probing classifiers, attention analysis, layer-wise studies—to protein sequences. The authors analyze five pretrained models spanning three architectures (BERT, ALBERT, XLNet) and discover that attention patterns correlate strongly with known structural and functional properties of proteins.
The models under investigation
The paper examines five Transformer models that represent the diversity of protein language model architectures circa 2020:
| Model | Architecture | Layers × Heads | Parameters | Training Data |
|---|---|---|---|---|
| TapeBert | BERT | 12 × 12 | 94M | Pfam (31M sequences) |
| ProtBert | BERT | 30 × 16 | 420M | UniRef100 (216M sequences) |
| ProtBert-BFD | BERT | 30 × 16 | 420M | BFD (2.1B sequences) |
| ProtAlbert | ALBERT | 12 × 64 | 224M | UniRef100 |
| ProtXLNet | XLNet | 30 × 16 | 409M | UniRef100 |
These models differ in both architecture and training data. BERT uses bidirectional masked language modeling, predicting randomly masked amino acids from surrounding context. ALBERT shares parameters across layers, enabling more attention heads (64 vs. 12-16) with fewer total parameters. XLNet employs permutation language modeling, a bidirectional autoregressive approach that considers all possible orderings of the input factorization.
What is bidirectional masked language modeling?
Bidirectional Masked Language Modeling (MLM) is a self-supervised pretraining technique where a model learns to predict hidden or “masked” tokens in a sequence by analyzing the entire context—both before and after the missing element.
This approach differs from traditional unidirectional models, which only process text sequentially (e.g., left-to-right), and is a core component of architectures like BERT.
How It Works
1. Masking: A random subset of tokens (typically 15%) in an input sequence is replaced with a special [MASK] token.
2. Bidirectional Processing: The model uses self-attention mechanisms to weigh every word in the sentence simultaneously, rather than processing them in a fixed order.
3. Reconstruction: The model attempts to predict the original identity of the masked tokens based on the surrounding “unmasked” context.
Key Advantages
-
Deep Contextual Understanding: By seeing “future” and “past” words at once, the model captures nuanced meanings and resolves ambiguities that unidirectional models might miss.
-
Contextual Embeddings: It generates word representations (embeddings) that vary based on the sentence they appear in (e.g., distinguishing “bank” in “river bank” vs. “bank account”).
-
Superior NLU Performance: In 2026, it remains the preeminent paradigm for pretraining models used in Natural Language Understanding (NLU) tasks like sentiment analysis, named entity recognition, and question answering.
Modern Applications
-
Multimodal Pretraining: High masking rates (over 60%) are now used in vision-language tasks to facilitate cross-modal alignment between images and text.
-
Tabular Data Synthesis: MLM is repurposed to reconstruct missing values in tabular datasets, enabling privacy-adjustable synthetic data generation.
-
Protein Modeling: As seen in benchmarks like TAPE, MLM is used to learn the “language” of amino acid sequences to predict protein stability and structure [Previous Turns].
-
Privacy-Preserving Training: “Privacy-by-design” MLM excludes sensitive identifiers from masking to prevent the model from memorizing and regurgitating private user data.
The largest model by dataset size—ProtBert-BFD—was trained on the Big Fantastic Database, which combines UniProt with metagenomic sequences to achieve an unprecedented 393 billion amino acid tokens, roughly 112 times larger than English Wikipedia. This scale proves consequential: ProtBert-BFD consistently shows the strongest alignment between attention and structural properties.
Methodology: how to analyze attention in protein Transformers
The attention alignment metric
The core analytical tool measures how attention aligns with known biological properties. Given a property $f$ defined for token pairs (e.g., whether amino acids $i$ and $j$ are in contact), the paper computes the proportion of high-confidence attention weights that connect tokens where the property holds:
\[p_\alpha(f) = \frac{\sum_{x \in X} \sum_{i=1}^{|x|} \sum_{j=1}^{|x|} f(i,j) \cdot \mathbb{1}_{\alpha_{i,j} > \theta}}{\sum_{x \in X} \sum_{i=1}^{|x|} \sum_{j=1}^{|x|} \mathbb{1}_{\alpha_{i,j} > \theta}}\]Here $\alpha_{i,j}$ denotes the attention weight from position $i$ to position $j$, $\theta = 0.3$ is a threshold selecting high-confidence attention, and $X$ is a dataset of protein sequences. The indicator function $f(i,j)$ returns 1 if the property is present (e.g., if residues $i$ and $j$ are within 8 Ångströms in the folded structure) and 0 otherwise.
For properties of individual tokens rather than pairs—such as binding sites—$f(i,j)$ returns 1 if token $j$ possesses the property, making $p_\alpha(f)$ the proportion of attention directed toward that property.
The paper also presents an alternative continuous formulation using attention-weighted averages:
\[p_\alpha(f) = \frac{\sum_{x \in X} \sum_{i=1}^{|x|} \sum_{j=1}^{|x|} f(i,j) \cdot \alpha_{i,j}(x)}{\sum_{x \in X} \sum_{i=1}^{|x|} \sum_{j=1}^{|x|} \alpha_{i,j}(x)}\]This formulation considers all attention weights rather than thresholding, providing a complementary view of attention-property alignment.
The BERT attention mechanism
Understanding these metrics requires revisiting how Transformers compute attention (Vaswani et al. 2017, The Illustrated Transformer, and The Illustrated Bert). BERT applies multi-head self-attention, where each head learns distinct query ($Q$), key ($K$), and value ($V$) matrices:
\[\text{Attention}(Q, K, V) = \text{softmax}\left(\frac{QK^T}{\sqrt{d_k}}\right)V\]The attention weights $\alpha_{i,j}$ emerge from the softmax over scaled dot-products of query and key vectors. In a multi-head setting with $h$ heads, queries, keys, and values are linearly projected $h$ times, attention is computed in parallel for each projection, and results are concatenated. TapeBert has 12 layers with 12 heads each, yielding 144 distinct attention mechanisms to analyze.
Statistical validation through dual checks
To ensure findings aren’t artifacts of chance, the authors implement two validation approaches:
Bonferroni-corrected significance testing: For each property, a two-proportion z-test compares (1) the proportion of high-confidence attention arcs where $f(i,j) = 1$ to (2) the background frequency of the property in the dataset. With $m$ attention heads constituting multiple hypotheses, the significance threshold is adjusted to $\alpha/m$, typically yielding thresholds around $p < 0.00001$ for the 144-head TapeBert model.
Null model comparison: Initial attempts to use randomly initialized models or models trained on shuffled sequences produced no attention weights exceeding the threshold $\theta$—evidence that high-confidence attention itself is meaningful. Instead, the authors implement a post-hoc randomization: attention weights from each position are randomly permuted while preserving their sum-to-one constraint. This null model reveals whether specific attention patterns, not just the existence of strong attention, drive the observed correlations.
Probing classifiers: embeddings vs. attention
Beyond attention analysis, the paper employs probing classifiers to quantify knowledge encoded in model representations. These diagnostic classifiers—trained on frozen model outputs to predict biological properties—reveal what information is accessible at each layer.
Embedding probes use layer outputs directly. For token-level properties (binding sites, secondary structure), each token’s embedding feeds into the classifier. For pairwise properties (contacts), the probe receives concatenated elementwise differences and products of the two tokens’ embeddings.
Attention probes treat attention weights as features. For contact prediction, attention weights across all heads in a layer form a feature vector for each token pair, and a classifier predicts whether those residues are in contact.
Evaluation metrics are task-specific: F1 score for secondary structure, precision@L/5 for contacts (where L is sequence length), and precision@L/20 for binding sites (reflecting their ~4.8% frequency).
Results: attention captures folding structure
Contact maps emerge in deep layers
The paper’s most striking finding concerns contact maps—binary matrices indicating which amino acid pairs are spatially close (<8Å apart) in the folded 3D structure but distant (>6 positions apart) in the sequence. These “long-range contacts” are precisely what makes protein folding difficult: predicting them requires understanding how distant sequence elements interact.
Attention aligns remarkably well with contact maps, particularly in deep layers. The most contact-aligned heads achieve:
- TapeBert: 44.7% of attention on contacts (head 12-4)
- ProtAlbert: 55.7% (head 11-3)
- ProtBert: 58.5% (head 30-14)
- ProtBert-BFD: 63.2% (head 30-9)
- ProtXLNet: 44.5% (head 26-6)
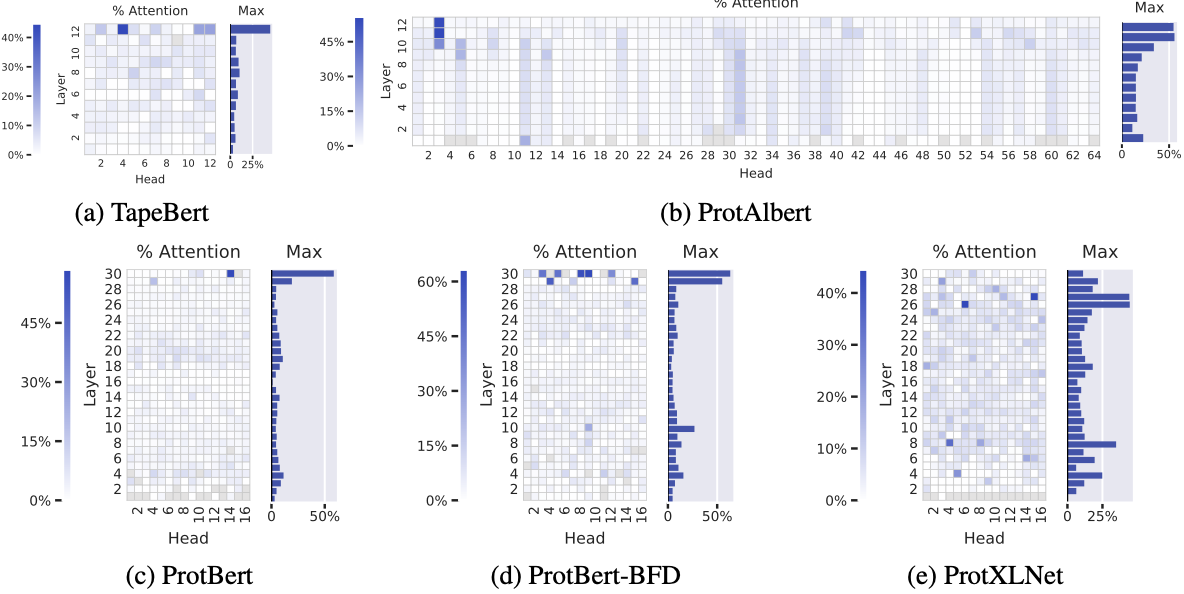
Figure 2: Contact map alignment across five pretrained Transformer models reveals consistent emergence of structure-aware attention in deep layers. Each heatmap shows the proportion of high-confidence attention weights ($α_{i,j} > 0.3$) from each head that connects amino acid pairs in contact ($\lt \text{8Å}$ apart in 3D structure but >6 positions apart in sequence). Background contact frequency is only 1.3%. (a) TapeBert (12 layers, 12 heads): maximum 44.7% in head 12-4. (b) ProtAlbert (12 layers, 64 heads): maximum 55.7% in head 11-3. Vertical striping pattern reflects ALBERT's cross-layer parameter sharing. (c) ProtBert (30 layers, 16 heads): maximum 58.5% in head 30-14. (d) ProtBert-BFD (30 layers, 16 heads): maximum 63.2% in head 30-9—the highest observed value, achieved by the largest training dataset (2.1B sequences). (e) ProtXLNet (30 layers, 16 heads): maximum 44.5% in head 26-6. The consistent concentration in final layers across all architectures suggests hierarchical representation learning where structural understanding emerges after lower-level features are established. Grayed cells indicate heads with $\lt 100$ high-confidence attention arcs in the dataset. Source: Vig et al. 2022, Figure 2.
These figures become extraordinary when compared to the background frequency of contacts at just 1.3%—a 50-fold enrichment in the best heads. The pattern is consistent: contact-aware attention concentrates in the deepest layers (Figure 2 in the paper), suggesting the models build hierarchical representations where structural understanding emerges late in processing.
ProtBert-BFD, trained on the largest dataset with 2.1 billion sequences, shows the strongest contact alignment. This supports the hypothesis that both model capacity (420M parameters, matching ProtBert) and training data scale contribute to learning structurally-informed representations.
Why does attention learn contacts?
The authors offer a compelling explanation rooted in coevolution. Amino acids in physical contact exert evolutionary pressure on each other—if one mutates, the other must often compensate to maintain the structural interaction. This creates statistical dependencies between contacting residue positions across evolutionary history.
When training with masked language modeling, a model predicting masked amino acids benefits from attending to positions that provide predictive information about the mask. If position $j$ tends to coevolve with position $i$—because they’re in contact—then attending to $j$ helps predict what amino acid occupies $i$. The language modeling objective thus naturally encourages attention patterns that reflect contact structure.
This explanation aligns with the broader principle that structure is encoded in evolutionary statistics. The success of coevolution-based methods like Direct Coupling Analysis (DCA) for contact prediction demonstrates that sequence alignments contain rich structural information. Transformers, with their ability to model long-range pairwise interactions through attention, appear to learn similar patterns from raw sequences.
What is Direct Coupling Analysis (DCA)?
Direct Coupling Analysis (DCA) is a global statistical framework used to predict spatial residue-residue contacts in proteins and RNA from sequence information alone. It is primarily a method for identifying which residues are physically close in a 3D folded structure by analyzing co-evolutionary signals in a Multiple Sequence Alignment (MSA).
Core Principle: Disentangling Direct vs. Indirect Correlations
Traditional methods like Mutual Information (MI) often fail because they cannot distinguish between direct interactions and indirect correlations.
-
Direct Interaction: Residues A and B are in physical contact; a mutation in A requires a compensatory mutation in B to maintain stability.
-
Indirect Correlation: If residue A interacts with B, and B interacts with C, residues A and C will appear correlated even if they are far apart in 3D space. DCA uses a global statistical model (typically a Potts model or generalized Ising model) to “disentangle” these signals, isolating only the direct couplings.
Common Implementation Methods
Several algorithmic strategies are used to solve the underlying inverse statistical problem, varying in speed and accuracy:
-
Mean-Field DCA (mfDCA): A fast approximation that is $10^{3}$ to $10^{4}$ times faster than earlier methods, making it suitable for high-throughput analysis.
-
Pseudo-likelihood Maximization (plmDCA): Generally more accurate than mfDCA and widely used as a standard benchmark in contact prediction software.
-
Boltzmann Learning: An exact but computationally expensive stochastic procedure that can outperform other methods in specific RNA and protein applications.
Applications
-
Monomeric Fold Prediction: Identifying the 3D structure of a single protein chain.
-
Protein-Protein Interactions (PPI): Predicting “interprotein” contacts that stabilize complex formation.
-
Alternative Conformations: Identifying signals for multiple functional states or alternative binding interfaces.
-
RNA Structure Prediction: Adapting the model to identify base-pair and tertiary interactions in RNA families.
Key Metrics and Outputs
-
Direct Information (DI) Score: A metric used to rank the strength of predicted couplings; higher scores indicate a greater likelihood of a spatial contact.
-
Contact Map: A 2D grid representing the predicted pairs, where clusters of top-scoring pairs often correspond to secondary structure elements (e.g., helices packing) or distant tertiary contacts.
Attention targets functional sites
Binding sites receive concentrated attention
Beyond structure, the paper examines whether attention captures functional properties—specifically binding sites, the regions where proteins interact with other molecules to perform their biological roles.
ProtAlbert shows particularly strong binding site attention, with 22 heads focusing >50% of attention on binding sites despite binding sites comprising only 4.8% of sequence positions. The three BERT models achieve top-head values of 48.2%, 50.7%, and 45.6% attention on binding sites.
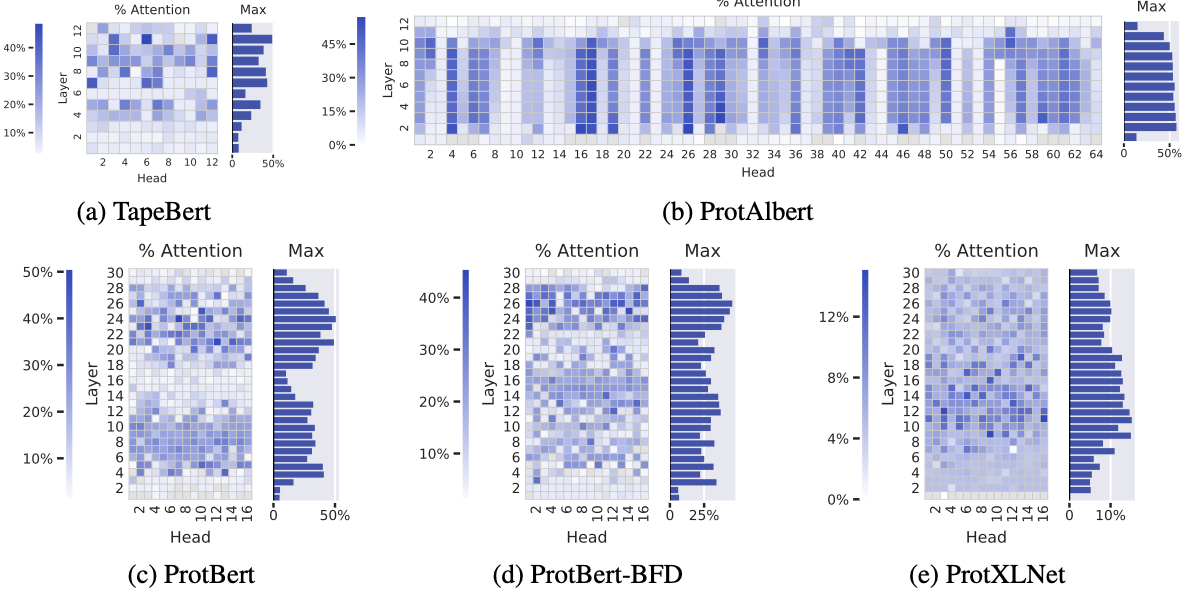
Figure 3: Binding site attention across five models demonstrates functional awareness learned from sequence statistics alone. Each heatmap shows the proportion of high-confidence attention directed toward experimentally annotated binding sites (background frequency: 4.8%).
(a) TapeBert: maximum 48.2% in head 11-6.
(b) ProtAlbert: exceptional performance with 22 heads exceeding 50% attention on binding sites (maximum 49% in multiple heads).
(c) ProtBert: maximum 50.7% in head 24-7.
(d) ProtBert-BFD: maximum 45.6% in head 26-1.
(e) ProtXLNet: notably weaker binding site attention (maximum 15.1% in head 11-16, average 6.2%) compared to masked language models, possibly reflecting differences in bidirectional autoregressive pretraining.
Unlike contact map attention which concentrates in final layers, binding site attention appears throughout middle and deep layers, suggesting these functional hotspots serve as anchoring points for contextual understanding across multiple stages of representation building. Source: Vig et al. 2022, Figure 3.
Figure 1b of the paper visualizes this dramatically: attention head 7-1 in TapeBert highlights position 27G in HIV-1 protease, a known binding site for protease inhibitor drugs. The model, trained only to predict masked amino acids, learned to “see” the functional hotspot that pharmaceutical chemists target.
Interestingly, ProtXLNet shows weaker binding site attention (maximum 15.1%, average 6.2%) compared to the masked language models. The authors speculate this may reflect differences in pretraining objectives—XLNet’s permutation language modeling may not create the same incentives to attend to conserved functional regions.
The evolutionary basis for binding site attention
Why would language modeling encourage attention to binding sites? The paper offers an evolutionary explanation: binding sites remain conserved even as surrounding sequences evolve, because mutations in these functional regions would disrupt protein activity and be selected against. This conservation creates a distinctive statistical signature—binding sites have lower sequence entropy and stronger coevolutionary couplings with the residues they contact.
By attending to these conserved regions, models gain access to a “high-level characterization of the protein that is robust to individual sequence variation.” Binding sites effectively anchor the protein’s identity, providing predictive context for masked positions throughout the sequence.
Post-translational modifications
A smaller number of heads concentrate attention on post-translational modification (PTM) sites—positions where proteins are chemically modified after translation. Head 11-6 in TapeBert focuses 64% of attention on PTM positions, though these occur at only 0.8% of sequence positions.
PTMs like phosphorylation and glycosylation critically regulate protein function, and their sites have characteristic sequence motifs that the model apparently learns to recognize. This attention pattern suggests potential applications in predicting modification sites for newly discovered proteins.
Layer-wise analysis reveals hierarchical learning
From simple to complex across depth
One of the paper’s most elegant findings concerns how different layers encode different complexity levels. Figure 4 shows attention to various properties averaged across heads within each layer of TapeBert:
- Secondary structure (helix, strand, turn/bend): attention distributed relatively evenly across layers
- Binding sites: attention increases in deeper layers
- Contacts: attention strongly concentrated in the deepest layers
This pattern—simple, local properties encoded early; complex, global properties emerging late—mirrors findings from NLP, where BERT’s layers progressively encode part-of-speech tagging, syntax, and semantics (Tenney et al., 2019).
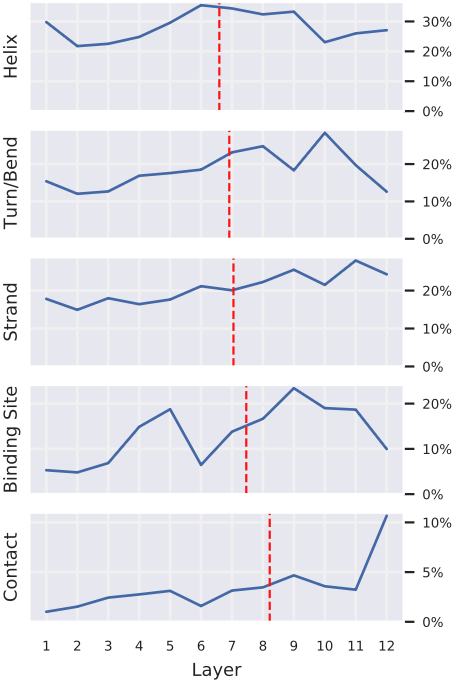
Figure 4: Attention to different protein properties exhibits distinct layer-wise progression, revealing hierarchical concept learning. Each subplot shows the percentage of attention (averaged over all heads within each layer) focused on a given property in TapeBert. Properties are sorted by "center of gravity" (red dashed lines)—the attention-weighted average layer depth.
- Helix, Turn/Bend, Strand (secondary structure): relatively flat distributions across layers, with slight early-layer emphasis for these local structural motifs.
- Binding sites: progressive increase in deeper layers (center of gravity: layer ~7).
- Contacts: sharp concentration in final layers (center of gravity: layer ~10).
This gradient from local to global mirrors findings in NLP where BERT progressively encodes part-of-speech → syntax → semantics. The deepest layers integrate information across the entire sequence to identify long-range structural relationships, while earlier layers capture local patterns that provide building blocks for higher-level representations. Source: Vig et al. 2022, Figure 4.
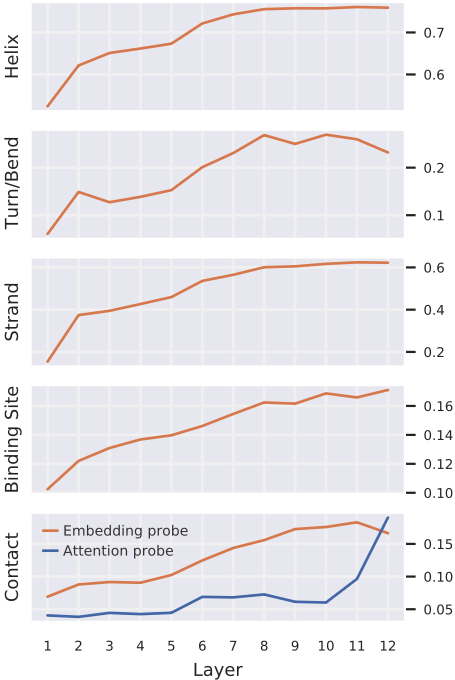
Figure 5: Embedding probes versus attention probes reveal different timescales for knowledge encoding versus knowledge deployment. Each subplot shows probing classifier performance by layer for a given property (same vertical ordering as Figure 4).
Embedding probes (orange): Linear classifiers trained on layer output embeddings to predict properties. Performance increases gradually across layers for all properties, with secondary structure knowledge building quickly in early layers and binding site/contact knowledge accumulating more slowly.
Attention probe (blue, contacts only): Classifier using attention weights as features for pairwise contact prediction. Performance remains near-baseline through layer 11, then jumps sharply in layer 12.
This divergence is methodologically significant: embeddings encode contact knowledge progressively across many layers (accumulated through feedforward transformations and residual connections), but attention only operationalizes this knowledge in the final layers. Standard embedding-only probing would miss this distinction between "what the model knows" versus "how it deploys that knowledge."
Evaluation metrics: F1 score (secondary structure), precision@L/5 (contacts), precision@L/20 (binding sites). Source: Vig et al. 2022, Figure 5.
Embedding probes vs. attention probes tell different stories
The probing analysis (Figure 5) reveals a fascinating distinction between what embeddings encode versus what attention operationalizes:
Embedding probes show gradual accumulation of contact information across layers. Secondary structure knowledge builds quickly in early layers; contact and binding site knowledge increases steadily but more slowly.
Attention probes for contacts show a dramatically different pattern: accuracy remains low through layers 1-11, then jumps sharply in layer 12. Knowledge of contacts appears in attention weights “only in the final layers.”
This divergence is methodologically significant. As the authors note, embedding probes reveal “what the model knows” while attention analysis reveals “how it operationalizes that knowledge.” A model might build up structural understanding gradually through layer computations, but only deploy that understanding through attention at the final layers. Standard layer-wise probing of embeddings alone would miss this distinction.
Amino acid attention aligns with evolutionary substitution
Specialized attention heads for specific amino acids
The paper’s fine-grained analysis reveals that individual attention heads specialize in particular amino acids. For 16 of the 20 standard amino acids in TapeBert, at least one head focuses >25% of its attention on that amino acid—far exceeding background frequencies ranging from 1.3% to 9.4%.
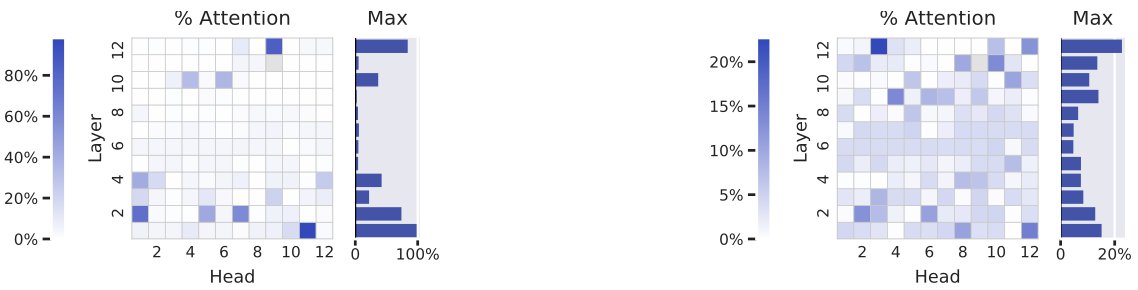
Figure 6: Individual attention heads specialize in specific amino acids, with some achieving near-complete specialization. Heatmaps show the percentage of each head's attention focused on proline (left) and phenylalanine (right) in TapeBert.
- Proline (left): Head 1-11 directs 98.3% of its attention to proline residues, despite proline's 4.6% background frequency. Proline is structurally unique—its cyclic side chain restricts backbone angles, often creating kinks and turns. This extreme specialization suggests the head has learned proline's distinctive structural role.
- Phenylalanine (right): Head 12-3 focuses 22.7% of attention on this aromatic amino acid (background: 3.9%). Multiple heads show moderate phenylalanine preference, consistent with its role in hydrophobic core packing and aromatic interactions.
Across all 20 amino acids in TapeBert, 16 have at least one head directing >25% attention to that residue—far exceeding chance. Similar patterns emerge in ProtBert (17/20), ProtBert-BFD (15/20), ProtAlbert (16/20), and ProtXLNet (18/20). Source: Vig et al. 2022, Figure 6.
Figure 6 illustrates this vividly for proline (Pro) and phenylalanine (Phe). Head 1-11 directs 98.3% of attention to proline, while head 12-3 focuses 22.7% on phenylalanine. Similar patterns emerge across all five models, with 15-18 amino acids receiving >25% attention from at least one head.
Correlation with BLOSUM62 substitution matrix
But has the model simply memorized amino acid identities, or has it learned meaningful biochemical relationships? The authors test this by computing pairwise attention similarity: for each amino acid pair, they calculate the correlation between their attention distributions across heads.

Figure 7: Attention-derived amino acid similarity strongly correlates with evolutionary substitution matrices, demonstrating learned biochemical relationships.
Left: Pairwise attention similarity matrix for TapeBert. For each amino acid pair, the value represents the Pearson correlation between their attention distributions across all 144 heads (e.g., the Pro-Phe entry correlates the heatmaps in Figure 6).
Right: BLOSUM62 substitution matrix, derived from evolutionary sequence alignments. Positive scores indicate amino acids that frequently substitute for each other while preserving protein function; negative scores indicate rarely observed substitutions.
Correlation: Pearson r = 0.73 between the two matrices for TapeBert. Similar values for ProtTrans models: 0.68 (ProtBert), 0.75 (ProtBert-BFD), 0.60 (ProtAlbert), 0.71 (ProtXLNet). Randomized null models yield r ≈ 0 (-0.05 to 0.21).
This alignment demonstrates that attention has learned biochemical properties—charge, hydrophobicity, size, aromaticity—that determine substitutability, purely from sequence statistics. The model hasn't memorized individual amino acid identities; it has internalized the dimensional structure of amino acid chemistry as encoded in evolutionary patterns. Source: Vig et al. 2022, Figure 7.
Comparing this attention-derived similarity matrix to BLOSUM62—a classical substitution matrix derived from evolutionary alignments—reveals striking agreement. TapeBert shows a Pearson correlation of 0.73 between attention similarity and BLOSUM62 scores. The ProtTrans models range from 0.60 (ProtAlbert) to 0.75 (ProtBert-BFD).
BLOSUM62 captures which amino acids can substitute for each other while preserving protein function—essentially encoding biochemical properties like charge, hydrophobicity, and size. The high correlations suggest that attention has learned these same biochemical relationships from sequence statistics alone, without explicit supervision.
As a control, randomized models yield near-zero correlations (-0.05 to 0.21), confirming that the BLOSUM alignment emerges from genuine learned patterns rather than architectural biases.
Visualizing attention in 3D structure
A methodological contribution of the paper is the 3D visualization of attention overlaid on protein structure (Figure 1). Using the NGL Viewer, the authors render proteins with attention weights as orange lines connecting amino acid positions, with line width proportional to attention magnitude.
This visualization reveals attention’s structural awareness in dramatic fashion. Figure 1a shows head 12-4 attending between residues 117D and 157I in a TIM-barrel protein—positions far apart in sequence but adjacent in the barrel’s characteristic architecture. Figure 1b shows head 7-1 targeting position 27G in HIV-1 protease, precisely the binding site for protease inhibitor drugs.
These visualizations serve as both validation and inspiration. They demonstrate that attention patterns correspond to meaningful structural features, and they suggest how domain experts might use attention analysis for biological discovery—identifying novel functional sites or structural relationships that haven’t been experimentally characterized.
Related work in context
Protein language models: a rapidly evolving landscape
The paper situates itself within an explosion of protein language model research. Early work applied Skip-gram embeddings to protein sequences (Asgari & Mofrad, 2015). TAPE (Rao et al., 2019) established benchmarks using a 12-layer Transformer trained on Pfam. ProtTrans (Elnaggar et al., 2020) scaled to 30-layer models trained on billions of sequences. ESM (Rives et al., 2021) from Facebook AI demonstrated that unsupervised embeddings capture structural and functional properties through linear transformations.
Concurrently, generative models emerged for protein design. ProGen (Madani et al., 2020), developed at Salesforce alongside the authors of this paper, demonstrated that autoregressive language models could generate novel protein sequences with plausible structures. This work directly connects to “Bertology Meets Biology”—Ali Madani appears as an author on both papers, and the interpretability insights here inform understanding of what generative models learn.
BERTology: a toolkit for neural network interpretation
The paper draws heavily on NLP interpretability methods, particularly the “BERTology” literature catalogued by Rogers et al. (2020). This includes:
- Probing classifiers (Veldhoen et al., 2016; Conneau et al., 2018): training simple classifiers on frozen representations to test what properties they encode
- Attention analysis (Clark et al., 2019): examining which attention heads correspond to syntactic dependencies
- Layer-wise studies (Tenney et al., 2019): demonstrating that BERT layers progressively encode POS tagging → syntax → semantics
The paper also engages with the “attention is not explanation” debate (Jain & Wallace, 2019; Wiegreffe & Pinter, 2019), carefully noting that all analyses are “purely associative” and don’t establish causal links between attention and predictions. This epistemic caution is important: high attention to contacts doesn’t prove the model uses contact information for predictions, only that attention correlates with contact structure.
From correlation to mechanism
The distinction matters for applications. If attention merely correlates with biologically meaningful features without driving model behavior, then interpreting attention could mislead researchers about what models actually use. The paper’s contribution is demonstrating strong, consistent correlations across multiple models and architectures—evidence that attention-property alignment is a robust phenomenon worth investigating, even if the causal story remains incomplete.
Implications for AI and biology
Scientific discovery through learned representations
The paper closes with a vision that extends beyond interpretation to discovery. While the analyses reconcile attention with known protein properties, the authors suggest attention might reveal novel relationships—”more nuanced forms of existing measures such as contact maps.”
This positions language models as potential tools for hypothesis generation. A head that strongly attends to specific residue pairs not annotated in current databases might point to uncharacterized functional relationships or structural contacts. Attention analysis becomes a means of surfacing what the model has learned from evolution that humans haven’t yet catalogued.
Enabling scientific trust
For protein language models to influence drug discovery, enzyme engineering, or therapeutic design, practitioners need confidence in what these models represent. The paper provides methodological infrastructure for building such trust—quantitative metrics for attention-property alignment, statistical validation against null models, and visualization tools for expert inspection.
Limitations and future directions
The work acknowledges important limitations. Attention analysis is correlational, not causal. The analyses focus on pretrained models without fine-tuning for specific tasks. And while the paper examines diverse architectures, the transformer landscape has evolved considerably since 2020, with models like ESM-2 scaling to 15 billion parameters and ESMFold achieving AlphaFold-competitive structure prediction.
Future work might extend these methods to larger models, examine attention patterns after task-specific fine-tuning, or develop causal interventions that probe whether attention-identified features actually drive predictions. The tools developed here—attention alignment metrics, probing protocols, 3D visualization—provide a foundation for this ongoing investigation.
Conclusion: structure emerges from sequence statistics
“Bertology Meets Biology” demonstrates something profound about the relationship between protein sequence and structure. Transformers trained solely to predict masked amino acids—with no supervision from experimental structures, no coevolution analysis, no biophysical simulations—spontaneously develop attention patterns that align with contact maps, binding sites, and amino acid substitution matrices.
This emergent structure awareness validates Anfinsen’s hypothesis computationally: the 3D structure of a protein is indeed determined by its amino acid sequence, and the statistical patterns across evolution encode sufficient information for neural networks to learn that determination. The models haven’t been taught protein biochemistry; they’ve inferred it from the patterns that evolution has written into sequence space.
For the machine learning community, the paper exemplifies how interpretability methods transfer across domains—the BERTology toolkit developed for natural language proves equally valuable for biological sequences. For computational biologists, it provides confidence that protein language models capture genuine biochemical knowledge, not merely statistical artifacts.
Perhaps most importantly, the work suggests a new mode of scientific interaction with AI systems. Rather than treating language models as black boxes that output predictions, we can interrogate their internal representations to understand—and potentially extend—biological knowledge. When attention lights up on a particular residue, that’s not just a number; it’s a hypothesis about function that domain experts can investigate.
The proteins have been speaking for billions of years through the language of evolution. Transformers are learning to listen. And through careful interpretability work, we’re beginning to understand what they hear.
Reference
The code and visualization tools described in this paper are available at github.com/salesforce/provis.
The paper was published at ICLR 2021 and is available on OpenReview.
Enjoy Reading This Article?
Here are some more articles you might like to read next: