BioEmu - A Biomolecular Emulator
Microsoft's breakthrough in protein dynamics prediction reshapes computational biology
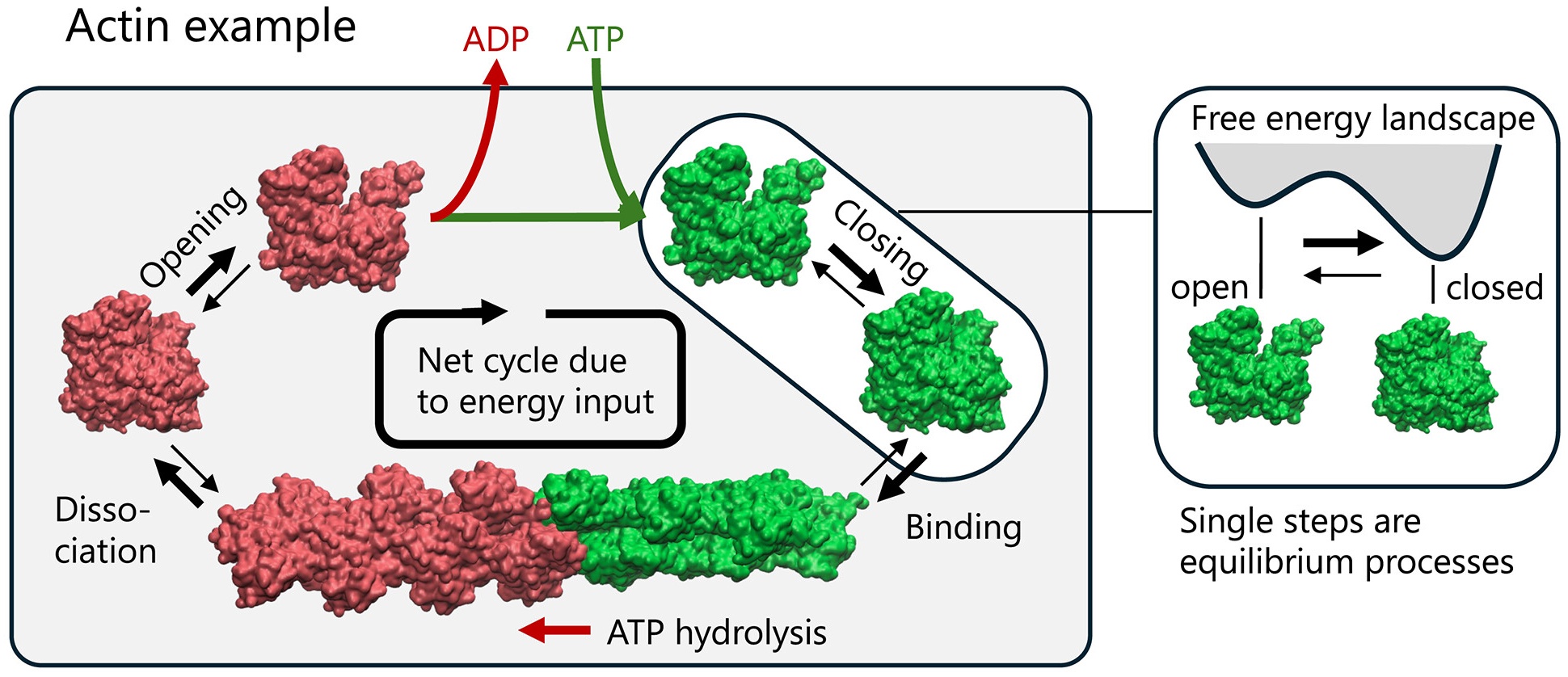
BioEmu marks the most significant advance in protein conformational prediction since AlphaFold, generating thousands of thermodynamically accurate protein structures per hour on a single GPU—100,000x faster than traditional molecular dynamics simulations while achieving ~1 kcal/mol free energy accuracy. Published in Science in August 2025 by Sarah Lewis, Tim Hempel, Cecilia Clementi, Frank Noé, and colleagues at Microsoft Research AI for Science, this generative deep learning system integrates over 200 milliseconds of MD simulation data, static structures, and experimental stability measurements through novel training algorithms. The community response has been overwhelmingly positive, with experts like Martin Steinegger declaring that “protein dynamics is the next frontier in discovery” and BioEmu “a significant step in this direction.” However, thoughtful critiques have emerged questioning whether computationally predicted ensembles can truly represent functionally relevant protein conformations in their native thermodynamic environments.
The architecture builds on diffusion foundations with novel training innovations
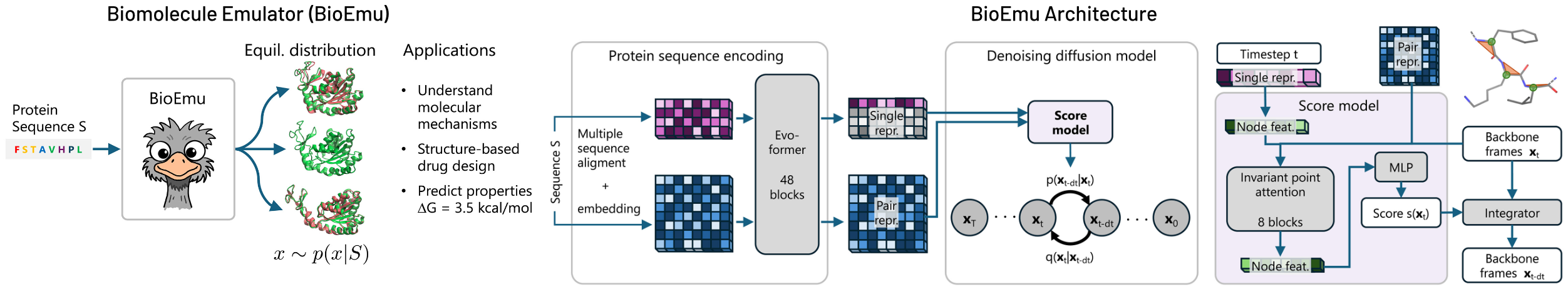
BioEmu is an E(3)-equivariant conditional diffusion model trained to sample residue-level protein conformations from a learned equilibrium distribution. Each protein is represented using a backbone frame parameterization where residue orientations are constructed via Gram-Schmidt orthogonalization of $\text{N–Cα}$ and $\text{C–Cα}$ bonds. The forward noising process operates as paired stochastic differential equations on positions (variance-preserving Gaussian) and orientations ($\text{IGSO(3)}$ distribution on $\text{SO(3)}$ manifold), ensuring the model learns orientation-equivariant denoising updates.
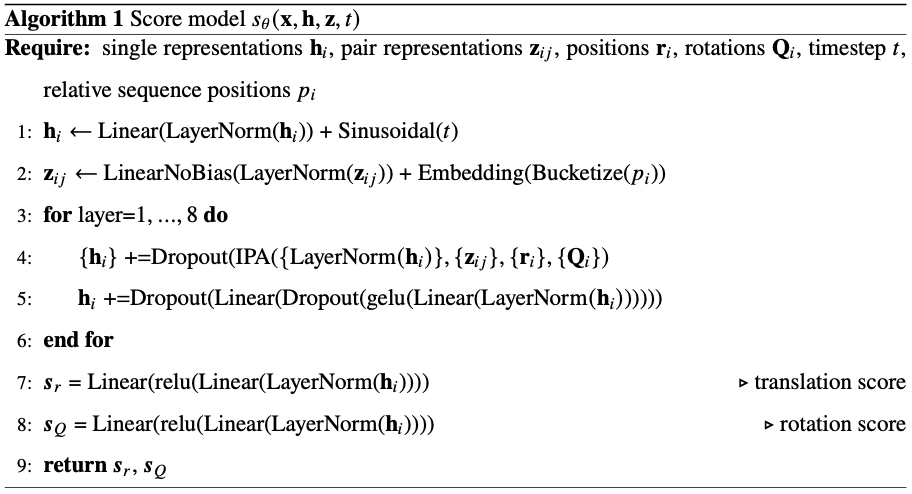

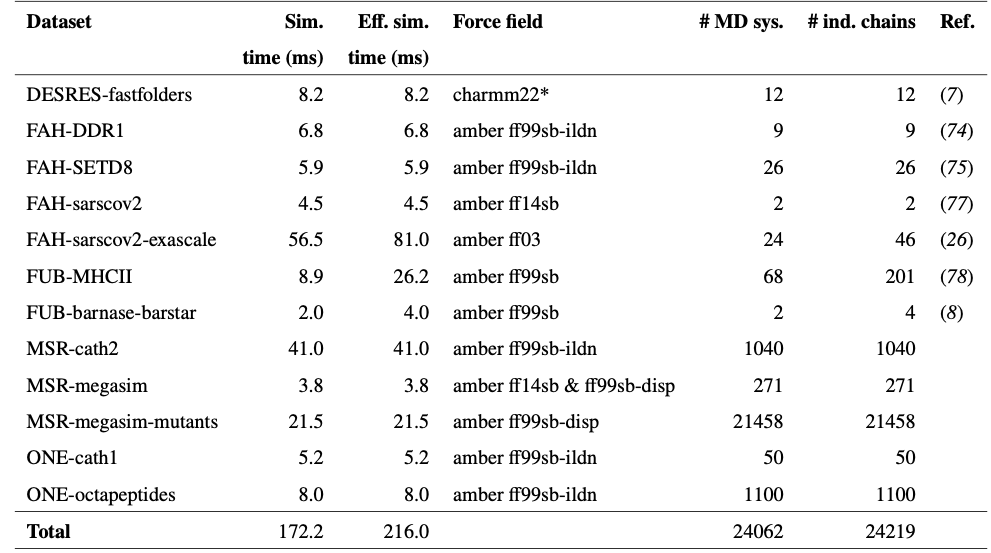
The key architectural innovation is a three-stage training pipeline. First, pre-training on clustered AlphaFold Database structures with data augmentation encourages diverse conformational associations from sequence. Second, continued training on more than 200 milliseconds of aggregate MD simulation data—reweighted using Markov state models to better approximate equilibrium populations—teaches realistic dynamics. Third, and most novel, Property Prediction Fine-Tuning (PPFT) aligns the learned distribution with experimental free energies from the MEGAscale protein stability dataset without requiring structural information for the measured proteins. This enables BioEmu to predict folding stability changes for mutations with remarkable accuracy.
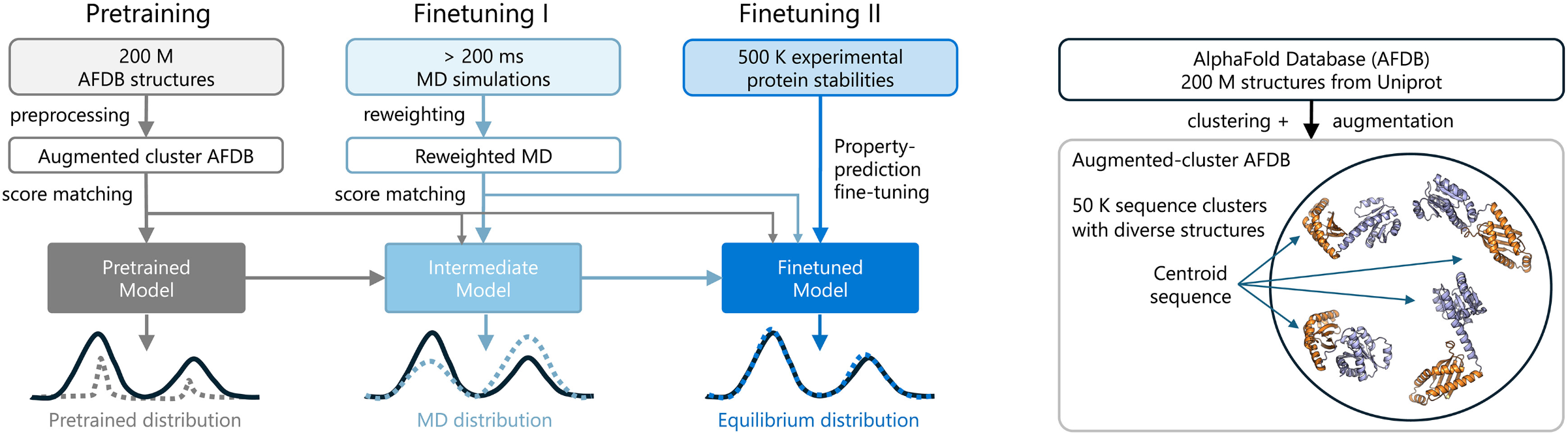
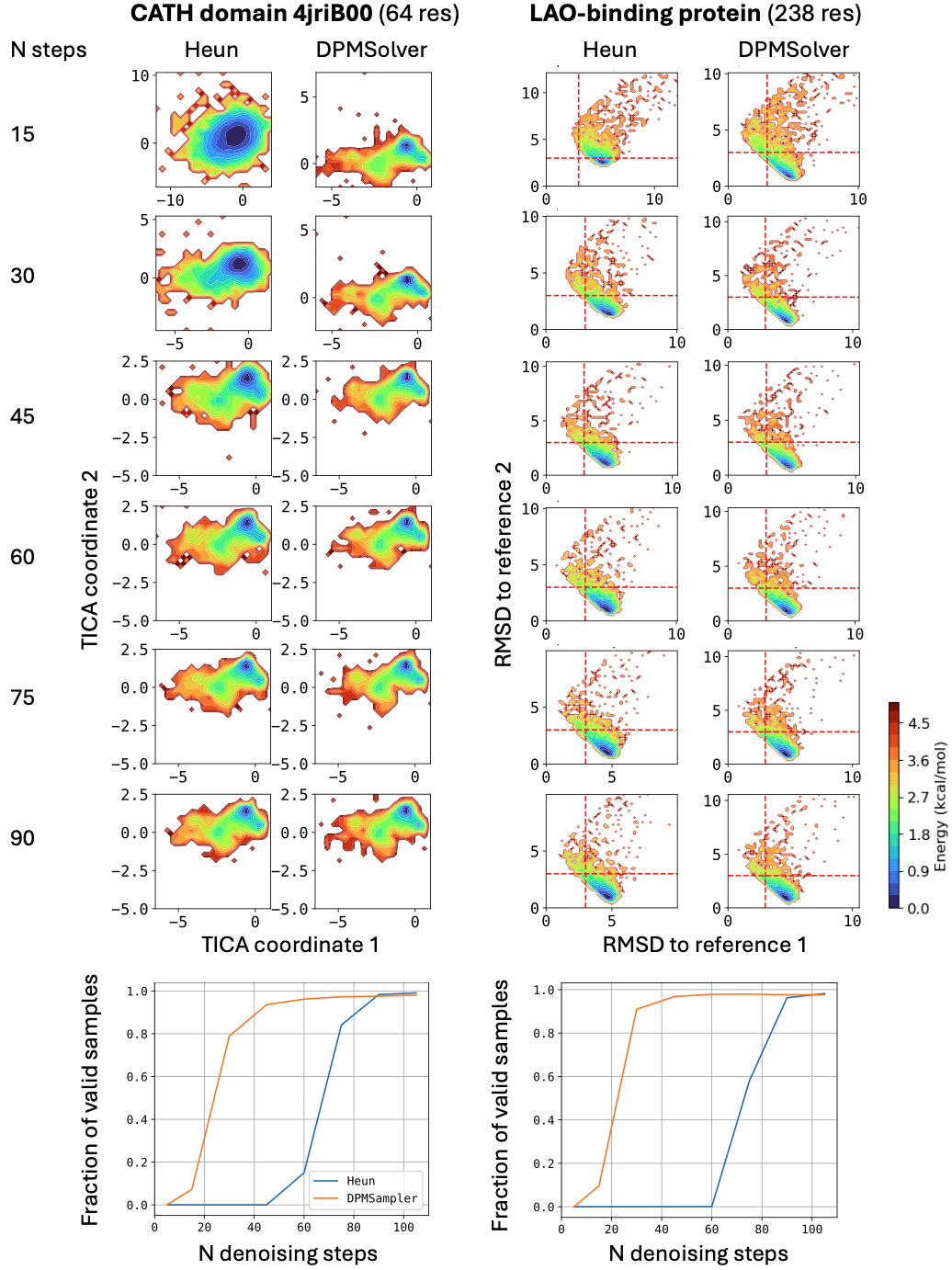
BioEmu leverages AlphaFold2’s Evoformer to compute single and pair representations, invoked only once per protein, then uses ~100 denoising steps with a second-order integration scheme (DPM-Solver) to generate structures. Performance benchmarks show 85% coverage of domain motions, 72-74% coverage of local unfolding states, and 49-85% coverage of cryptic pocket formation—all at ~1.9 GPU-seconds per sample compared to AlphaFlow’s ~32 GPU-seconds.
Technical comparisons reveal BioEmu’s unique positioning
BioEmu occupies a distinctive niche in the rapidly evolving landscape of protein conformational ensemble methods. AlphaFlow (Jing et al., ICML 2024) uses flow matching rather than diffusion, fine-tuning AlphaFold2/ESMFold with a polymer-structured harmonic prior. While AlphaFlow achieves excellent Pearson correlation (r=0.92 with templates) on the ATLAS benchmark, it was trained on only ~82 proteins and lacks explicit thermodynamic validation. BioEmu’s training scale of thousands of proteins with >200ms of MD data provides substantially broader coverage.
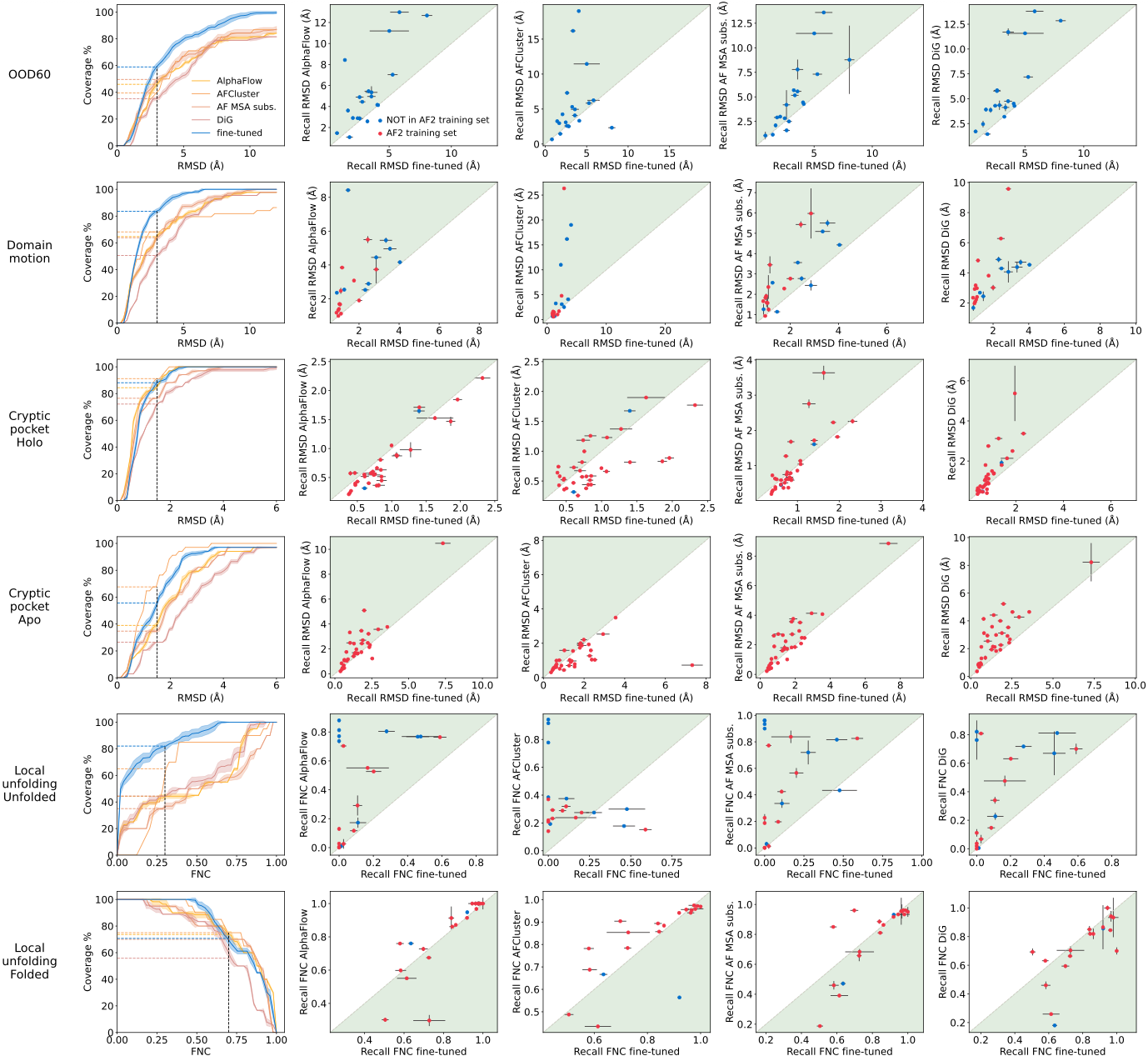
Distributional Graphormer (DiG) (Zheng et al., Nature Machine Intelligence 2024) provides BioEmu’s architectural foundation, using diffusion inspired by thermodynamic annealing and Physics-Informed Diffusion Pre-training (PIDP) for data-scarce cases. DiG demonstrated ~72% coverage of SARS-CoV-2 RBD conformational space but was not systematically validated for free energy accuracy. BioEmu extends DiG with the multi-stage training pipeline and experimental data integration.
AFCluster (Wayment-Steele et al., Nature 2024) represents a fundamentally different approach—clustering MSA sequences with DBSCAN to separate evolutionary signals encoding different conformational states, then running AlphaFold2 on each cluster. While successful for metamorphic proteins like KaiB and RfaH, AFCluster provides no thermodynamic predictions and requires diverse MSAs. Notably, recent work by Schafer et al. (Nature 2025) challenged AFCluster’s claims, arguing it underperforms random sequence sampling.
Boltzmann generators (Noé et al., Science 2019) from Frank Noé’s earlier work use normalizing flows for one-shot equilibrium sampling with tractable probability densities enabling exact reweighting. However, they remain limited to small systems (alanine dipeptide, BPTI at 58 residues) due to coordinate transformation challenges. BioEmu represents the conceptual successor—trading exact probability densities for scalability to hundreds of residues.
| Method | Architecture | Training Data | Free Energy | Speed (GPU-sec) |
|---|---|---|---|---|
| BioEmu | Diffusion | AFDB+200ms MD+Exp | ~1 kcal/mol | 1.9 |
| AlphaFlow | Flow matching | PDB/ATLAS | Not quantified | 32.0 |
| DiG | Diffusion | PDB+MD | Not systematic | Fast |
| AFCluster | MSA perturbation | None (AF2 pretrained) | None | Fast |
| Boltzmann Gen. | Normalizing flows | Energy functions | Exact | Fast (small systems) |
The broader literature positions BioEmu as the culmination of rapid progress
The past two years have seen explosive growth in diffusion and flow-matching models for proteins. RFdiffusion (Watson et al., Nature 2023) demonstrated that fine-tuning RoseTTAFold on denoising tasks enables outstanding protein backbone design, achieving picomolar-affinity binders. FrameDiff (Yim et al., ICML 2023) established theoretical foundations for SE(3)-invariant diffusion on multiple frames using Invariant Point Attention modules from AlphaFold2, generating designable monomers up to 500 amino acids without pretrained structure prediction networks.
Flow matching approaches have emerged as efficient alternatives. AlphaFlow demonstrated that fine-tuning structure prediction models under custom flow matching yields faster wall-clock convergence to equilibrium properties than MD. The MIT course “Flow Matching and Diffusion Models” (diffusion.csail.mit.edu) provides comprehensive tutorials explaining that “diffusion models and Gaussian flow matching are the same” mathematically but differ in network output specifications and sampling schedules.
SE(3)-equivariant architectures underpin all modern protein generative models. The e3nn library and tutorials (e3nn_tutorial) provide foundations for implementing spherical tensor operations. NequIP demonstrated remarkable data efficiency for interatomic potentials, while EGNN (E(n) Equivariant Graph Neural Networks) offers computationally efficient implementations without full spherical harmonics machinery.
For training with MD data, Markov state model reweighting has become essential. The approach recovers unbiased equilibrium populations from biased or short simulations through likelihood reweighting or path reweighting methods. BioEmu’s use of MSM-reweighted MD data represents the most ambitious application of this approach, aggregating over 200 milliseconds of simulation time across thousands of protein systems.
Property Prediction Fine-Tuning (PPFT): A novel training paradigm
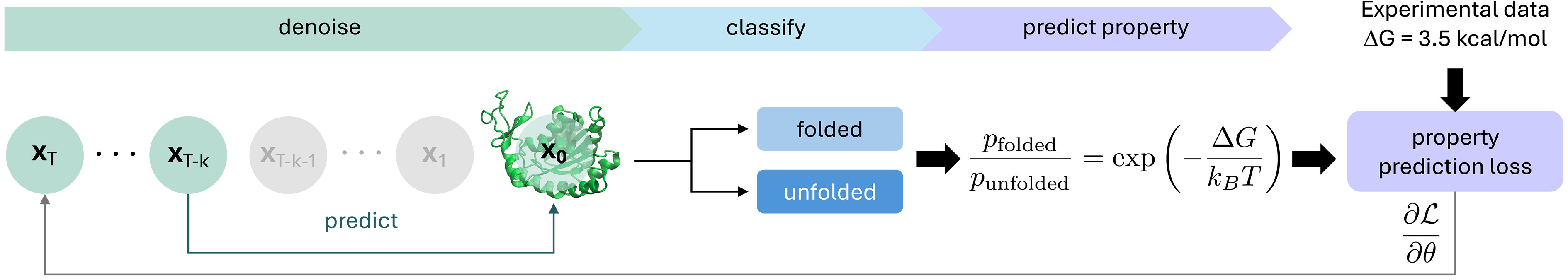

The PPFT innovation enables training on experimental observables without structural data. For the MEGAscale dataset (500,000+ stability measurements), BioEmu generates small ensembles (M~8-16 samples) using rapid approximate sampling, classifies each as folded/unfolded based on fraction of native contacts, computes ensemble-averaged foldedness, and minimizes squared error against the target derived from experimental ΔG via Boltzmann weighting.
Key technical tricks prevent mode collapse and reduce computational cost:
- Cross-target matching loss: Instead of minimizing $\mathrm{E[(f(x) - f_{target})^2}]$, minimize $\mathrm{(E[f(x)] - f_{target})^2}$ using pairs of samples, avoiding variance penalty that would collapse the distribution
- Partial denoising: Only 8 of 35 denoising steps executed, with clean sample extrapolation via reparameterization trick
- Selective backpropagation: Gradients computed only through final 3-5 steps, treating earlier steps as frozen
- Parameter freezing: Only layers 1 and 8 of the score model updated during PPFT, preserving conformational diversity learned in earlier stages
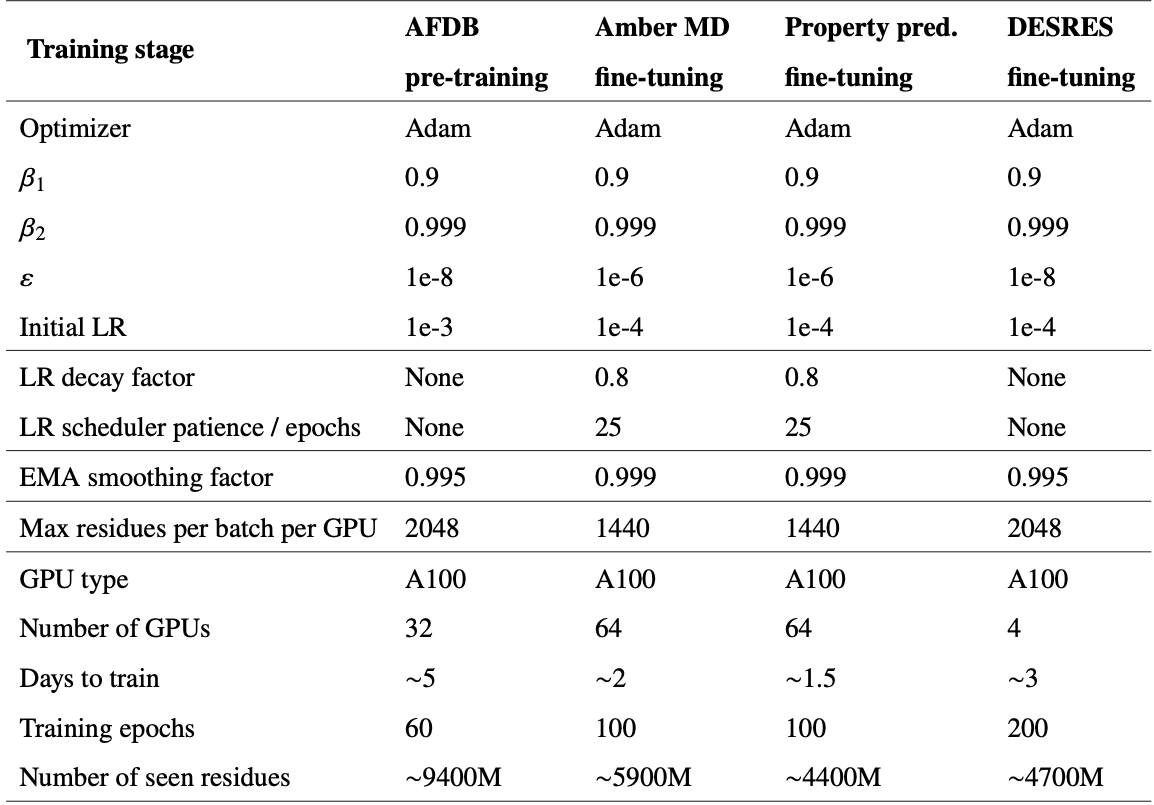
Emulating molecular dynamics equilibrium distributions
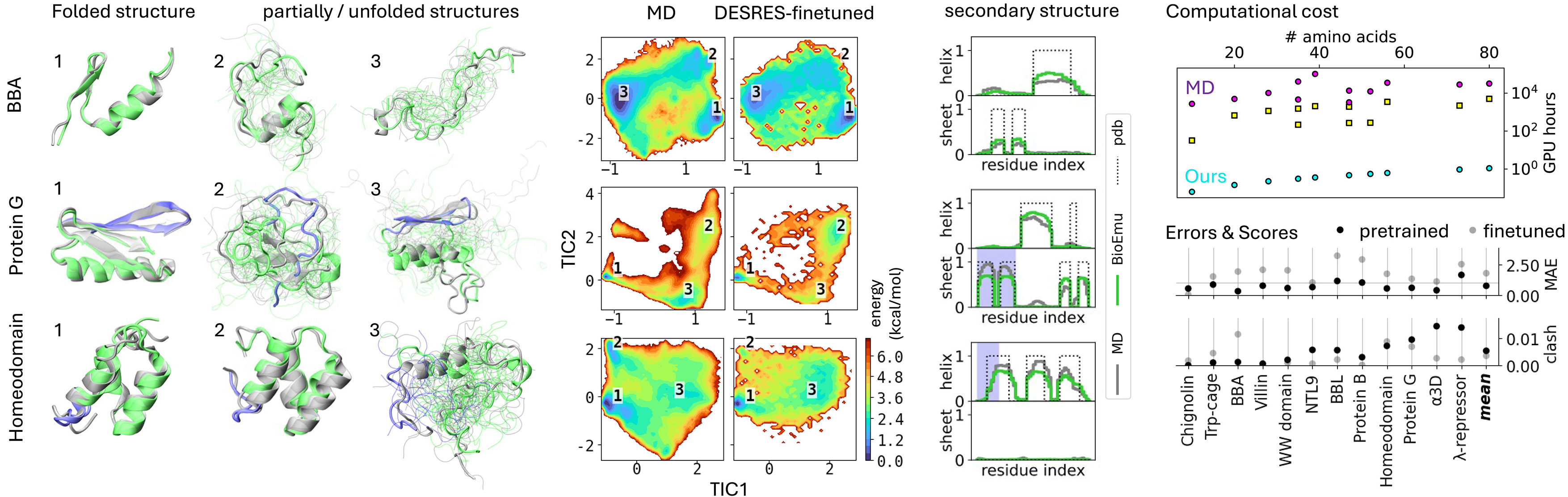
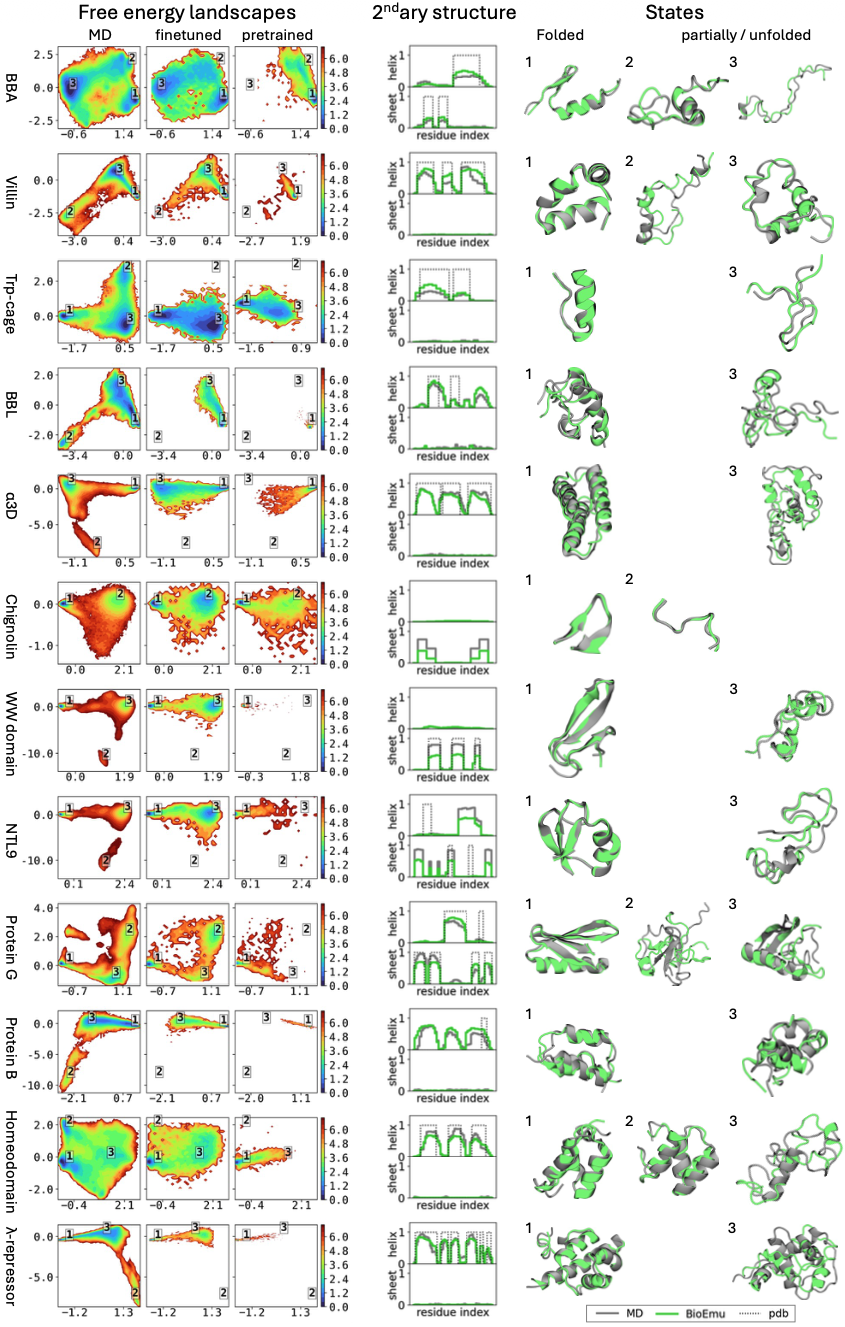
For the DESRES benchmark, BioEmu achieved 0.74 kcal/mol mean absolute error in free energy differences between states, comparable to differences between classical MD force fields. Critically, BioEmu predicts not just folded and unfolded basins but also folding intermediates visible in 2D projections—for Protein G, both MD and BioEmu sample intermediates with partial β-sheet formation.
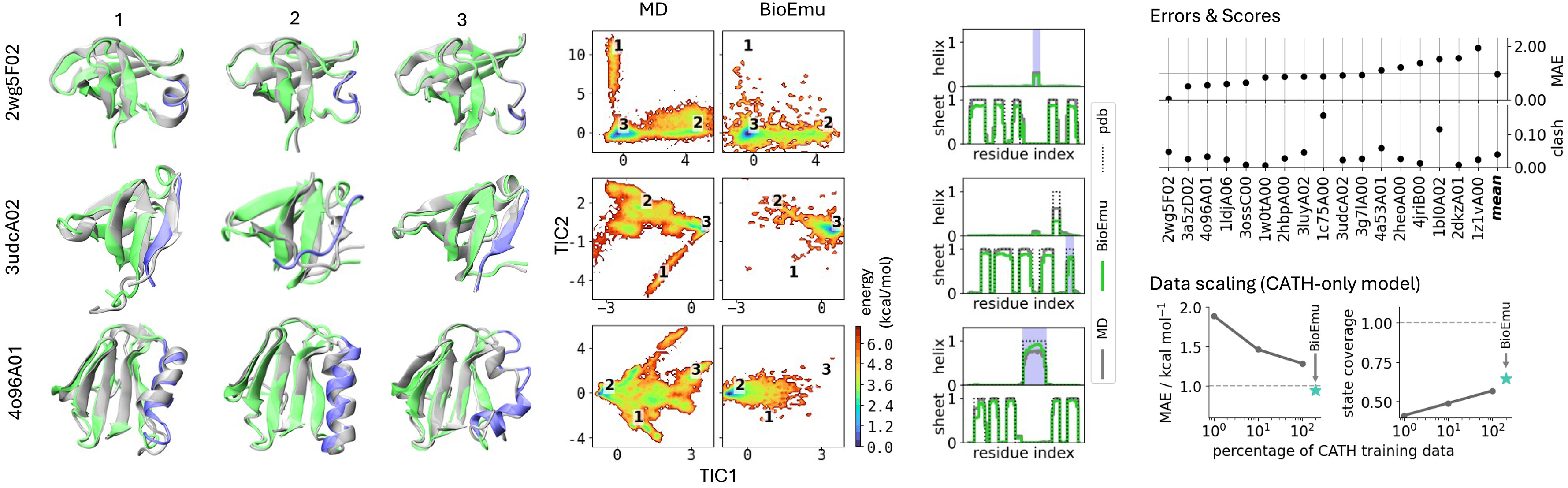
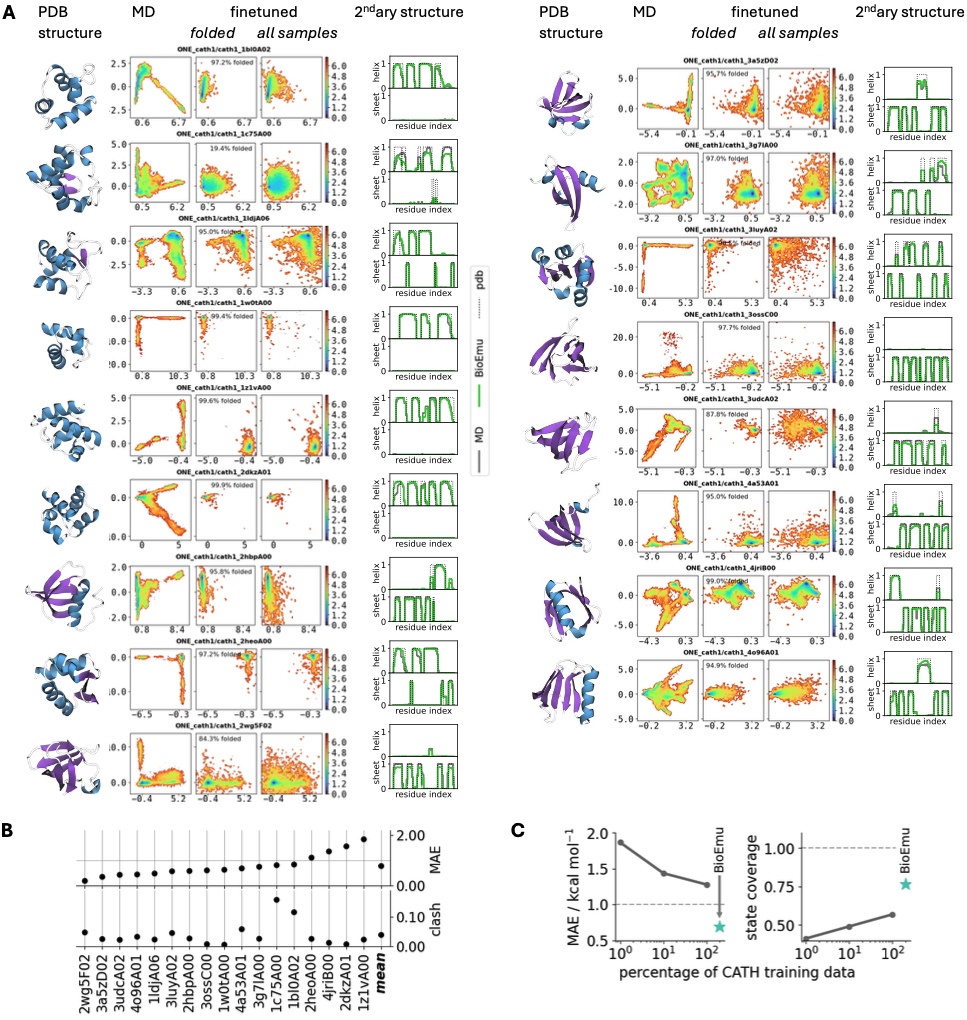
The CATH benchmark demonstrates generalization beyond training proteins. For 1040 CATH domains spanning diverse structural topologies, BioEmu was trained on varying fractions (1%, 10%, 100%) and evaluated on held-out domains. Free energy MAE decreased from ~1.5 to ~1.0 kcal/mol as training data increased, while state coverage improved from ~0.5 to ~0.75, suggesting continued improvement with more simulation data.
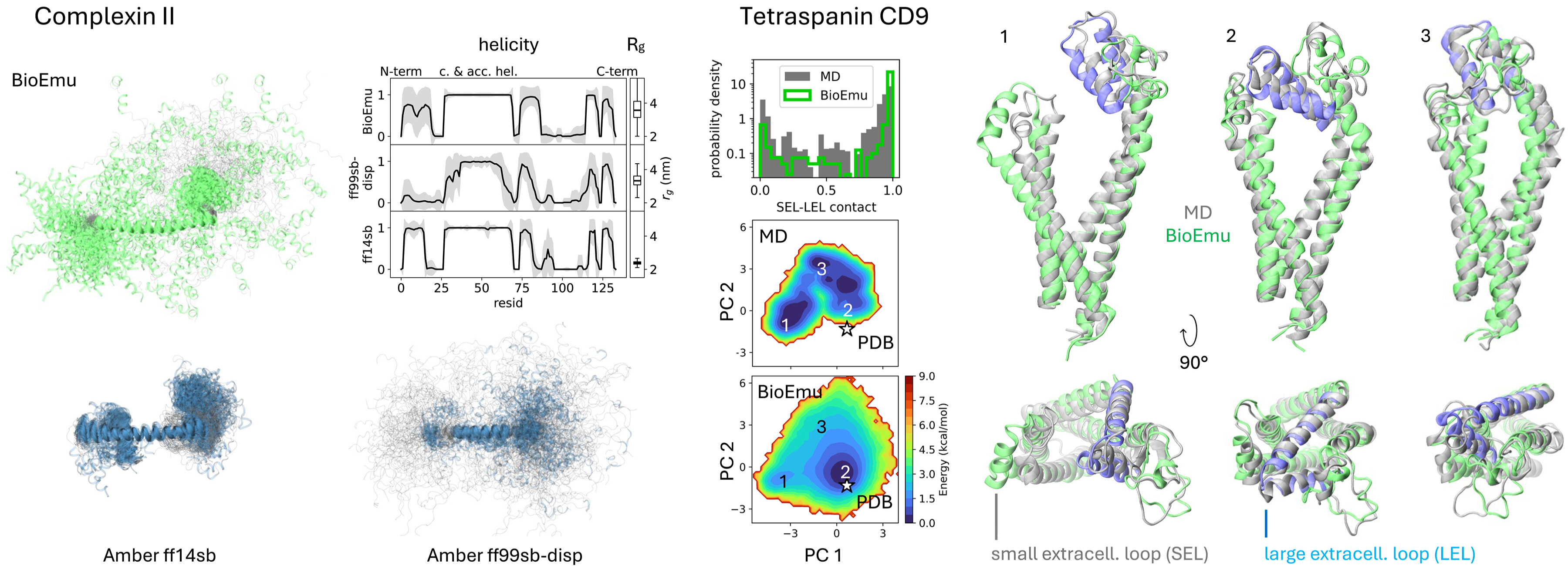
Multi-conformation sampling captures functional transitions

Performance varies by transition type:
- Domain motions: 83% coverage within 3Å RMSD (19 of 22 systems)
- Local unfolding: 70% coverage of folded states, 81% of unfolded states
- Cryptic pockets: 86% coverage of holo (bound) states, 56% of apo (unbound) states
The cryptic pocket asymmetry reveals a bias toward bound states, likely reflecting PDB composition (proteins often crystallized with multiple ligands but few apo structures). This suggests future training data curation should emphasize apo state diversity.
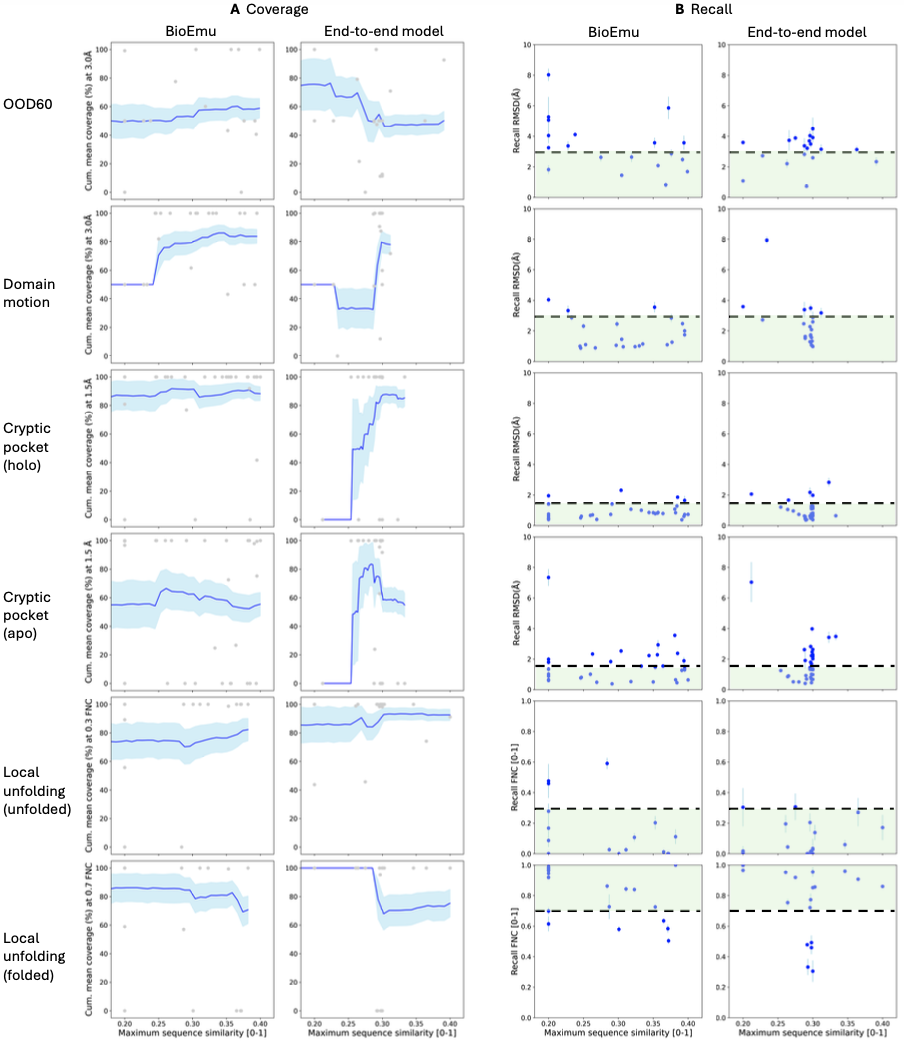
Predicting protein stability from equilibrium ensembles
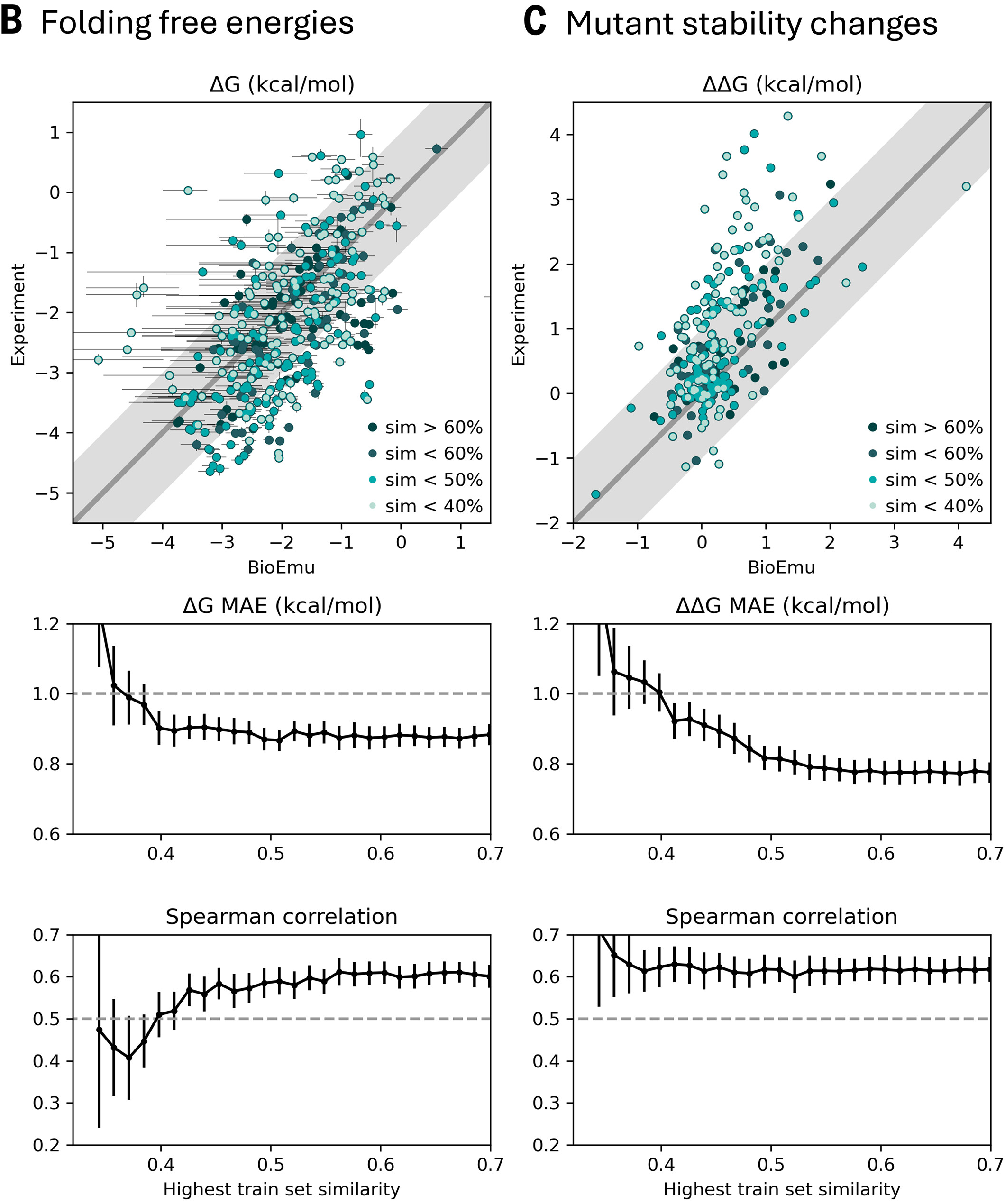
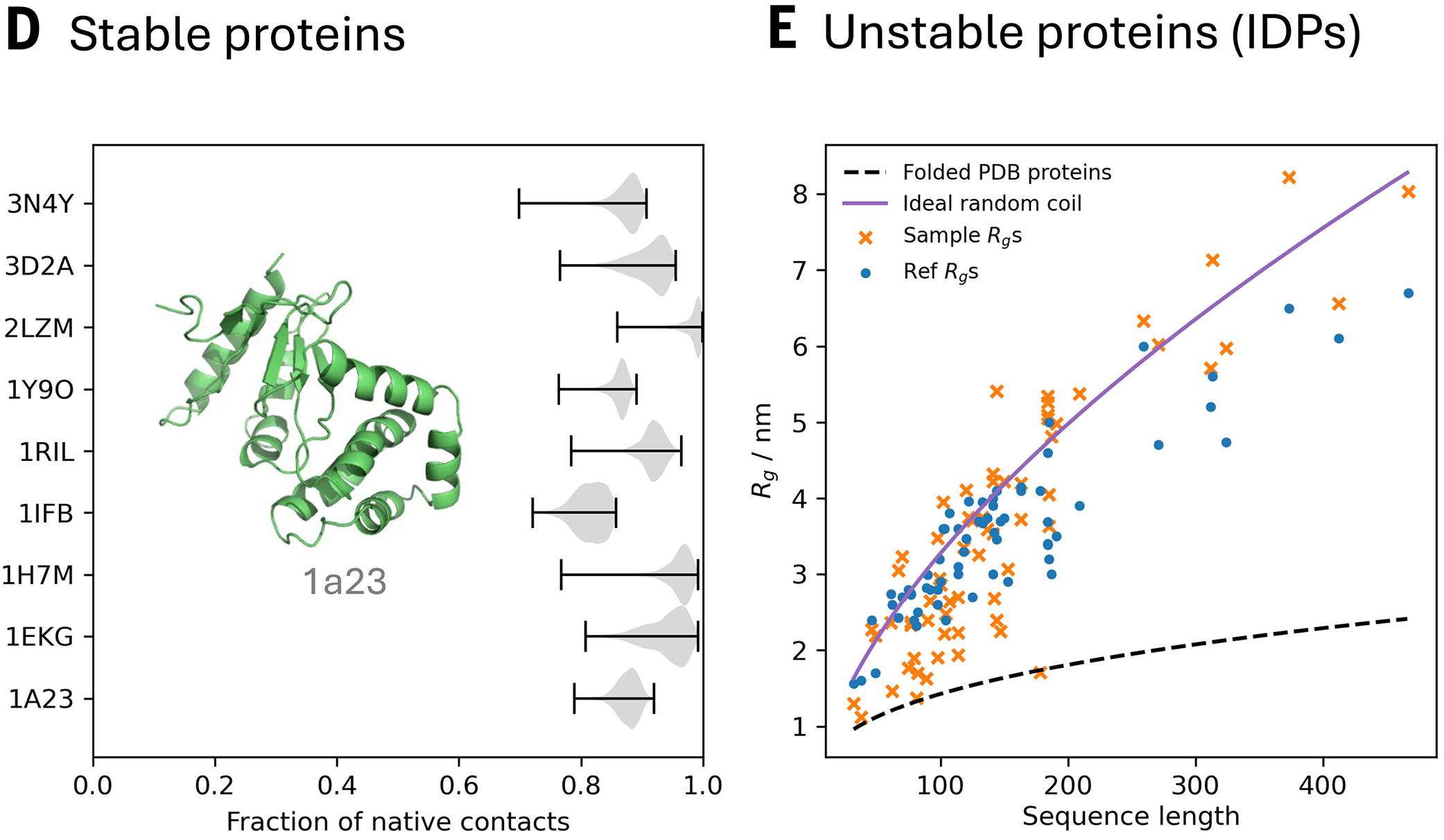
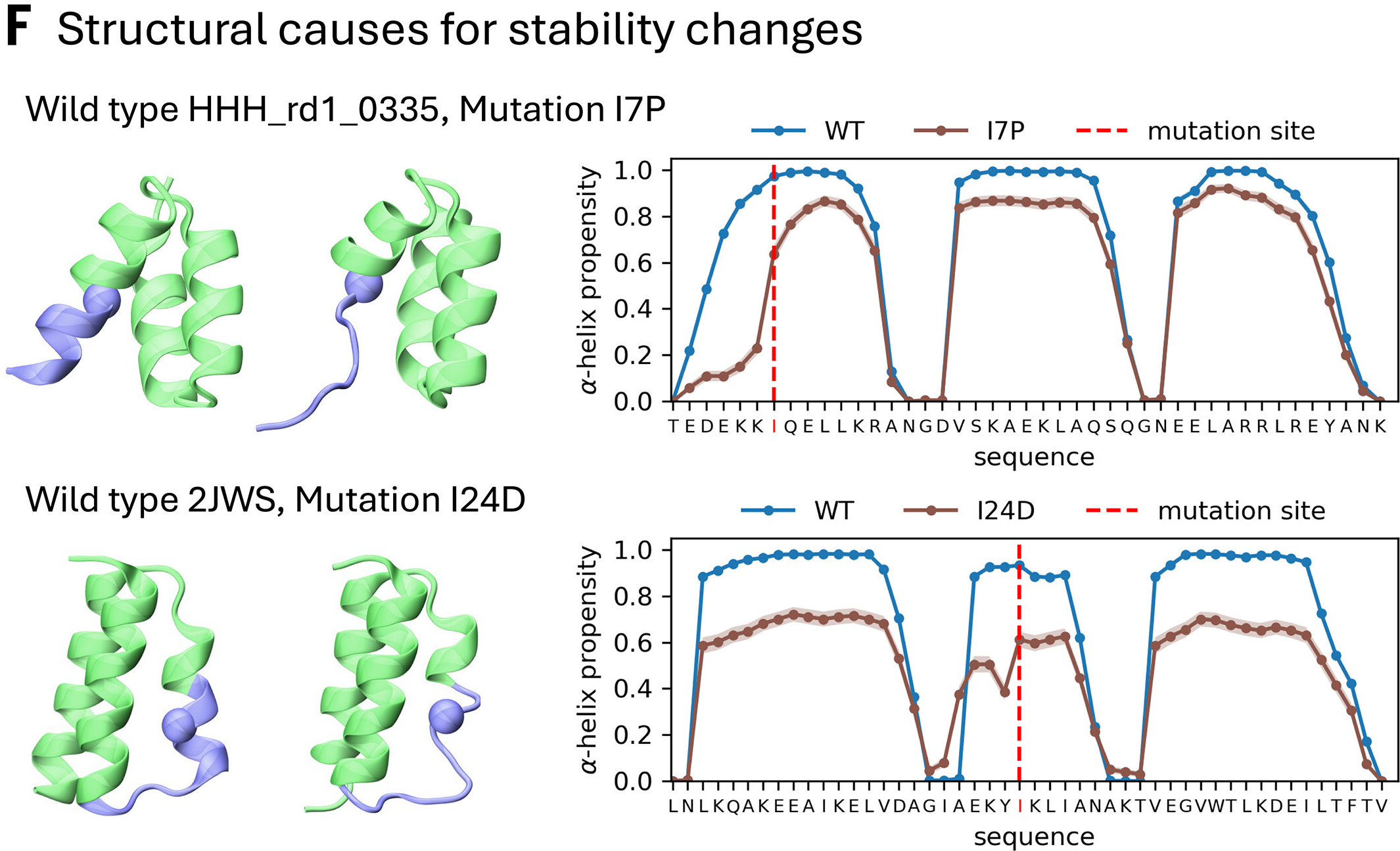
The key advantage over black-box ΔΔG predictors (ThermoMPNN, ESM-1v, ProteinMPNN) is interpretability—BioEmu explains why mutations destabilize by showing structural changes in the ensemble, enabling rational design iterations.
Community reception combines enthusiasm with thoughtful critique
The scientific community has responded with significant enthusiasm. Martin Steinegger (Seoul National University) stated that “protein dynamics is the next frontier in discovery” and BioEmu “marks a significant step in this direction by enabling blazing-fast sampling of the free-energy landscape.” Zhidian Zhang (MIT) praised BioEmu for predicting “the distribution of different conformations, which is a much more difficult problem” than static structure prediction. Alberto Perez (University of Florida) expressed intention to “use BioEmu in my own work” and appreciation for the open-source release.
Nature Methods published a research highlight noting the “urgent need for methods that predict protein structural changes at scale.” Chemical & Engineering News positioned BioEmu as going “beyond AlphaFold.” Microsoft CEO Satya Nadella mentioned BioEmu in a July 2025 post about reducing protein motion analysis from years to hours.
Understanding protein motion is essential to understanding biology and advancing drug discovery.
— Satya Nadella (@satyanadella) July 10, 2025
Today we’re introducing BioEmu, an AI system that emulates the structural ensembles proteins adopt, delivering insights in hours that would otherwise require years of simulation. https://t.co/tZDJ3tUjZx
Papers citing BioEmu have already emerged. eRMSF (Arantes et al., J Chem Inf Model 2025) provides a Python package specifically designed for ensemble-based RMSF analysis including BioEmu-generated ensembles. ESMDynamic (Kleiman et al., bioRxiv 2025) benchmarks against BioEmu, claiming to match or outperform it for transient contact prediction while offering orders-of-magnitude faster inference.
However, Sarfaraz K. Niazi (University of Illinois) published a formal critique in Science arguing that BioEmu’s
Drug discovery applications center on cryptic pockets and protein stability
BioEmu’s ability to sample conformational changes has immediate drug discovery relevance. Cryptic pocket prediction—identifying binding sites absent in ground-state structures—could potentially double the druggable proteome. PocketMiner (Meller et al., Nature Communications 2023) achieved ROC-AUC of 0.87 for cryptic pocket identification, but BioEmu enables direct sampling of pocket-forming conformations rather than prediction from static features.
Traditional MD simulations require 100 microseconds to 10 milliseconds of simulation time for comprehensive conformational discovery, achievable only with special-purpose supercomputers (D.E. Shaw’s Anton) or massive distributed computing (Folding@home). BioEmu generates equilibrium ensemble snapshots in minutes, enabling proteome-scale analysis. The critical trade-off: BioEmu generates statistical samples from the equilibrium distribution, not time-ordered trajectories, so kinetic pathways between states cannot be modeled.
For protein stability prediction, BioEmu’s PPFT-derived thermodynamic accuracy complements existing methods. ThermoMPNN (Dieckhaus et al., PNAS 2024) achieves state-of-the-art benchmark performance through transfer learning, while RaSP provides deep learning predictions at >10,000x the speed of Rosetta. BioEmu’s unique contribution is explaining structure-stability relationships by analyzing generated ensembles—revealing mechanistic causes of mutant destabilization.
Industry adoption is accelerating. Relay Therapeutics uses its Dynamo platform for protein dynamics in drug discovery. Recursion conducts 2M+ experiments weekly with AI integration. Cradle raised $73M in 2024 for AI-powered protein engineering with major pharma partnerships. The investment thesis is compelling: AI-discovered drugs show 80-90% Phase 1 success rates compared to industry averages.
Technical resources enable practitioners to implement these methods
Yang Song’s foundational blog post provides the definitive introduction to score-based generative models, with Google Colab tutorials in JAX and PyTorch. For SE(3) equivariance, the official e3nn tutorial covers spherical tensor data types and practical implementation. The MIT course on flow matching (diffusion.csail.mit.edu) offers systematic development with hands-on Colab labs.
BioEmu’s code, model weights, and the largest sequence-diverse protein simulation dataset publicly available (>200ms of trajectories) have been released through Microsoft Research. The bioRxiv preprint provides full methodological details. For MSM reweighting, the Girsanov reweighting implementation guide (Schäfer and Keller, J Phys Chem B 2024) offers integration with OpenMM and the Deeptime time-series analysis package.
Conclusion: A transformative tool with clearly defined boundaries
BioEmu represents a genuine breakthrough in computational biology—not as a replacement for MD simulation, but as a complementary tool enabling new scales of hypothesis generation. The combination of AlphaFold-derived sequence encoding, diffusion-based structure generation, and experimental data integration through PPFT achieves what neither pure physics-based simulation nor pure data-driven learning could accomplish alone.
The key insight is amortization: the enormous cost of generating 200+ milliseconds of MD trajectories and half a million stability measurements is paid once during training, then amortized across infinite future predictions. For any new protein sequence, BioEmu provides rapid access to thermodynamically calibrated conformational ensembles without new simulations.
Critical limitations deserve emphasis. BioEmu generates equilibrium snapshots, not dynamics—kinetic mechanisms remain inaccessible. The restriction to soluble, single-chain proteins at 300K excludes membrane proteins, protein complexes, and temperature-dependent phenomena crucial for many therapeutic targets. The empirical learning of energy landscapes, rather than physics-based potential energy functions, means extrapolation beyond training data may be unreliable.
These boundaries are clearly defined, enabling appropriate use. For high-throughput screening of conformational changes across protein families, cryptic pocket discovery, and stability prediction, BioEmu offers transformative capability. For detailed mechanistic understanding of specific drug-target interactions, traditional MD with explicit ligands and membranes remains essential. The future likely lies in hybrid workflows combining both approaches—ML for rapid exploration, physics for detailed characterization.
Main Reference
Lewis et al., Scalable emulation of protein equilibrium ensembles with generative deep learning, Science 2025.
Enjoy Reading This Article?
Here are some more articles you might like to read next: