BoltzGen Redefines Protein Binder Design
with Unified All-Atom Architecture
BoltzGen represents a fundamental shift in computational protein design by unifying structure prediction and binder generation in a single all-atom diffusion model. Released in November 2025 by MIT’s Jameel Clinic, this fully open-source tool (MIT license) achieves 66% experimental success rates on truly novel targets—proteins with less than 30% sequence identity to any bound structure in the PDB. Unlike the dominant two-stage pipeline of RFdiffusion + ProteinMPNN, BoltzGen simultaneously generates atomic coordinates and residue identities through a novel geometric encoding scheme, eliminating the information loss between backbone generation and sequence design. Early validation includes the first demonstrated binding to an intrinsically disordered protein region in living cells.
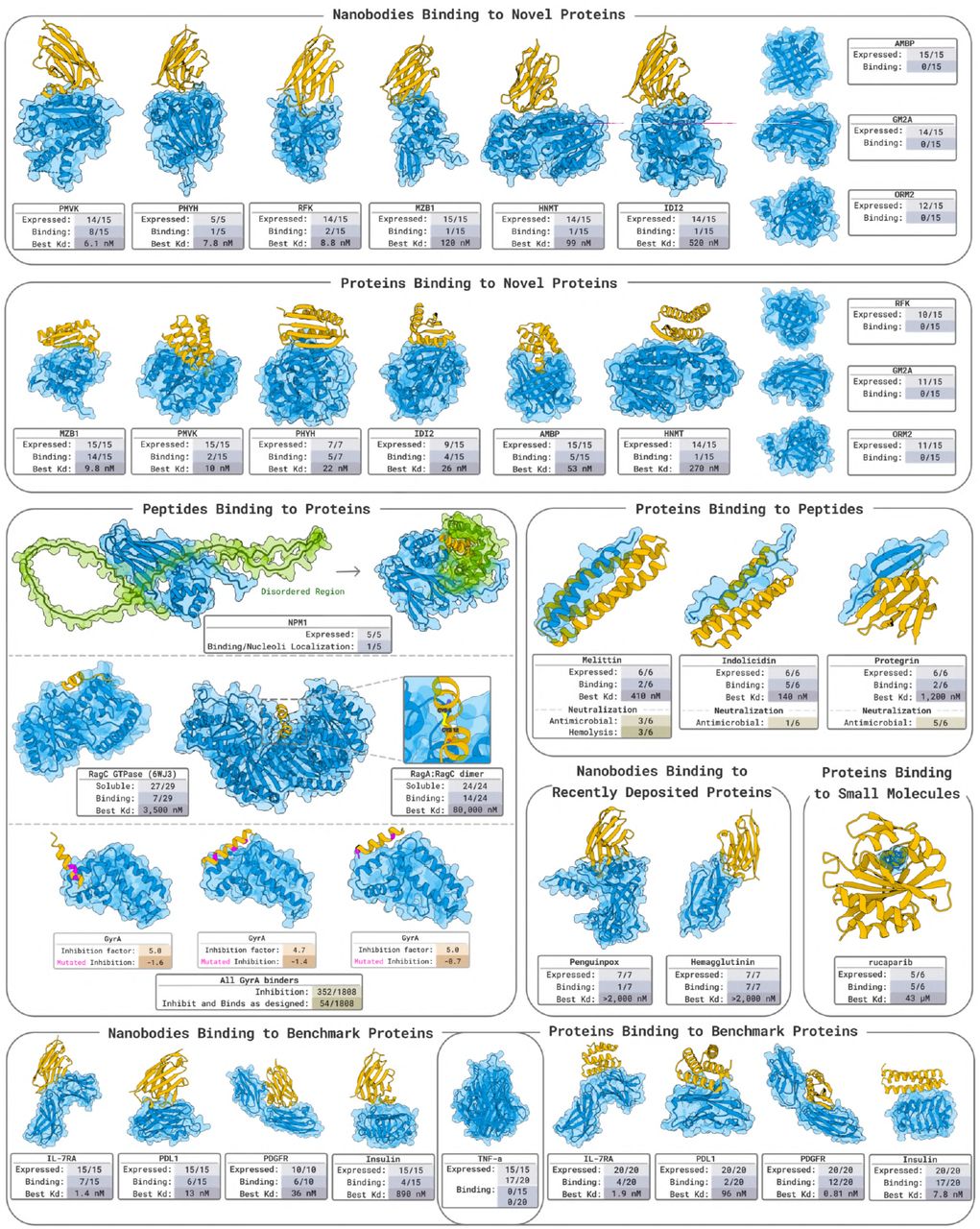

Figure 1: BoltzGen experimental validation campaign. (a) Nine novel target proteins with $\lt 30%$ sequence identity to bound PDB structures. (b) Design success rates for nanobodies (66%, 6/9 targets) and miniproteins (66%, 6/9 targets) with representative structures. (c) Best validated affinities: 1.9 nM for AMBP, 6.1 nM for PMVK. Source: Stark et al. 2025, Figure 1.
The 14-atom breakthrough solves the continuous-discrete problem
BoltzGen’s core innovation addresses a fundamental tension in protein design: structure exists in continuous 3D space while sequence is discrete. Previous approaches either decouple these problems (RFDiffusion generates backbone, ProteinMPNN designs sequence) or use discrete tokens with limited geometric reasoning. BoltzGen solves this with a 14-atom representation where each designed residue is encoded as exactly 14 virtual atoms. The first four atoms occupy fixed backbone positions ($\mathrm{N, Cα, C, O}$), while the remaining 10 encode residue identity through their geometric configuration.
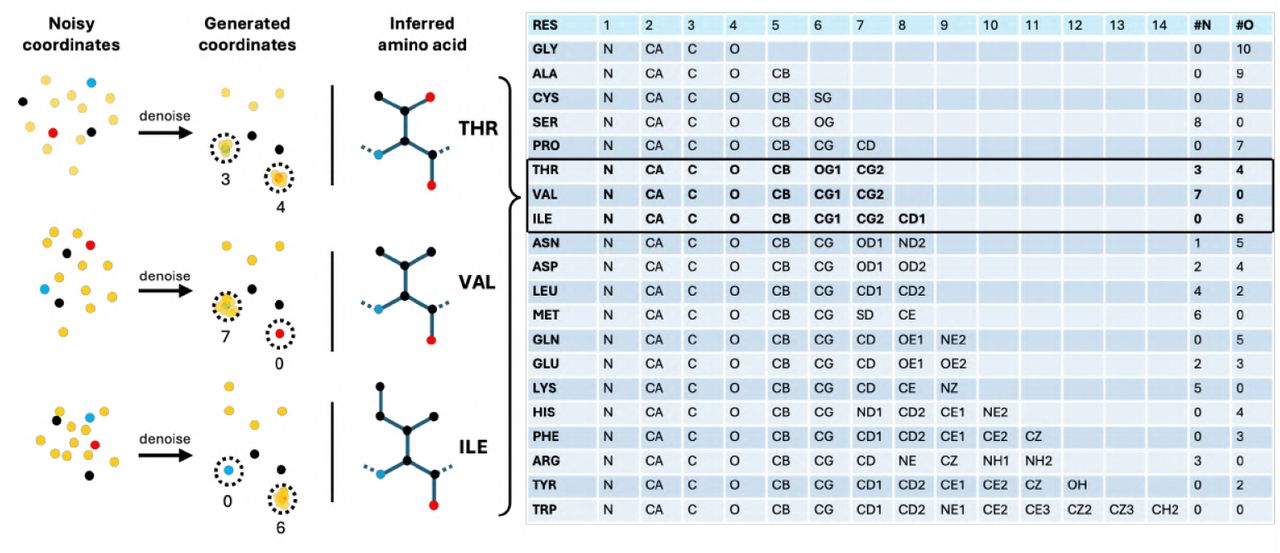
Figure 2: Geometric residue encoding via 14 virtual atoms. Each designed residue uses 4 backbone atoms ($\mathrm{N, Cα, C, O}$) plus 10 virtual atoms superposed on backbone positions to encode amino acid identity. Example: proline = 7 atoms on oxygen, threonine = 3 on nitrogen + 4 on oxygen. Residue type decoded by counting atoms within 0.5Å of each backbone position. Source: Stark et al. 2025, Figure 7.
The model learns to superpose virtual atoms onto designated backbone positions to signal amino acid type—for example, proline is encoded by 7 atoms placed on the backbone oxygen, while threonine uses 3 atoms on nitrogen and 4 on oxygen. Residue identity is recovered by counting atoms within 0.5Å of each backbone position, enabling fully differentiable joint training. This scheme allows BoltzGen to match state-of-the-art on structure prediction while also generating novel designs—a capability no previous design model achieved.
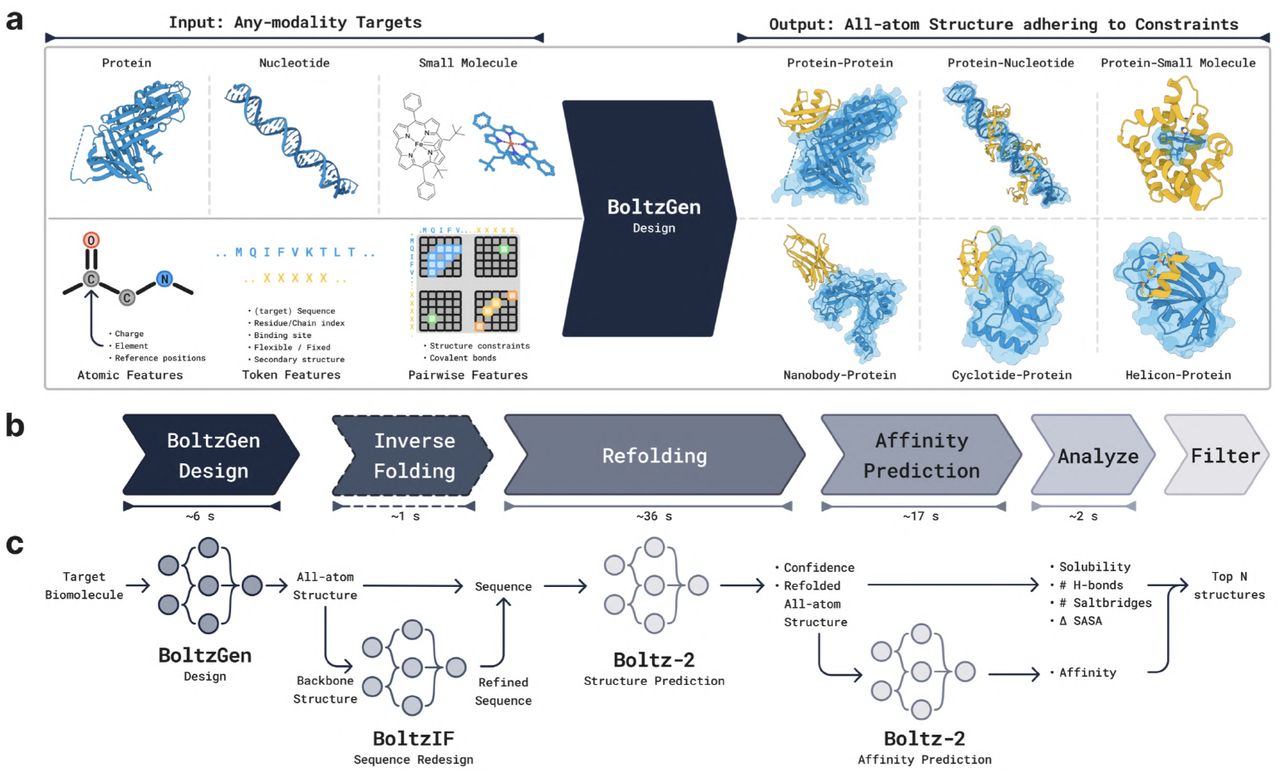
Figure 3: BoltzGen unified architecture. (a) PairFormer trunk processes tokenized structures to generate pairwise representations. (b) Diffusion module with atom-level and token-level Transformers iteratively denoises 3D coordinates. (c) Training alternates between structure prediction, binder design, and structure completion tasks. Source: Stark et al. 2025, Figure 6.
The architecture inherits components from AlphaFold3 and Boltz-2: a PairFormer trunk with Triangle Attention processes tokenized structures once per design, generating pairwise representations, while the diffusion module iteratively denoises 3D coordinates using a Transformer operating at atom and token levels. Training randomly alternates between structure prediction, binder design, and structure completion tasks, enabling multi-task synergy across ~6GB of model weights.
How BoltzGen compares to the competitive landscape
The protein design field has stratified into distinct methodological camps, each with characteristic strengths and limitations:
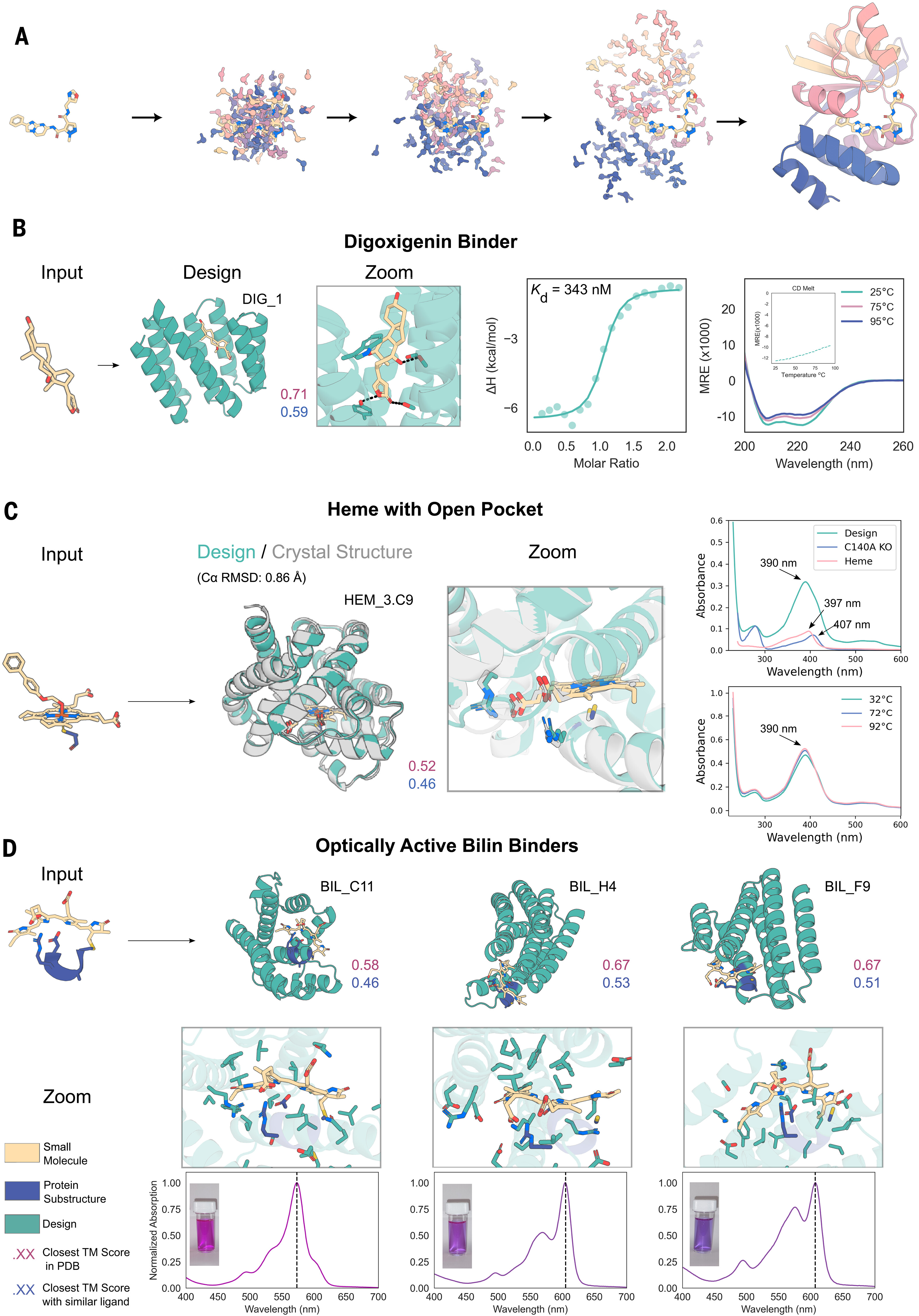
Figure 4: RFDiffusion two-stage pipeline. (a) Backbone diffusion generates $\text{Cα}$ traces. (b) ProteinMPNN designs sequences from backbone. (c) AlphaFold2 validates designs. Contrast with BoltzGen's single-stage approach. Source: Watson et al., Nature 2023, Figure 2.
RFdiffusion (Baker Lab, Nature 2023) established diffusion-based design by fine-tuning RoseTTAFold on structure denoising. It generates backbone-only coordinates, requiring ProteinMPNN for sequence design—a two-stage pipeline that achieved picomolar binders (340 pM to PTH) and approximately 18% experimental success rates. The all-atom variant, RFDiffusionAA (Science 2024), extends to small molecules and nucleic acids, generating custom binding pockets with validated affinities like 343 nM for digoxigenin binders. The latest RFDiffusion3 (November 2025) repurposes AlphaFold3’s architecture for joint backbone and side-chain generation, achieving 9% success for DNA-binding proteins.
AlphaFold3 and Chai-1 represent prediction-focused architectures. AlphaFold3’s all-atom diffusion predicts structures across proteins, nucleic acids, and small molecules with remarkable accuracy but offers no generative design capability—it evaluates rather than creates. Chai-1 similarly provides multi-modal prediction with 77% success on PoseBusters benchmarks but cannot generate novel binders. Both serve as validation tools for designs from generative models.
ProteinMPNN remains the workhorse for inverse folding, achieving 52% sequence recovery from backbone coordinates—essential in RFDiffusion pipelines but creating the information gap BoltzGen eliminates. Extensions like LigandMPNN handle small molecules and metals, while AntiFold specializes in antibody CDRs.
| Tool | Approach | All-Atom | Joint Struct/Seq | Best Validated Affinity | Open Source |
|---|---|---|---|---|---|
| BoltzGen | Unified diffusion | $\surd$ | $\surd$ | 1.9 nM (AMBP) | $\surd$ MIT |
| RFDiffusion | Backbone diffusion | Backbone only | Two-stage | 340 pM (PTH) | $\surd$ BSD |
| RFDiffusionAA | All-atom diffusion | $\surd$ | Partial | 343 nM (digoxigenin) | $\surd$ |
| BindCraft | AF2 backprop | Side chains | $\surd$ | Sub-nM | $\surd$ MIT |
| Chroma | Polymer diffusion | $\surd$ | Two-stage | µM range | Academic |
| AlphaFold3 | Prediction only | $\surd$ | N/A | N/A | Academic only |
Emerging flow matching approaches like FrameFlow achieve comparable designability with 5× fewer sampling steps than diffusion, while OriginFlow claims 95% wet-lab success rates on benchmark targets—though independent validation remains pending.
Evolution from hallucination to unified models spans four distinct eras
The field’s transformation from 2022-2025 follows a clear trajectory through four paradigms:
Hallucination (2020-2022): trRosetta hallucination iteratively optimized random sequences via Monte Carlo sampling, requiring 20,000-40,000 steps to converge. Success rates were low—27/129 designs folded correctly—and compute costs high. RFjoint Inpainting offered faster deterministic generation but limited diversity.
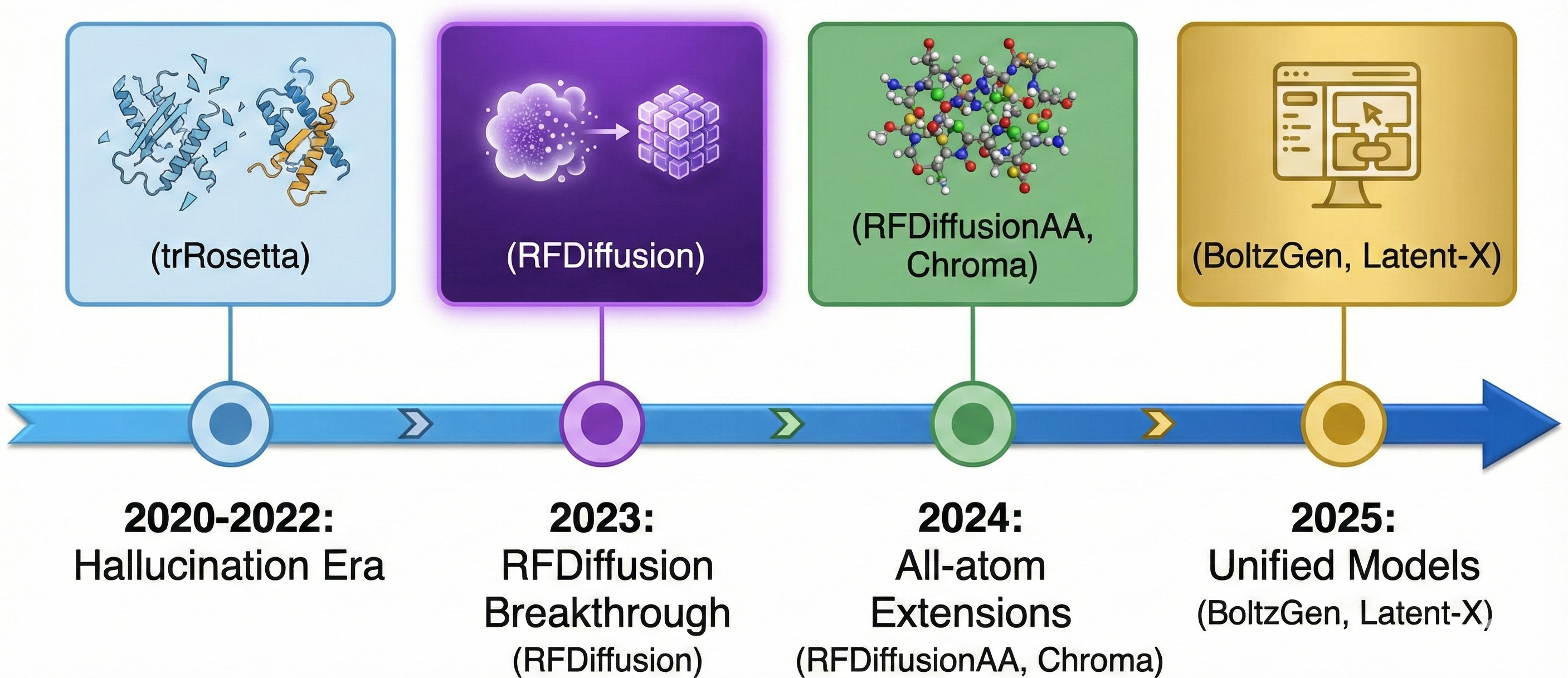
Backbone diffusion (2023): RFdiffusion’s Nature publication established the paradigm of fine-tuning structure prediction networks on denoising tasks. The critical insight was that pre-trained structure predictors contain geometric priors essential for generation—training from scratch performs substantially worse. This era validated hundreds of designs at single-digit nanomolar affinities but required separate inverse folding.
All-atom extension (2024): RFDiffusionAA, Chroma, and similar tools expanded to full atomic representations, enabling small molecule binder design and multi-modal targets. However, most still separated structure and sequence generation.
Unified models (2025): BoltzGen, Latent-X, and refined BindCraft workflows now jointly generate structure and sequence. Latent-X reports 91-100% hit rates for macrocycles and 10-64% for mini-binders, claiming an order of magnitude speed improvement over multi-step pipelines. BoltzGen’s geometric encoding uniquely enables single-model training across prediction and design tasks.
Wet lab validation reveals target-dependent success rates
Experimental characterization remains the ultimate test, with success rates varying dramatically by target and modality. BoltzGen’s validation campaign tested 9 novel targets (less than 30% sequence identity to bound PDB structures) with both nanobodies and miniproteins, achieving 66% target coverage (6/9 targets) for each modality. Best affinities reached 1.9 nM for AMBP and 6.1 nM for PMVK—competitive with therapeutic-grade candidates.
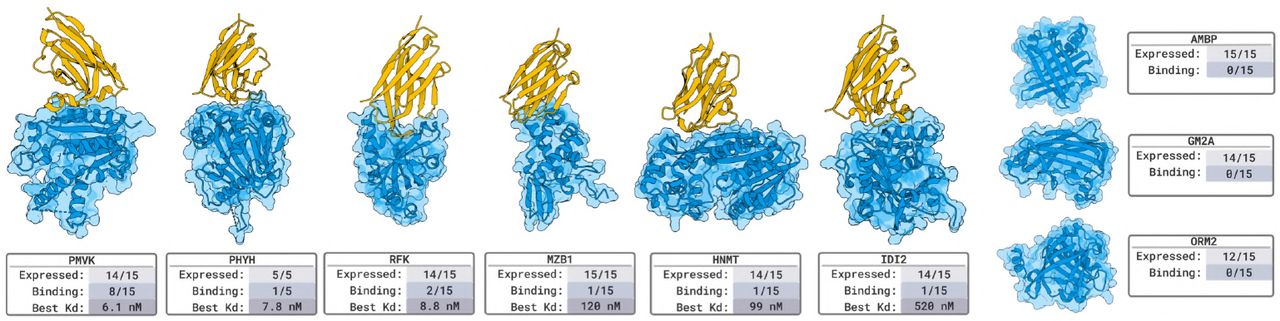
Figure 6: Nanobody binder validation. (a-i) BLI binding curves for designed nanobodies against 9 novel targets. Success on 6/9 targets (AMBP, PMVK, RFK, MZB1, CD22, HLA-A). Best affinity: 1.9 nM to AMBP. Each target tested with ~15 designs. Source: Stark et al. 2025, Figure 2.
Standard validation assays include:
- Bio-layer interferometry (BLI): High-throughput kinetic screening in 96-well format, measuring association/dissociation rates
- Surface plasmon resonance (SPR): Gold-standard sensitivity for affinity determination
- Isothermal titration calorimetry (ITC): Thermodynamic characterization requiring larger samples
- Structural validation: X-ray crystallography or cryo-EM confirming design accuracy
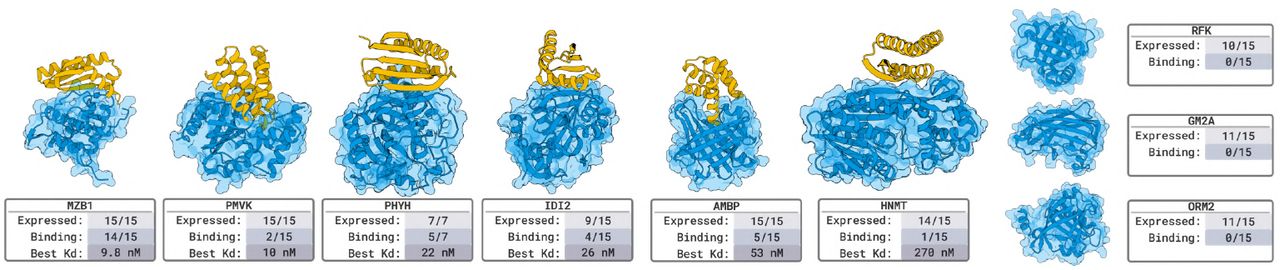
Figure 7: Miniprotein binder validation. Designed proteins (50-150 residues) against the same 9 novel targets. 6/9 target success rate. Highlights complementary success patterns to nanobodies, demonstrating modality-independent design capability. Source: Stark et al. 2025, Figure 3.
The Adaptyv Bio competition provides real-world benchmarking: Round 2 tested 400 proteins with 95% expression success but only 14% binding success (53/378 binders). Top designs matched or exceeded Cetuximab, the approved EGFR therapeutic. This gap between expression (~95%) and binding (~10-50%) represents the field’s central challenge—computational metrics predict foldability far better than functional binding.
Reported success rates span enormous ranges: BindCraft claims 10-100% depending on target, AlphaProteo achieves 5-88%, while pre-deep-learning Rosetta methods achieved less than 1%. This variability reflects target difficulty—charged polar interfaces and glycan-proximal sites remain particularly challenging.
Design specification language enables precise control

Figure 8: Design specification language (DSL) examples. (a) YAML syntax for cyclic peptide with disulfide bonds. (b) Helicon specification with backbone cyclization. (c) Binding site conditioning syntax. DSL enables precise control over covalent bonds, secondary structure, and interface residues. Source: Stark et al. 2025, Figure 9.
BoltzGen introduces a YAML-based design specification language (DSL) offering granular control over generation:
- Covalent bonds: Atom-level specification for cyclic peptides, disulfide staples, and helicons
- Binding site conditioning: Label residues as binding or non-binding to target or avoid specific regions
- Secondary structure constraints: Restrict designed residues to α-helices, β-sheets, or coils
- Structure groups: Control visibility and relative positioning with multi-chain assemblies
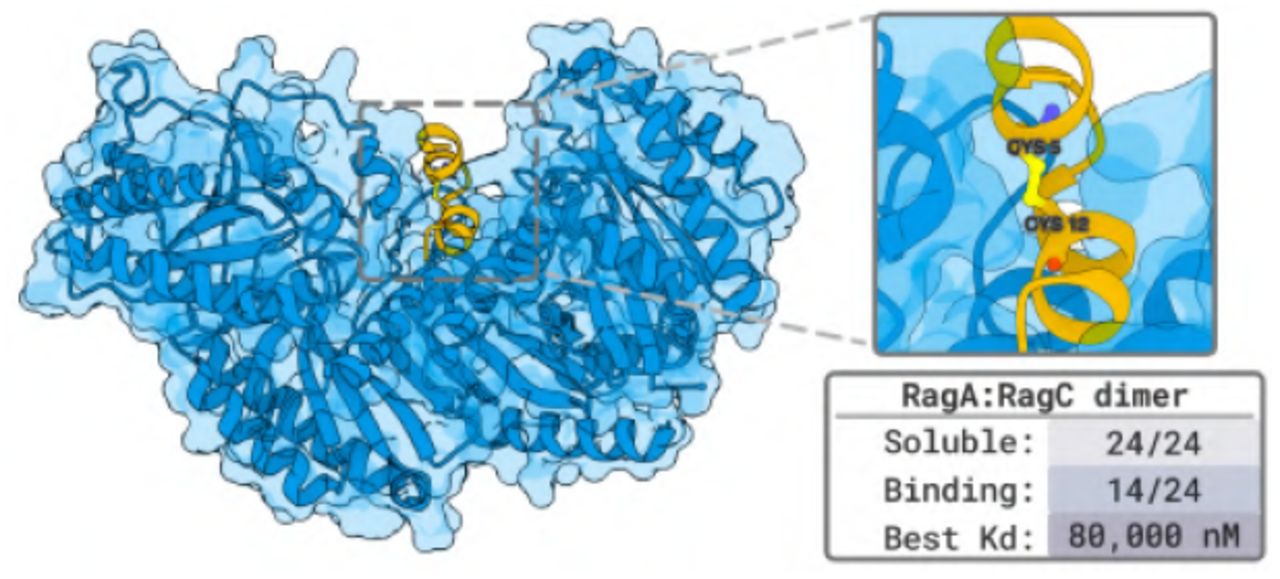
Figure 9: Disulfide-bonded cyclic peptide designs against RagA:RagC. 14/28 designs showed binding (50% success). Demonstrates DSL capability for constrained peptide geometries. Source: Stark et al. 2025.
This DSL supports multi-modal design across proteins (50-150 residues), peptides (8-30 residues), nanobodies, and cyclic peptides with various stapling chemistries. Binding site conditioning during training (30% of iterations, residues within 6Å of binder) enables pocket-directed generation while maintaining robustness to partial specification.
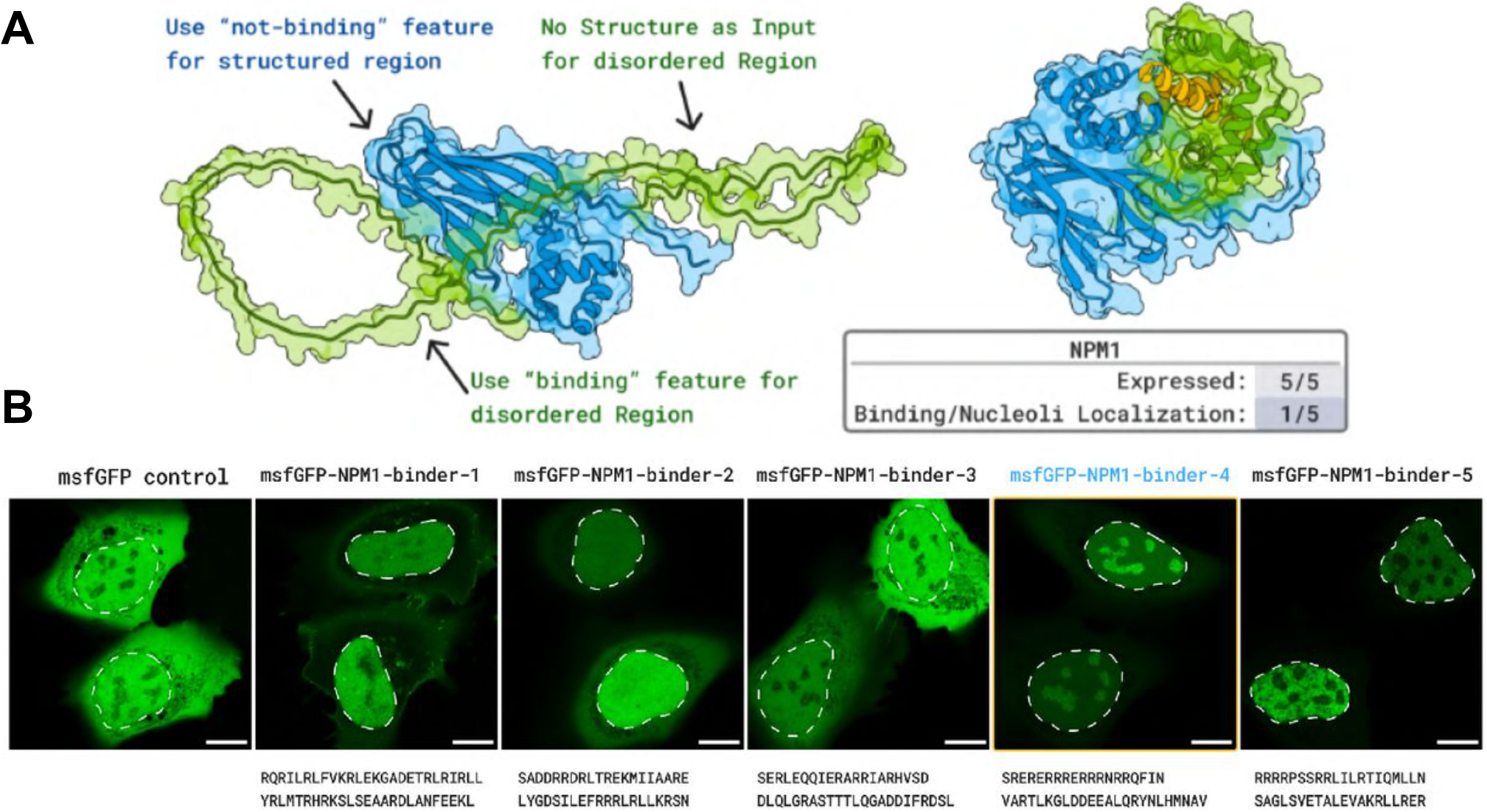
Figure 10: First de novo binder to intrinsically disordered protein in living cells. (a) Designed peptide targeting NPM1 C-terminal disordered region. (b) Confocal microscopy showing nucleolar localization in HeLa cells (Scale bar = 10 $\mathrm{\mu m}$). 1/5 designs showed specific enrichment. Validates BoltzGen on targets beyond structured proteins. Source: Stark et al. 2025, Figure S16.
A landmark validation demonstrated peptide binding to NPM1’s intrinsically disordered C-terminal region, a driver of acute myeloid leukemia. One of five tested designs showed nucleolar localization in live human cells—the first evidence of de novo designed proteins binding disordered targets in vivo.
Benchmarking metrics correlate imperfectly with experimental success
Computational filtering relies on structure prediction confidence metrics that imperfectly predict experimental outcomes. Primary metrics include pLDDT (per-residue confidence, threshold >80), ipTM (interface predicted TM-score, >0.5), and iPAE (interface predicted aligned error, <10Å). A 2025 meta-analysis of 3,766 binders found ipSAE—an AF3-derived metric—provides 1.4× improvement in average precision over iPAE.
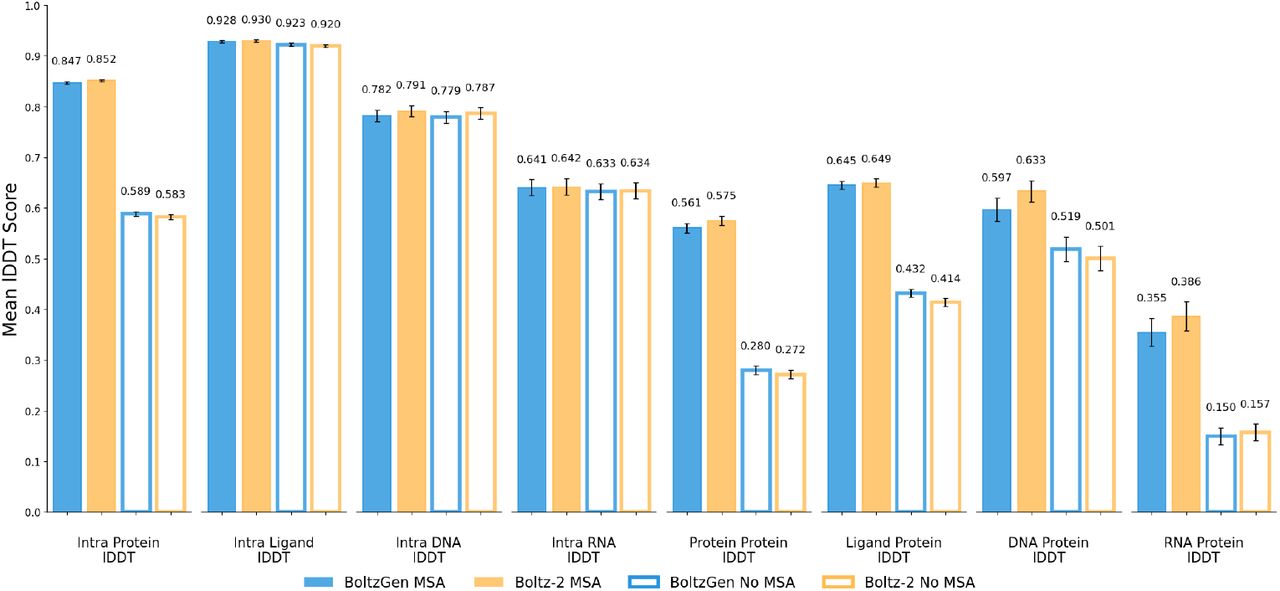
Figure 11: BoltzGen structure prediction benchmarks. (a) pLDDT distribution on the test set matches Boltz-2 baseline. (b) Interface metrics (ipTM, iPAE) for binder-target complexes. (c) Self-consistency scRMSD $\lt 2.0$Å for 80% of designs. Multi-task training preserves prediction accuracy while enabling generation. Source: Stark et al. 2025, Figure 20.
Self-consistency metrics validate that inverse-folded sequences recapitulate designed structures: scRMSD <2.0Å and scTM >0.5 using the best of 8 ProteinMPNN sequences refolded by AlphaFold2 or ESMFold. Critically, multiple predictors should be used—sequences optimized for AF2 confidence alone can create adversarial examples that fail with ESMFold.
Rosetta-based physical constraints remain valuable: shape complementarity >0.5, buried surface area >1, at least 6 interface residues, at least 2 hydrogen bonds, and favorable binding energy. The combination of folding model metrics with physics-based constraints outperforms either alone.
The fundamental limitation: no reliable metric correlates with binding affinity. Current approaches predict binary binding (yes/no) but not strength. Many designs pass computational filters yet fail experimentally; others achieve only micromolar affinity requiring experimental maturation for therapeutic use.
Testing on novel targets validates genuine generalization
BoltzGen’s emphasis on targets with less than 30% sequence identity to bound structures in PDB represents rigorous generalization testing. This threshold—the “twilight zone” of sequence-structure relationships—means homology modeling cannot infer binding modes. PDB clustering at 30% identity identifies unique protein folds, so designs against such targets demonstrate genuine novelty rather than pattern matching to training data.
Previous tools often validated on well-characterized therapeutic targets (PD-L1, TNFα, IL-7Rα) with numerous known binders. While useful for benchmarking, success on these targets may reflect similarity to training examples. BoltzGen’s 66% success on truly novel targets—including PMVK, AMBP, RFK, and MZB1—provides stronger evidence of generalizable design capability.
Conclusion
The protein binder design field has undergone remarkable transformation, from sub-1% success rates with Rosetta-era methods to 10-100% with modern deep learning approaches depending on target difficulty. BoltzGen’s unified architecture eliminates the structure-sequence information gap that limited two-stage pipelines, while its geometric residue encoding enables single-model training across prediction and design tasks. Key innovations—the design specification language, disordered protein targeting, and rigorous novel-target validation—position it as a significant advance. However, the gap between computational metrics and experimental outcomes persists: expression is largely solved (~95%), but binding success remains target-dependent (~10-66%), and affinity optimization still requires experimental maturation. The fully open-source ecosystem (BoltzGen, RFDiffusion, ProteinMPNN, BindCraft under MIT/BSD licenses) now rivals proprietary tools, democratizing access to capabilities that seemed impossible three years ago.
Main Reference:
Hannes Stark et al., BoltzGen: Toward Universal Binder Design, bioRxiv 2025.
Enjoy Reading This Article?
Here are some more articles you might like to read next: