Deep Learning for Protein-Ligand Binding
Are We Learning Physics or Memorizing the PDB?
The emergence of Boltz-2 claiming to approach free energy perturbation (FEP) performance marks a watershed moment for computational drug discovery—but skeptics are questioning whether these impressive benchmarks reflect genuine understanding of molecular recognition or sophisticated pattern-matching against memorized training data. The answer will determine whether deep learning can truly accelerate drug discovery or merely recapitulate what we already know.
This question sits at the heart of a growing debate in computational biology. A landmark 2025 study in Nature Machine Intelligence revealed that 49% of the widely-used CASF benchmark is contaminated with highly similar structures from the PDBbind training set (DOI: 10.1038/s42256-025-01124-5)—enabling models to achieve impressive scores through simple memorization rather than learned physics. Meanwhile, adversarial analyses of AlphaFold3 show that the model frequently places ligands in identical poses even when binding sites are mutated to impossibility, suggesting it has memorized poses rather than learned electrostatics.
Yet the story isn’t entirely bleak. Prospective validations—including the discovery of the novel antibiotic halicin—demonstrate that some models genuinely extrapolate to novel chemical space. Understanding when to trust these predictions is now the central challenge for practitioners.
The new generation of biomolecular foundation models
Boltz-2’s architectural innovations enable binding affinity prediction
Boltz-2, released by MIT and Recursion in June 2025, represents a significant architectural evolution from its predecessors. While AlphaFold3 and the original Boltz-1 focused on structure prediction, Boltz-2 introduces a dedicated affinity module that predicts binding strength alongside pose.

The architecture comprises four main components: a trunk with 64 PairFormer layers (up from 48 in Boltz-1) that builds rich pairwise representations, a diffusion-based denoising module for structure generation, a confidence module predicting reliability scores (pTM, ipTM, PAE), and the novel affinity module. This last component takes the trunk’s pair representation after five recycling iterations and processes it through 4-8 additional PairFormer layers focused exclusively on protein-ligand and intra-ligand interactions. Two prediction heads then output a binding likelihood (binary classifier) and an affinity value (regression to $\log_{10} \mathrm{IC_{50}}$ in μM).
Tokenization differs meaningfully from AlphaFold3: while AF3 atomizes non-canonical residues, Boltz-2 keeps them as single tokens—a design choice that preserves chemical identity. The model also introduces controllability features including method conditioning (specifying X-ray, NMR, or MD-derived structures), multimeric template support, and contact/pocket steering for physics-informed constraints.
Training leveraged ~5 million binding affinity measurements from ChEMBL, BindingDB, PubChem assays, and synthetic decoys. Crucially, the affinity module uses pairwise intra-assay differences during training—strongly weighting relative rankings within assays rather than absolute values—which may explain its lead optimization performance.
AlphaFold3’s diffusion approach enables ligand binding but not affinity
AlphaFold3, published in Nature in May 2024, marked DeepMind’s pivot from structure prediction to biomolecular interaction modeling. The key architectural innovation is replacing AlphaFold2’s structure module with a diffusion-based generator operating directly on raw atomic coordinates.
Where AF2 required torsion-based parameterization specifically designed for amino acid geometry, AF3’s diffusion approach naturally handles arbitrary chemical components—proteins, DNA, RNA, small molecules, ions, and modified residues—within a unified framework. The model receives “noised” atomic coordinates during training and learns to predict true coordinates; at inference, random noise is iteratively denoised to generate structures.
The Pairformer replaces AF2’s Evoformer, comprising 48 blocks operating on pair (n × n × 128) and single (n × 384) representations. A crucial simplification: MSA information is compressed into the pair representation after just 4 blocks and discarded, rather than maintained throughout processing. This design choice may contribute to both strengths (handling diverse molecular types) and weaknesses (reduced leverage of evolutionary information).
Critical limitation: AF3 predicts binding poses, not binding affinities. Its confidence metrics (pLDDT, ipTM, PAE) indicate structural reliability but do not correlate with binding strength. This fundamentally limits its direct utility for drug discovery ranking tasks.
How these models represent molecular interactions
Both models build on similar representation strategies but with important differences:
| Feature | AlphaFold3 | Boltz-2 |
|---|---|---|
| Protein input | Sequence → MSA → pair representation | Sequence → MSA → pair representation |
| Ligand input | SMILES → atomic tokens | SMILES → atomic tokens |
| Non-canonical handling | Atomic tokenization | Single token (preserves identity) |
| Core processing | 48 Pairformer blocks | 64 Pairformer blocks |
| Structure generation | Diffusion on raw coordinates | Diffusion with steering potentials |
| Affinity prediction | Not supported | Dedicated module |
| Template support | Single-chain only | Multi-chain templates |
Both architectures leverage ESM-style protein language model embeddings to capture evolutionary and structural information, but Boltz-2 explicitly incorporates ESM2 representations into its affinity module—a design choice the GEMS model paper demonstrated improves generalization on debiased benchmarks.
Performance benchmarks reveal promises and limitations
Boltz-2’s claim of approaching FEP performance
Boltz-2’s headline claim—approaching free energy perturbation performance at 1,000× lower computational cost—deserves careful scrutiny. On the FEP+ benchmark (4 targets: CDK2, TYK2, JNK1, P38), Boltz-2 achieves Pearson R = 0.66 versus FEP+’s 0.78 and ABFE’s 0.75. On the broader OpenFE benchmark, Boltz-2 achieves R = 0.62 versus OpenFE’s 0.63 and FEP+’s 0.72.
These results are impressive—but context matters. As computational chemist David Pearlman noted in his analysis: “For the OpenFE benchmark, it’s a move from R² = 0.52 to 0.38… For a chemist whose success depends on the results, this can easily be the difference between ‘intriguing, yes please’ and ‘sorry, next.’”
More convincingly, Boltz-2 achieved Pearson R = 0.65 on the CASP16 affinity challenge (140 protein-ligand pairs across 2 targets), outperforming all submitted competition entries (best competitor: 0.54). A prospective virtual screen against TYK2 found 8 of 10 top-scoring compounds validated as binders by ABFE simulations, with correlation R = 0.74 between Boltz-2 scores and ABFE predictions.
Structure prediction benchmarks show incremental progress
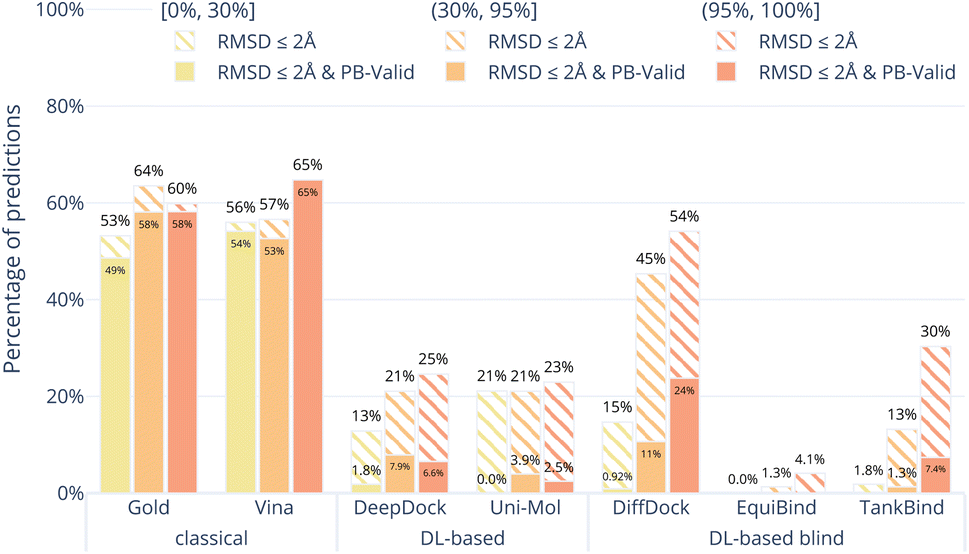
On the PoseBusters benchmark (308 complexes from 2021+), which evaluates both RMSD accuracy and physical/chemical validity, current methods perform as follows:
| Method | RMSD <2Å + PB-Valid | Key Issues |
|---|---|---|
| NeuralPLexer3 | 77.9% | Flow-based, physics-informed |
| AlphaFold3 | 76.4% (blind) | Chirality violations ~4.4% |
| SurfDock | ~65% | Volume overlap issues |
| DiffDock-L | ~50% | Best among DL docking |
| Vina | 59.7% | Consistent but slower |
| DiffDock | ~25% | Significant PB failures |
| EquiBind | <5% | Near-zero PB compliance |
Notably, over 50% of deep learning-generated poses fail basic physical validity checks including steric clashes, incorrect chirality, and impossible bond geometries. Post-processing with force field relaxation improves validity by 20-40 percentage points—suggesting these models capture approximate binding modes but not detailed chemistry.
The metrics matter enormously
Standard metrics can obscure important limitations:
- Pearson R measures linear correlation but ignores ranking quality and regression-to-mean effects
- RMSE (root mean square error) in pK units varies widely: FEP achieves ~0.8-1.2 kcal/mol, ML methods typically 1.2-2.0 kcal/mol
- Enrichment factors for virtual screening reveal practical utility better than correlation metrics
- PB-validity (physical validity) matters as much as RMSD for drug discovery applications
Boltz-2 demonstrates a concerning regression-to-mean tendency, predicting binding affinities within a narrow ~2 kcal/mol range across diverse targets. Community evaluations on 75 targets showed significant performance variance and struggles with conformationally flexible systems.
The generalization vs. memorization debate: what the evidence shows
Compelling evidence that standard benchmarks dramatically overestimate performance
The most damaging evidence comes from the PDBbind CleanSplit study. By comparing all CASF complexes with all PDBbind training complexes using structure-based clustering (TM scores for proteins, Tanimoto + RMSD for ligand poses), researchers identified that 49% of CASF benchmark complexes have highly similar structures in training data—sharing similar proteins, similar ligands, and similar binding poses.
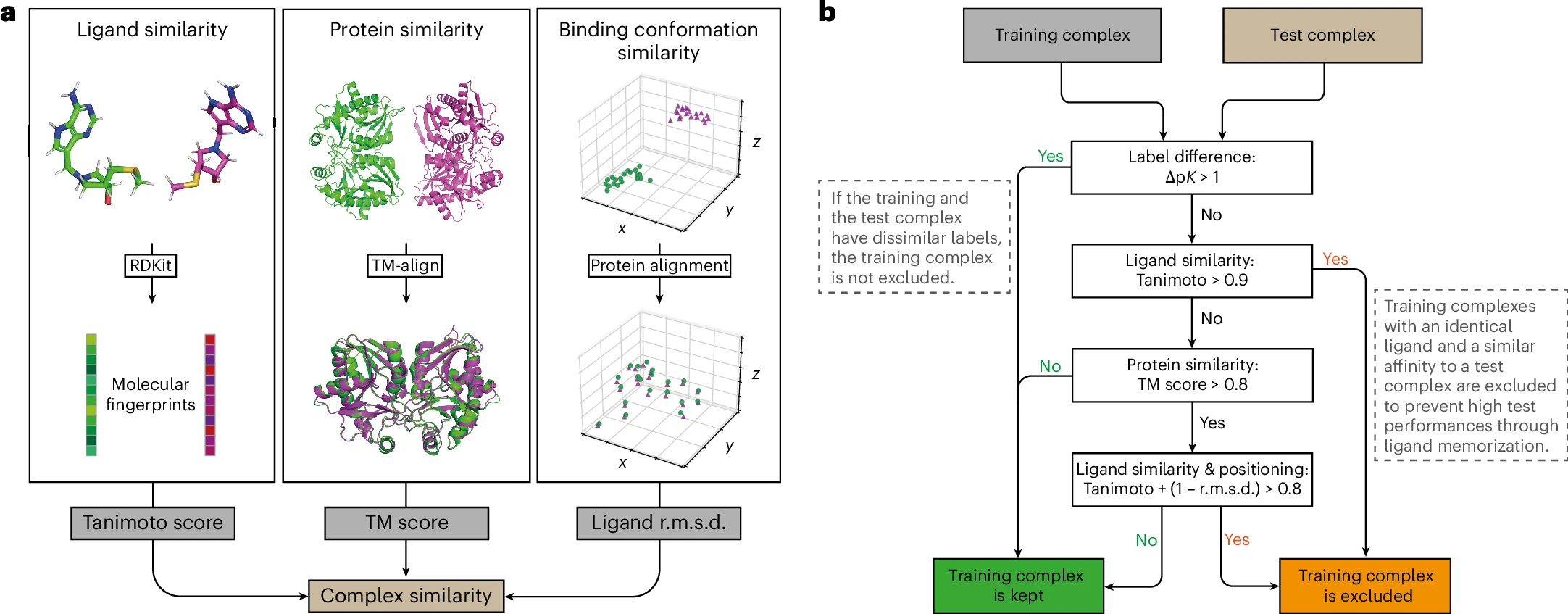
The implications are stark: when top-performing models were retrained on the leak-free CleanSplit dataset, their benchmark performance dropped substantially. Most alarmingly, some models achieved competitive CASF-2016 performance “even after removing all protein information from the input data“—demonstrating they had learned to predict affinities from ligand features alone, exploiting the fact that similar ligands often have similar affinities regardless of target.
Adversarial experiments reveal limited physics understanding
A 2025 study in Nature Communications (DOI: 10.1038/s41467-025-63947-5) subjected AlphaFold3 to adversarial stress tests:
- Binding site removal: All binding site residues mutated to glycine
- Pocket blocking: Binding sites filled with bulky phenylalanines
- Chemical inversion: Binding site residues changed to opposite charge/polarity
The result: “AlphaFold 3 frequently placed the ligand in the exact same pose as it did with the unmodified protein. In some cases, this led to the prediction of impossible structures with severe steric clashes.”
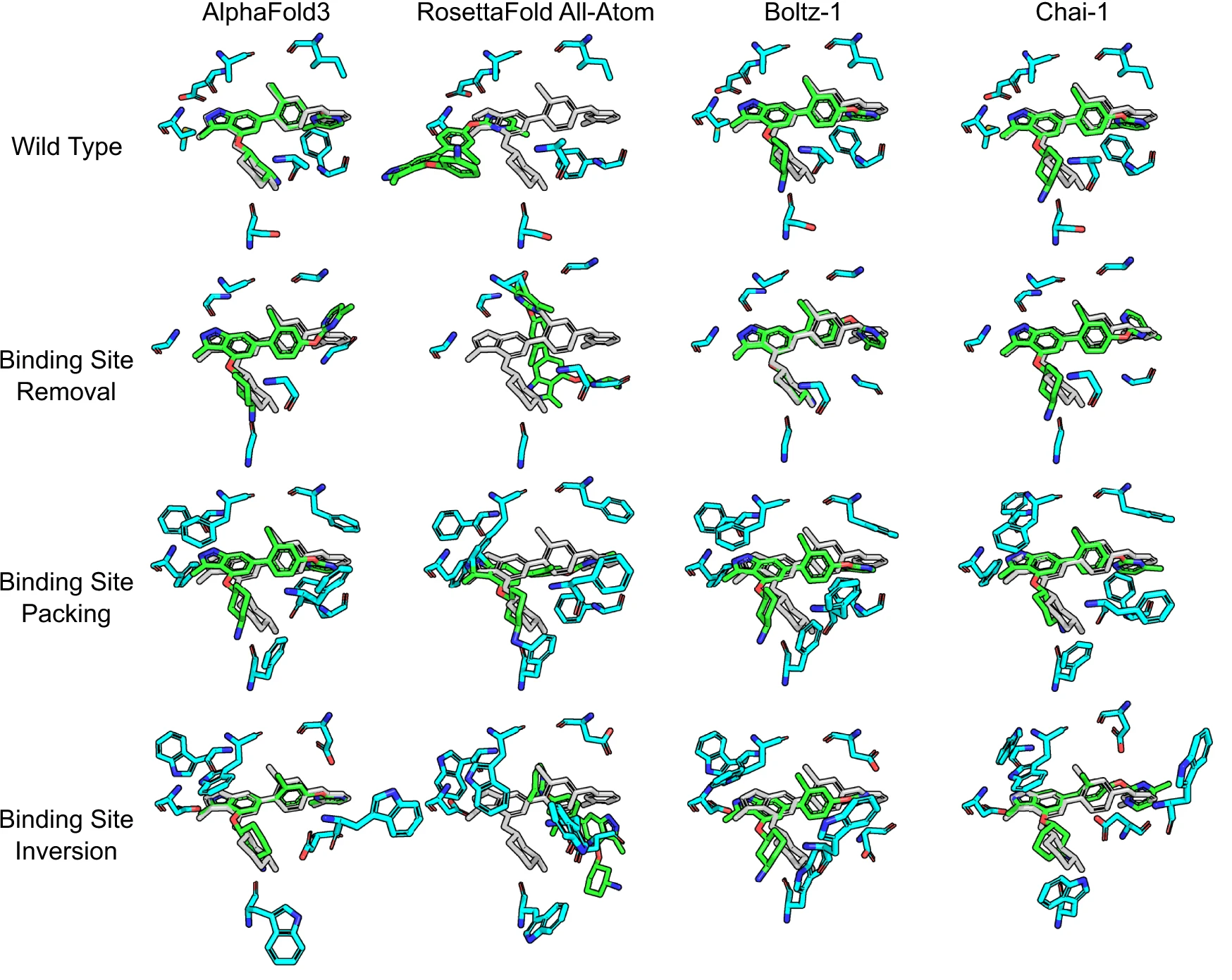
The conclusion was unambiguous: “These stress experiments show quite clearly that the AlphaFold 3 model did not learn any kind of physical principles such as electrostatics or van der Waals forces, and not even about clashes.”
Fold-switching proteins reveal memorization of structural templates
A Nature Communications study examining AlphaFold2’s predictions of fold-switching proteins found the model succeeded only 35% of the time for structures likely in its training set and just 1 of 7 times for experimentally-confirmed fold-switchers outside training. The authors documented that “AF2 ultimately predicted [one protein’s] helical conformation” even though coevolutionary information consistently indicated β-sheet structure—suggesting memorized templates override sequence-derived features.
Performance correlates with training set similarity
Multiple studies demonstrate that model performance correlates with similarity to training data:
- AlphaFold3 beats docking baselines on common ligands (appearing >100 times in training) but loses to docking on uncommon ligands (<100 occurrences)
- A kinase inhibitor study found models “memorize kinase phylogeny and match chemical analogues to make predictions instead of explicitly learning the factors driving high-affinity protein-ligand interactions”
- Temporal splits (training on pre-cutoff data, testing post-cutoff) consistently show 20-40% performance drops versus random splits
Counter-evidence: some models genuinely generalize
The picture isn’t entirely pessimistic. The GEMS model maintains strong performance (Pearson R = 0.803) on CASF-2016 even after removing all train-test leakage—suggesting that with proper architecture design and training data curation, genuine learning is possible. Key to GEMS’s success: sparse graph modeling of protein-ligand interactions and transfer learning from language models (ESM2, ChemBERTa-2).
The OpenFold retraining study (DOI: 10.1038/s41592-024-02272-z) found the model “remarkably robust at generalizing even when the size and diversity of its training set is deliberately limited, including near-complete elisions of classes of secondary structure elements.” Models trained primarily on α-helices could still predict β-sheet structures—evidence of transferable principles.
Most compellingly, prospective validations have succeeded. The discovery of halicin, a novel antibiotic structurally distinct from known antibiotics (Tanimoto similarity ~0.39 to nearest training compound), demonstrated that GNN-based models can identify genuinely novel active compounds (DOI: 10.1038/s41586-023-06887-8).
How researchers test for memorization versus generalization
Temporal splits remain the gold standard
Time-split cross-validation trains models on data before a cutoff date and tests on later data—mimicking real drug discovery scenarios where we predict future compounds. Studies consistently show temporal splits provide realistic performance estimates while random splits dramatically overestimate and even scaffold splits can overestimate.
Scaffold splits have limitations
Scaffold splitting—grouping molecules by shared core structure—is widely considered more rigorous than random splitting. However, a 2024 analysis demonstrated that scaffold splits also overestimate performance because “molecules with different chemical scaffolds are often similar” in descriptor space. The recommended alternative: UMAP-based clustering that considers full molecular similarity.
Structure-based filtering catches deeper leakage
The CleanSplit approach combines multiple criteria:
- TM score > 0.8 (protein structural similarity)
- Tanimoto + (1 - ligand RMSD) > 0.8 (ligand + pose similarity)
- Affinity labels within ±1 pK units
- Additional ligand identity filter (Tanimoto > 0.9)
This multi-pronged filtering catches cases where proteins and ligands appear dissimilar by sequence/scaffold but adopt highly similar binding geometries.
Ablation studies reveal what models actually learn
A particularly revealing test: remove protein information entirely and evaluate if models still perform well. On the standard CASF benchmark, several models maintained competitive performance using ligand features alone—impossible if they were learning protein-ligand interactions. On CleanSplit, protein information becomes essential, confirming the benchmark contamination hypothesis.
Practical implications: when can we trust these predictions?
High-confidence scenarios
Lead optimization within known chemotypes: When optimizing a series with substantial training data coverage and similar binding modes to known examples, models like Boltz-2 likely provide useful rankings. The strong intra-assay training signal makes relative predictions more reliable than absolute values.
Virtual screening for enrichment: Even imperfect models can dramatically improve hit rates in large-scale virtual screening. Boltz-2’s MF-PCBA results (enrichment factor 18.4 at 0.5% versus ~2-3 for docking) suggest meaningful utility for hit-finding, even if some success derives from memorization.
Well-characterized targets with many PDB structures: Targets extensively represented in training data (kinases, proteases, common enzyme families) should see better predictions—precisely because memorization works when test cases resemble training data.
High-risk scenarios
Novel targets with limited homologs: For truly novel protein folds or binding sites dissimilar to training data, predictions may be unreliable. The adversarial mutation experiments suggest models won’t adapt appropriately to unprecedented binding environments.
Scaffold hops to dissimilar chemotypes: Predictions for novel scaffolds structurally dissimilar to training compounds show consistently degraded performance. The ~40% performance drop on uncommon ligands in AF3 benchmarks should give pause.
Quantitative affinity predictions for decision-making: The regression-to-mean tendency means Boltz-2 may not reliably distinguish tight binders (nM) from weak ones (μM). For go/no-go decisions based on potency thresholds, FEP or experimental validation remains essential.
Flexible proteins and conformational selection: Both AlphaFold3 and Boltz-2 struggle with targets requiring large conformational changes. DynamicBind and NeuralPLexer attempt to address this but performance remains inconsistent.
A decision framework for practitioners
| Scenario | Recommended Approach |
|---|---|
| Large library screening | ML models (accept memorization benefits) |
| Lead optimization, known series | Boltz-2 or hybrid ML+FEP |
| Small modifications, quantitative ranking | FEP+ |
| Novel scaffolds, high stakes | FEP + experimental validation |
| Novel targets, structural exploration | AF3/Boltz-2 with low confidence threshold |
Uncertainty quantification is essential but underutilized
Recent work demonstrates that Bayes by Backprop provides well-calibrated uncertainty estimates for binding affinity predictions, distinguishing cases where models are confident from those where predictions are unreliable. Incorporating uncertainty quantification into production workflows would help practitioners identify when to trust predictions versus seek experimental validation.
The path forward: building models that learn physics
Architectural approaches that may improve generalization
The GEMS model’s success on CleanSplit suggests that sparse graph representations of protein-ligand interactions—rather than dense voxel grids or fully-connected attention—may better capture transferable interaction patterns. Integration of protein language model embeddings (ESM2) and chemical language models (ChemBERTa-2) provides rich pre-trained representations that encode general molecular properties.
Physics-informed constraints during training or inference (Boltz-2’s steering potentials, NeuralPLexer3’s optimal transport corrections) may help models respect physical plausibility rather than simply interpolating memorized poses.
Data strategies matter as much as architecture
The CleanSplit work demonstrates that training data curation dramatically affects generalization. Future benchmarks should adopt structure-based filtering as standard practice. Meanwhile, data augmentation using physics-based methods—generating synthetic binding data via docking or FEP—can expand training coverage into underrepresented regions of chemical space.
The need for honest benchmarking
The field needs standardized protocols that:
- Use temporal splits with enforced training cutoffs
- Apply structure-based filtering to prevent pose memorization
- Report ablation studies (performance with protein/ligand information removed)
- Include prospective validation as the ultimate test
Until benchmarking practices improve, published performance numbers should be treated skeptically.
Conclusion
The generalization versus memorization debate reveals a fundamental tension in applying deep learning to drug discovery. Current evidence suggests that standard benchmarks dramatically overestimate model capabilities, with up to half of test examples solvable through memorization alone. Adversarial experiments demonstrate that even state-of-the-art models like AlphaFold3 have not learned basic physics of molecular recognition.
Yet the story isn’t one of failure. Prospective validations demonstrate that well-designed models can discover genuinely novel compounds. The GEMS model maintains performance on rigorously debiased benchmarks. And even imperfect models offer tremendous value for high-throughput screening, where ranking many compounds approximately beats ranking few compounds precisely.

For practitioners, the key insight is contextual trust: these tools work best when predictions resemble training data (lead optimization within known series) and worst when extrapolating to unprecedented chemistry or biology. The 1,000× speed advantage over FEP creates clear value even with lower accuracy—but only if users understand the reliability boundaries.
The path forward requires honest benchmarking, physics-informed architectures, and careful curation of training data to encourage genuine learning over sophisticated memorization. The ultimate test remains prospective validation: can these models discover drugs that work in patients? That question remains open, but the tools to answer it are finally emerging.
References
-
Passaro S, Corso G, Wohlwend J, et al. Boltz-2: Towards Accurate and Efficient Binding Affinity Prediction. bioRxiv 2025.06.14.659707. DOI: 10.1101/2025.06.14.659707
-
Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024;630:493-500. DOI: 10.1038/s41586-024-07487-w
-
Graber D, Stockinger P, Meyer F, et al. Resolving data bias improves generalization in binding affinity prediction. Nature Machine Intelligence 2025;7:1713-1725. DOI: 10.1038/s42256-025-01124-5
-
Chakravarty D, et al. AlphaFold predictions of fold-switched conformations are driven by structure memorization. Nature Communications 2024;15:7296. DOI: 10.1038/s41467-024-51801-z
-
Investigating whether deep learning models for co-folding learn the physics of protein-ligand interactions. Nature Communications 2025. DOI: 10.1038/s41467-025-63947-5
-
Ahdritz G, et al. OpenFold: retraining AlphaFold2 yields new insights into its learning mechanisms and capacity for generalization. Nature Methods 2024;21:1514-1524. DOI: 10.1038/s41592-024-02272-z
-
Buttenschoen M, Morris GM, Deane CM. PoseBusters: AI-based docking methods fail to generate physically valid poses or generalise to novel sequences. Chemical Science 2024;15:3130-3139. DOI: 10.1039/d3sc04185a
-
Wong F, et al. Discovery of a structural class of antibiotics with explainable deep learning. Nature 2024;626:177-185. DOI: 10.1038/s41586-023-06887-8
-
Lu W, et al. DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model. Nature Communications 2024;15:1071. DOI: 10.1038/s41467-024-45461-2
-
Qiao Z, et al. State-specific protein-ligand complex structure prediction with a multiscale deep generative model. Nature Machine Intelligence 2024;6:195-208. DOI: 10.1038/s42256-024-00792-z
-
Cao D, et al. SurfDock is a surface-informed diffusion generative model for reliable and accurate protein-ligand complex prediction. Nature Methods 2025;22:310-322. DOI: 10.1038/s41592-024-02516-y
-
Corso G, et al. DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking. ICLR 2023. arXiv:2210.01776
-
Valsson O, et al. Narrowing the gap between machine learning scoring functions and free energy perturbation using augmented training data. Communications Chemistry 2025. DOI: 10.1038/s42004-025-01428-y
-
Ross GA, et al. Maximal and current accuracy of rigorous protein-ligand binding free energy calculations. Communications Chemistry 2023. DOI: 10.1038/s42004-023-01019-9
-
Li J, et al. Leak Proof PDBBind: A Reorganized Dataset of Protein-Ligand Complexes for More Generalizable Binding Affinity Prediction. arXiv:2308.09639v2
-
Guo Q, et al. Scaffold Splits Overestimate Virtual Screening Performance. arXiv:2406.00873 (2024)
-
David Pearlman, Boltz-2 vs. FEP? The Wrong Question. Here’s Why They’re Stronger Together, Medium.
Enjoy Reading This Article?
Here are some more articles you might like to read next: