Deep Learning for Computational Structural Biology
A Roadmap for Enthusiasts
The transition from traditional computational biology to AI-driven structural biology requires mastering four pillars: (1) foundational machine learning, (2) deep learning architectures specific to 3D molecular structures (transformers, diffusion models, equivariant neural networks, GNNs), (3) hands-on experience with landmark tools (AlphaFold, ESMFold, RFDiffusion, ProteinMPNN), and (4) engagement with a rapidly evolving research community. This roadmap provides a 16-24 week structured pathway from ML fundamentals to implementing state-of-the-art protein structure prediction and design—prioritizing free, actively maintained resources vetted by the computational biology community.
The field underwent revolutionary change when AlphaFold2 achieved near-experimental accuracy at CASP14 in 2020, earning its creators the 2024 Nobel Prize in Chemistry. Since then, the landscape has expanded dramatically: ESMFold demonstrates that protein language models alone can predict structures without multiple sequence alignments, RFDiffusion enables de novo protein design through diffusion models, and AlphaFold3/Boltz2 extends predictions to protein-ligand-nucleic acid complexes. Graduate students entering this field now have unprecedented opportunities—but also face the challenge of synthesizing knowledge across machine learning, structural biology, and software engineering.
“A journey of a thousand miles begins with a single step.“ – Lao Tzu
The complete learning pathway spans four phases
The roadmap divides into distinct phases, each building systematically on the previous. Phase 1 (Weeks 1-6) establishes ML/DL fundamentals through courses, hands-on coding, and mathematical foundations. Phase 2 (Weeks 7-12) introduces the specialized architectures—transformers, diffusion models, equivariant networks, and GNNs—that underpin modern structural biology AI. Phase 3 (Weeks 13-18) focuses on mastering specific tools: running AlphaFold predictions, designing proteins with ProteinMPNN and RFDiffusion, and extracting embeddings from protein language models. Phase 4 (Weeks 19-24+) transitions to independent research through projects, paper reading, and community engagement.
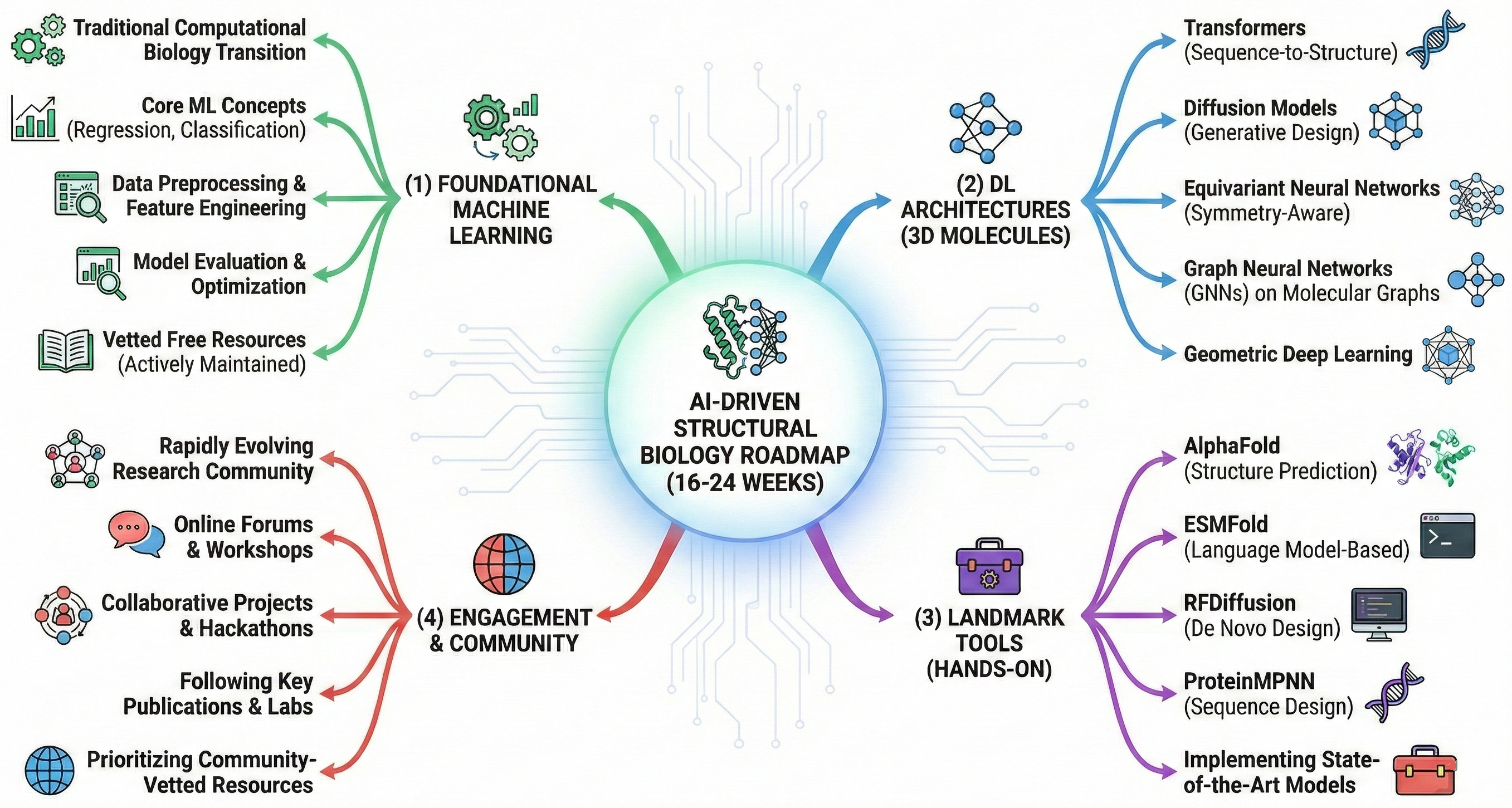
Prerequisites vary by phase. Phase 1 requires only Python programming proficiency (one year experience), undergraduate-level calculus, and basic linear algebra. No prior biology knowledge is strictly necessary, though familiarity with protein structure basics accelerates progress. By Phase 3, learners should be comfortable with PyTorch tensor operations, loss functions, and optimization, while Phase 4 assumes the ability to read and implement methods from research papers.
Time estimates assume 15-20 hours per week of dedicated study. Full-time learners (40+ hours/week) can compress the timeline to 10-12 weeks. The path accommodates different learning styles: visual learners benefit from 3Blue1Brown and lecture-based courses, while hands-on learners should prioritize Colab notebooks and coding exercises throughout.
Phase 1: ML and deep learning foundations
Core machine learning concepts (Weeks 1-2)
Begin with MIT 6.S191: Introduction to Deep Learning, a one-week intensive course covering neural network fundamentals, CNNs, RNNs, transformers, and generative AI. The course uniquely includes biological applications and is taught by researchers with computational biology backgrounds (e.g., Ava Amini, a researcher at Microsoft Research). All materials—lectures, slides, and TensorFlow/Google Colab labs—are freely available on the course website and GitHub.
For deeper mathematical foundations, Stanford CS229 provides rigorous coverage of supervised and unsupervised learning, optimization, and regularization. The 2018 lecture videos featuring Andrew Ng remain excellent (YouTube playlist). Expect approximately 30-40 hours across both courses.
Visual learners should start with 3Blue1Brown’s Neural Networks series, which provides unparalleled intuition for backpropagation and gradient descent in approximately 4 hours. The recent chapters on attention and transformers directly prepare learners for protein language models.
Deep learning frameworks (Weeks 3-4)
PyTorch is the dominant framework for structural biology AI—ESMFold, OpenFold, RoseTTAFold, ProteinMPNN, and RFDiffusion all use it. Begin with the official PyTorch tutorials, starting with the 60-Minute Blitz covering tensors, autograd, and basic neural networks. The “Learn the Basics” tutorial series then builds complete training workflows. The Zero to Mastery: Learn PyTorch for Deep Learning course offers comprehensive free video-based training.
JAX is essential for AlphaFold work—both AlphaFold2 and AlphaFold3 are written in JAX. After gaining PyTorch fluency, spend one week on JAX fundamentals via the official tutorials and the excellent UvA (University of Amsterdam) JAX introduction. Focus on JIT compilation, automatic differentiation (grad), vectorization (vmap), and the functional programming paradigm that differs from PyTorch’s imperative style.
Practical coding and hands-on practice (Weeks 5-6)
Kaggle Learn offers free micro-courses in Python, Pandas, data visualization, and intro ML/DL—each taking approximately 4 hours. Complete at minimum Intro to Machine Learning and Intermediate Machine Learning.
Andrej Karpathy’s Neural Networks: Zero to Hero represents the gold standard for building deep understanding. Across ~25 hours of video, Karpathy builds neural networks from scratch, culminating in a GPT implementation (nanoGPT). The transformer understanding gained here directly transfers to comprehending ESM, AlphaFold’s attention mechanisms, and all protein language models (GitHub repository and YouTube playlist).
Recommended books
“Dive into Deep Learning” (d2l.ai) stands out as the most practical resource—completely free, interactive with runnable code, and available in PyTorch, JAX, and TensorFlow implementations. The ~900-page book covers everything from linear networks to transformers, with excellent chapters on attention mechanisms (Chapter 11) directly relevant to protein modeling.
“Deep Learning” by Goodfellow, Bengio, and Courville serves as the theoretical reference. The free HTML version covers mathematical foundations comprehensively; use it as a reference rather than reading cover-to-cover.
Phase 2: Advanced architectures for structural biology
Transformers and attention mechanisms (Week 7)
Understanding transformers is non-negotiable—they power AlphaFold’s Evoformer, ESM protein language models, and virtually all modern sequence-to-structure approaches. The key insight: transformers treat amino acid sequences as token sequences, with self-attention capturing long-range dependencies between residues critical for protein folding.
Begin with “The Illustrated Transformer“ by Jay Alammar , featured in courses at Stanford, MIT, and Harvard. Follow with “The Annotated Transformer“ from Harvard NLP, providing line-by-line PyTorch implementation. 3Blue1Brown’s attention visualization solidifies intuition.
Stanford CS224n: NLP with Deep Learning provides comprehensive coverage of transformers, attention, BERT, and GPT—all directly applicable to protein language models (2021 YouTube playlist). This course is particularly valuable because protein language models (ESM, ProtTrans) use nearly identical architectures to NLP transformers.
The foundational paper, “Attention Is All You Need” (arXiv:1706.03762), should be read after gaining intuition from the tutorials.
Diffusion models (Week 8)
Diffusion models underpin RFDiffusion (protein design), AlphaFold3’s structure module, and emerging protein generation methods. The core concept: learn to reverse a gradual noising process to generate new samples.
Lilian Weng’s “What are Diffusion Models?“ provides the definitive introduction, covering DDPM, score matching, and classifier guidance in a comprehensive 32-minute read. Yang Song’s “Generative Modeling by Estimating Gradients” explains score-based approaches from first principles.
For implementation, Yang Song’s official PyTorch code offers tutorials on sampling, likelihood computation, and controllable synthesis. The foundational paper is “Denoising Diffusion Probabilistic Models” by Ho et al. (arXiv:2006.11239).
The application to proteins—RFDiffusion—demonstrates how fine-tuning structure prediction networks (RoseTTAFold) for denoising enables generation of novel protein backbones. The Nature 2023 paper with GitHub repo should be studied after understanding diffusion fundamentals.
Equivariant neural networks (Weeks 9-10)
SE(3) and E(3) equivariance is perhaps the most conceptually challenging topic—but essential for understanding why modern architectures handle 3D structures so effectively. The key insight: physical properties of molecules (energy, forces, binding affinity) don’t depend on how the molecule is rotated or translated in space. Equivariant networks encode this symmetry directly, dramatically improving data efficiency.
Start with the “Geometric Deep Learning” proto-book by Bronstein, Bruna, Cohen, and Veličković (arXiv:2104.13478). This comprehensive resource unifies CNNs, GNNs, and transformers through the lens of symmetry and group theory. The accompanying lecture series from AMMI makes the mathematical concepts accessible.
For implementation, the e3nn library provides E(3)-equivariant neural network operations in PyTorch and JAX. The tutorials at e3nn_tutorial cover geometric tensors, spherical harmonics, and equivariant convolutions. The TeachOpenCADD tutorial T036 specifically explains why equivariance matters for molecular modeling.
Key architectures to understand include NequIP (Nature Communications 2022), which demonstrated that equivariant networks achieve state-of-the-art accuracy with 3 orders of magnitude fewer training samples than conventional approaches.
Graph neural networks (Weeks 11-12)
Proteins and molecules are naturally represented as graphs—residue graphs (nodes = amino acids, edges = spatial proximity or sequence connectivity) or atom graphs (nodes = atoms, edges = covalent bonds). GNNs process these representations through message passing.
PyTorch Geometric (PyG) is the dominant library. Start with the “Introduction by Example” tutorial , then the UvA Deep Learning Tutorial 7 on GNNs, which includes molecular property prediction examples.
The Deep Graph Library (DGL) offers an alternative with DGL-LifeSci (ACS Omega 2021) specifically for molecular property prediction—7 models for property prediction with a command-line interface requiring no coding.
For converting molecular representations, the Oxford BLOPIG tutorial explains SMILES to graph conversion using RDKit.
Phase 3: Mastering structural biology AI tools
Protein structure prediction (Week 13-14)
ColabFold democratized AlphaFold2 access, providing 40-60x faster predictions through MMseqs2-based MSA generation. The main Colab notebook runs on Google Colab’s free tier (T4 GPU) for sequences up to ~1,000-1,400 residues. The Nature Methods paper and 2024 Nature Protocols paper detail best practices.
ESMFold offers even faster predictions (~60x faster than AlphaFold2) without requiring MSAs. The Colab notebook predicts a 252-residue protein in ~21 seconds. ESMFold accuracy is slightly below AlphaFold2 but sufficient for many applications—particularly when speed matters or MSAs are unavailable (e.g., orphan proteins, metagenomic sequences).
The EMBL-EBI AlphaFold training course, co-developed with DeepMind, provides authoritative guidance on interpreting predictions, understanding limitations, and accessing the AlphaFold Database containing 200+ million predicted structures.
For local installation with full control, OpenFold provides a trainable PyTorch reimplementation matching AlphaFold2 accuracy. The 2024 Nature Methods paper includes released weights, training code, and ~400K MSAs via OpenProteinSet on AWS Open Data.
AlphaFold3 (Nature 2024) extends to proteins, DNA, RNA, small molecules, and ions using a diffusion-based architecture. Inference code was released November 2024; weights require application to DeepMind under CC-BY-NC-SA 4.0 (non-commercial). Open alternatives—Boltz-1 (MIT license) and Chai-1—provide comparable accuracy with more permissive licensing.
Protein design tools (Weeks 15-16)
ProteinMPNN performs inverse folding—designing amino acid sequences that fold into target backbone structures. The Science 2022 paper achieved 52.4% sequence recovery versus 32.9% for Rosetta. Colab notebooks include a quick demo and a version with AlphaFold2 validation. The Meiler Lab tutorial provides practical guidance on redesigning protein interfaces.
LigandMPNN (Nature Methods 2025) extends ProteinMPNN to explicitly model small molecules, nucleotides, and metals—achieving 63.3% sequence recovery for small molecule interactions versus 50% for ProteinMPNN.
RFDiffusion generates novel protein backbones through diffusion, enabling binder design, symmetric assembly creation, and enzyme active site scaffolding. Colab notebooks are available via ColabDesign. The typical workflow chains RFDiffusion (backbone generation) → ProteinMPNN (sequence design) → AlphaFold2 (validation). The Data-Driven Life Sciences (DDLS) course notebook provides hands-on exercises.
Chroma (Nature 2023) from Generate:Biomedicines offers programmable protein generation with composable constraints for symmetry, shape, and substructure.
Protein language models (Week 17)
ESM-2 from Meta FAIR provides protein embeddings for downstream ML tasks. Models range from 8M to 15B parameters; the 650M parameter model balances performance and computational requirements. Use cases include extracting sequence embeddings for function prediction, variant effect prediction (ESM-1v), contact prediction, and zero-shot fitness scoring. HuggingFace integration enables easy model loading.
ProtTrans offers multiple architectures (ProtBERT, ProtT5, ProtAlbert, ProtXLNet), all available on HuggingFace. ProtT5 is generally recommended for best performance.
RNA and ligand prediction (Week 18)
RhoFold+ (Nature Methods 2024) predicts RNA 3D structures using an RNA language model, achieving top performance on CASP15 natural RNA targets with ~0.14 second inference on A100 (Online server).
DiffDock (ICLR 2023) applies diffusion models to molecular docking, achieving 38% top-1 success rate within 2Å RMSD versus 23% for traditional methods like Glide.
Essential academic papers organized by reading order
Foundation reading list (Weeks 1-4)
The first paper to read is “Attention Is All You Need” (arXiv:1706.03762)—understanding transformers is prerequisite to everything that follows. Next, “Denoising Diffusion Probabilistic Models” (arXiv:2006.11239) establishes diffusion fundamentals. Then “Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences” (ESM-1b; PNAS 2021) demonstrates that transformer pre-training on protein sequences yields emergent structure information.
Core structural biology AI (Weeks 5-8)
AlphaFold2 (Nature 2021) is the landmark paper—read it carefully, studying the Evoformer architecture, MSA processing, and invariant point attention. Follow with ESMFold (Science 2023) to understand how language models alone can predict structures. Then read RFDiffusion (Nature 2023) for diffusion-based design.
Current frontiers (Weeks 9-12)
AlphaFold3 (Nature 2024) represents the current state-of-the-art for biomolecular complexes. Boltz-1(2) (bioRxiv 2024, bioRxiv 2025) and Chai-1(2) (bioRxiv 2024, bioRxiv 2025) provide open alternatives. ESM-3 (bioRxiv 2024, Science 2025) demonstrates multimodal sequence-structure-function modeling.
Review papers for context
“AI-Driven Deep Learning Techniques in Protein Structure Prediction” (Int J Mol Sci 2024) provides comprehensive overview for newcomers. “Emerging frontiers in protein structure prediction following the AlphaFold revolution” (J R Soc Interface 2025) covers post-AlphaFold developments including conformational ensembles and remaining challenges.
Key research groups shaping the field
DeepMind’s AlphaFold team (London)—led by 2024 Nobel laureates Demis Hassabis and John Jumper—created AlphaFold and the AlphaFold Database. Meta FAIR / EvolutionaryScale (New York)—with Alexander Rives, Tom Sercu, and Zeming Lin—developed the ESM series and ESM Metagenomic Atlas with 600M+ structures. The group spun out as EvolutionaryScale in 2023. David Baker’s Institute for Protein Design (University of Washington, Seattle)—with Minkyung Baek and Joseph Watson—pioneered computational protein design, created Rosetta, RoseTTAFold, RFDiffusion, and ProteinMPNN. Baker won the 2024 Nobel Prize alongside Hassabis and Jumper. MIT’s Jameel Clinic—with Regina Barzilay and Tommi Jaakkola—developed Boltz-1(2) and DiffDock with emphasis on open-source accessibility.
Computing requirements and practical setup
For beginners, Google Colab’s free tier (T4 GPU, 16GB VRAM) handles ColabFold predictions up to ~1,000-1,400 residues, ESMFold up to ~900 residues, and ProteinMPNN design. This suffices for learning and small-scale projects.
For serious work, a local GPU with 24GB VRAM (RTX 3090/4090, ~$1,500-2,000) enables most inference tasks. Optimal setup includes 80GB A100 access for long sequences, AlphaFold3, and training—available via Google Cloud (a2-ultragpu-1g instances), AWS, or academic clusters.
Academic users can access COSMIC2 for free AlphaFold/ColabFold; NIH Biowulf; or XSEDE/ACCESS allocations. Many institutions provide JupyterHub environments with GPU access.
Full AlphaFold2 installation requires 1-1.25 TB SSD for sequence databases. ColabFold avoids this by using remote MSA servers. ESMFold requires no databases at all.
Community engagement and staying current
Key conferences and workshops
MLSB (Machine Learning in Structural Biology) at NeurIPS (December annually) is the premier venue—non-archival, emphasizing open-source implementations. MLCB (Machine Learning in Computational Biology) (September) hosts 1000+ participants with rigorous peer review and PMLR proceedings. CASP (Critical Assessment of Structure Prediction) (biennial) provides blind evaluation that drove AlphaFold development—though currently facing funding challenges.
Essential online resources
Set up bioRxiv alerts for keywords: “protein structure prediction,” “AlphaFold,” “protein design,” “deep learning.” Follow arXiv cs.LG and q-bio categories. Subscribe to newsletters: Sebastian Raschka’s “Ahead of AI“ for technical deep learning; TheSequence for ML/AI updates.
MLCB YouTube hosts all conference talks. Stanford MLSys Seminars regularly feature structural biology topics. The ColabDesign Discord provides community support for protein design tools.
Reddit communities r/bioinformatics (~180K members) and r/MachineLearning (~3M members) offer active Q&A. BioStars remains the major bioinformatics forum.
Project-based learning progression
Beginner projects (After Phase 2)
- Predict and validate a well-characterized protein: Use ColabFold on a protein with known experimental structure (e.g., lysozyme, PDB: 1LYZ), compare pLDDT scores and RMSD to experiment using PyMOL
- Redesign a miniprotein: Apply ProteinMPNN to 1QYS (56-residue three-helix bundle), validate designed sequences with AlphaFold—targeting >70% predicted structure recovery
- Extract ESM embeddings for function prediction: Use ESM-2 to embed a set of enzymes, train a simple classifier to predict enzyme class
Intermediate challenges (After Phase 3)
- Predict a protein complex: Use AlphaFold-Multimer on a known heterodimer, analyze interface contacts
- Design a binder: Use RFDiffusion to design binders for a target of interest (e.g., viral spike protein epitope)
- Conformation sampling: Generate multiple conformations using ColabFold seed sampling for a flexible protein
Kaggle competitions
CAFA 5 Protein Function Prediction challenges participants to predict GO terms from sequences, typically using ESM-2 embeddings and AlphaFold structural features. Past Novozymes Enzyme Stability competition used ESM embeddings with AlphaFold structures for thermostability prediction.
Advanced directions
Participate in CASP when the next experiment opens (monitor predictioncenter.org). Reproduce methods from recent papers—implementing a small component of AlphaFold2 or RFDiffusion builds deep understanding. Contribute to open-source projects: OpenFold, ColabFold, and ESM welcome contributions.
Complete resource index by category
Courses (Free)
- MIT 6.S191: https://introtodeeplearning.com
- Stanford CS229: https://cs229.stanford.edu (YouTube videos)
- Stanford CS224n: https://web.stanford.edu/class/cs224n
- Fast.ai: https://course.fast.ai
- EMBL-EBI AlphaFold: https://www.ebi.ac.uk/training/online/courses/alphafold
Framework tutorials
- PyTorch: https://docs.pytorch.org/tutorials
- JAX: https://docs.jax.dev/en/latest/tutorials.html
- PyTorch Geometric: https://pytorch-geometric.readthedocs.io
Books (Free online)
- Dive into Deep Learning: https://d2l.ai
- Deep Learning (Goodfellow): https://www.deeplearningbook.org
- Geometric Deep Learning: https://geometricdeeplearning.com
Key GitHub repositories
- ColabFold: https://github.com/sokrypton/ColabFold
- ColabDesign: https://github.com/sokrypton/ColabDesign
- OpenFold: https://github.com/aqlaboratory/openfold
- ESM: https://github.com/facebookresearch/esm
- ProteinMPNN: https://github.com/dauparas/ProteinMPNN
- RFDiffusion: https://github.com/RosettaCommons/RFdiffusion
- Boltz-1(2): https://github.com/boltz-bio/boltz
- e3nn: https://github.com/e3nn/e3nn
Video resources
- 3Blue1Brown Neural Networks: https://www.3blue1brown.com/topics/neural-networks
- Andrej Karpathy Zero to Hero: https://karpathy.ai/zero-to-hero.html
- StatQuest: https://www.youtube.com/c/joshstarmer
- MLCB Conference: https://www.youtube.com/@mlcbconf
Community
- MLSB Workshop: https://www.mlsb.io
- CASP: https://predictioncenter.org
- AlphaFold Database: https://alphafold.ebi.ac.uk
- BioStars: https://www.biostars.org
Conclusion
The path from ML novice to competent practitioner in AI-driven structural biology requires approximately 16-24 weeks of dedicated study—but the investment yields extraordinary capabilities. Learners who complete this roadmap can predict protein structures with near-experimental accuracy, design novel proteins computationally, extract meaningful embeddings from protein sequences, and engage productively with cutting-edge research literature.
Three principles guide successful learning: prioritize depth over breadth in the early phases (truly understanding transformers and diffusion models pays dividends throughout), maintain hands-on practice alongside theory (run ColabFold on day one, even before understanding the architecture), and engage with the community (the field moves too fast to learn in isolation).
The tools and techniques covered here—AlphaFold, ESMFold, RFDiffusion, ProteinMPNN—represent the current state-of-the-art, but the field continues rapid evolution. The foundational understanding of transformers, diffusion models, equivariant networks, and GNNs prepares learners to adapt as new methods emerge. The Nobel Prize recognition in 2024 signals that AI-driven structural biology has entered the mainstream—and the best opportunities lie ahead for those equipped to contribute.
Enjoy Reading This Article?
Here are some more articles you might like to read next: