ESMFold - Protein Language Model
From masked language models to protein generation
Protein structure prediction underwent two revolutions in rapid succession. First, AlphaFold2’s triumph at CASP14 in 2020 demonstrated that deep learning could achieve experimental-level accuracy through sophisticated processing of evolutionary information encoded in multiple sequence alignments. Then, in 2022, Meta AI’s ESMFold challenged a fundamental assumption: the massive language model ESM-2, trained solely to predict masked amino acids, had learned enough about proteins to predict their three-dimensional structures from single sequences alone—eliminating the costly MSA computation entirely. This discovery opened high-throughput structure prediction at unprecedented scale, culminating in the prediction of 617 million metagenomic protein structures in just two weeks.
The philosophical implications run deep. AlphaFold extracts co-evolutionary signals explicitly from thousands of aligned homologous sequences. ESM-2 stores these same statistical patterns implicitly in its 15 billion parameters, having compressed the evolutionary record of life during unsupervised training on 250 million protein sequences. Now, with ESM-3’s multimodal architecture capable of generating novel proteins with functions never seen in nature, the field faces a pivotal question: are language models merely pattern-matching, or have they genuinely learned something fundamental about protein biology?
This post traces the technical evolution from ESM-1’s initial proof of concept through ESM-3’s generative capabilities, comparing each advance against the parallel developments in the AlphaFold family. For computational biologists deploying these tools, understanding their architectural foundations, training paradigms, and failure modes is essential for choosing the right method for each application.
ESM-1 and ESM-1b established that transformers learn protein structure
The original ESM-1 model, released as a preprint in 2019 and formally published in PNAS in 2021 by Rives et al., adapted the BERT architecture to protein sequences with a deceptively simple training objective: mask 15% of amino acids in a protein sequence and train the model to predict them. This masked language modeling (MLM) approach, borrowed directly from natural language processing, required no structural labels whatsoever.
The flagship ESM-1b model comprises 33 transformer layers with 1280-dimensional embeddings totaling 650 million parameters, trained on UniRef50’s approximately 30 million unique sequences. The key architectural modification from standard BERT was pre-activation layer normalization, which stabilizes training for deeper networks. Training proceeded for 500,000 updates on 128 NVIDIA V100 GPUs, processing batches of 131,072 tokens with sequences cropped to 1024 residues.
What made this model scientifically significant was an emergent property: transformer attention patterns corresponded directly to residue-residue contact maps. Rao et al. (2021) demonstrated that a simple logistic regression trained on attention weights from just 20 structures could perform unsupervised contact prediction—the attention mechanism had learned spatial relationships despite never seeing a single 3D coordinate during training. This finding suggested that the evolutionary constraints forcing co-variation between spatially proximal residues were being captured by the language modeling objective.
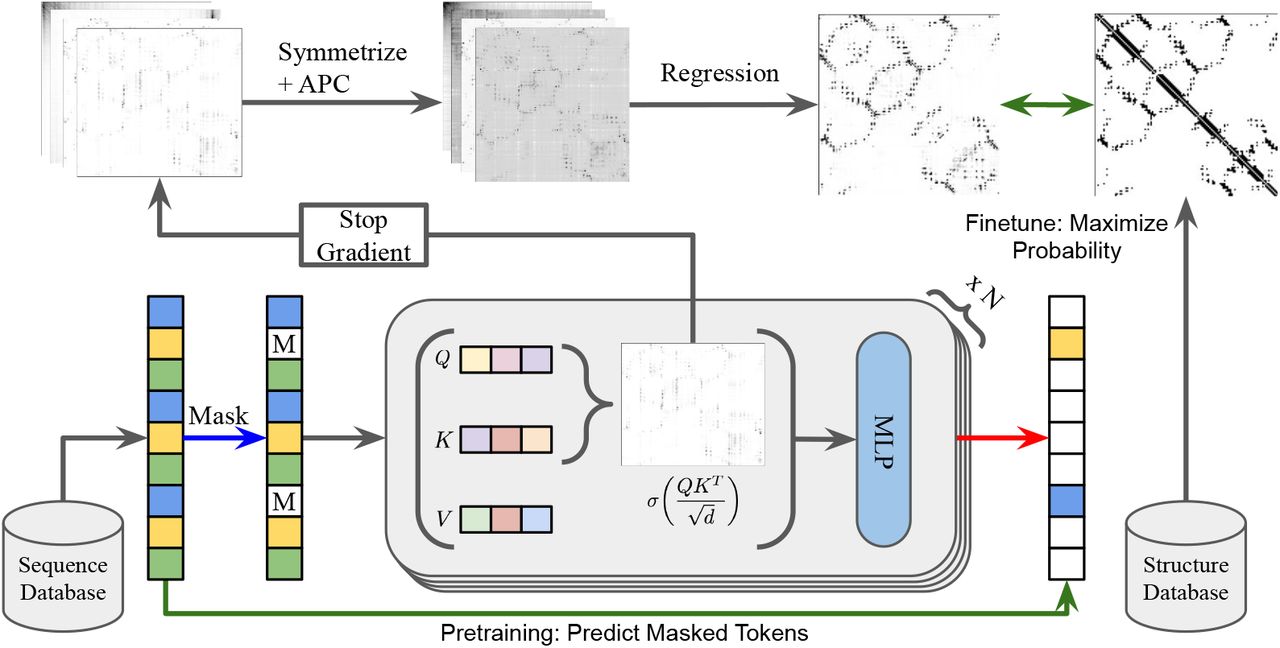
ESM-2 revealed dramatic scaling laws for structural information
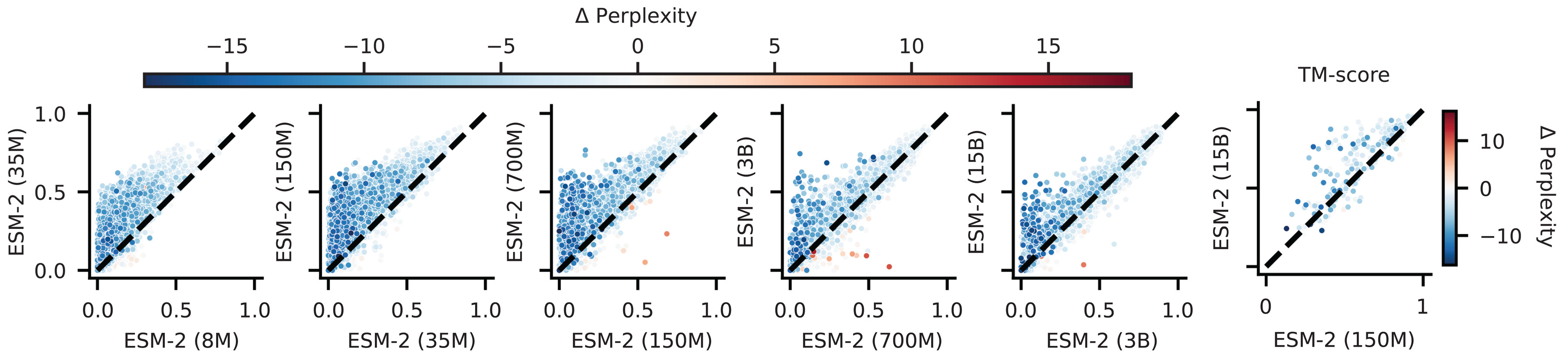
ESM-2, released in August 2022 and published in Science in March 2023, systematically explored how structural information scales with model capacity. The research team trained models spanning three orders of magnitude: 8 million to 15 billion parameters across 6 to 48 transformer layers.
| Model | Parameters | Layers | Embedding Dim | Perplexity |
|---|---|---|---|---|
| esm2_t6_8M | 8M | 6 | 320 | 10.45 |
| esm2_t33_650M | 650M | 33 | 1280 | 7.26 |
| esm2_t36_3B | 3B | 36 | 2560 | 6.82 |
| esm2_t48_15B | 15B | 48 | 5120 | 6.37 |
Two architectural improvements enabled this scaling. First, rotary positional embeddings replaced absolute position encodings, allowing the model to generalize to arbitrary sequence lengths without the 1024-token ceiling of ESM-1b. Second, improved training hyperparameters and increased data—now 65 million sequences from UniRef50/D 2021_04—pushed performance further.
The scaling results revealed a striking pattern: contact prediction accuracy showed nonlinear jumps at specific scale thresholds, and a near-perfect correlation ($r = -0.99$) emerged between validation perplexity and structure prediction accuracy on CASP14. This correlation between language modeling performance and structural understanding suggests that predicting the next amino acid requires, at sufficient scale, implicit modeling of the structural constraints that make certain residues fit better than others. The 150M parameter ESM-2 already outperformed the 650M parameter ESM-1b on structure prediction benchmarks, demonstrating that architectural improvements compound with scale.
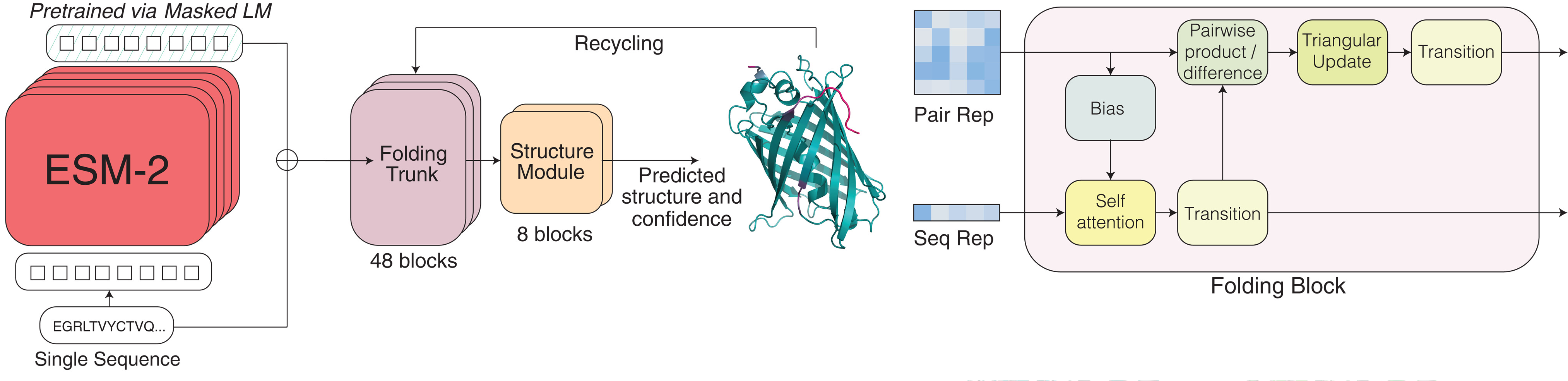
ESMFold converted language model representations into atomic coordinates
The ESMFold architecture, detailed in the same Science 2023 publication, bridges the gap between learned representations and three-dimensional structures through three integrated modules that process a single input sequence without any MSA computation.
The language model backbone (ESM-2 with either 3B or 15B parameters) processes the input sequence and outputs two representations: a per-residue single representation encoding individual amino acid contexts, and a pairwise representation encoding relationships between all residue pairs. These representations emerge from the attention patterns that already encode structural information.
The folding trunk comprises 48 simplified Evoformer blocks that iteratively refine these representations. Unlike AlphaFold2’s Evoformer, which jointly processes MSA and pair representations, ESMFold’s trunk operates only on the single and pair representations derived from the language model. This trunk employs recycling—feeding outputs back as inputs for multiple iterations—to progressively refine structural predictions.
The structure module contains 8 blocks of equivariant transformers employing Invariant Point Attention (IPA), adapted directly from AlphaFold2. Each residue is represented as a rigid body frame (rotation and translation for the N-Cα-C backbone) plus torsion angles for side chains. The IPA mechanism generates 3D query, key, and value points in local residue frames, projects them to global coordinates for attention computation, then projects results back—maintaining SE(3) equivariance throughout. The module outputs atomic coordinates, per-residue confidence scores (pLDDT), and predicted template modeling score (pTM).
The training strategy combined 25,000 experimentally determined structures from the PDB with 12 million AlphaFold2 predictions in a 25:75 ratio, addressing the limited quantity of experimental data. This approach exploits the accuracy of AlphaFold2 predictions to provide abundant training signal while grounding the model in experimental reality.
The speed advantage enabled metagenomic-scale prediction
ESMFold’s elimination of MSA computation yields dramatic speedups. Predicting a 384-residue protein takes approximately 14 seconds on a single V100 GPU, compared to the 30-40 minutes required for AlphaFold2’s full pipeline including jackhmmer MSA search. For shorter sequences, ESMFold achieves 60-fold speedups. This efficiency difference is not merely quantitative—it enables qualitatively different applications.
The ESM Metagenomic Atlas, released alongside ESMFold, predicted structures for 617 million metagenomic proteins using approximately 2,000 GPUs over two weeks. Of these predictions, 29% achieved mean pLDDT above 0.7 (good confidence), with 5.9% exceeding pLDDT 0.9 (high confidence). The atlas identified 7,271 high-confidence structures with TM-score below 0.5 to any PDB entry—potentially novel folds that experimental structural biology had never characterized. Such large-scale structural surveys would be computationally impractical with MSA-dependent methods.
The accuracy tradeoff is real but context-dependent. On CASP14, ESMFold achieved TM-score 0.68 compared to AlphaFold2’s 0.85—a substantial gap. However, when AlphaFold2 runs without MSA (single-sequence mode), its TM-score plummets to 0.37, demonstrating that ESM-2’s language model representations genuinely capture structural information unavailable from raw sequences alone. For high-confidence predictions (pLDDT > 0.85), ESMFold achieves median backbone RMSD of 1.33Å and all-atom RMSD of 1.91Å, approaching experimental accuracy.
ESM-3 represents a paradigm shift to multimodal generative modeling
ESM-3, developed by EvolutionaryScale (the company founded by former Meta AI researchers in 2023) and published in Science in January 2025, fundamentally reconceives the ESM approach. Rather than a discriminative language model followed by a separate folding module, ESM-3 is a unified multimodal generative model that simultaneously reasons over sequence, structure, and function.
The architecture accepts seven input tracks: amino acid tokens, atomic coordinates, discretized structure tokens, 8-class secondary structure labels, quantized solvent-accessible surface area, function keywords (InterPro annotations, GO terms), and residue annotation features. The first transformer block incorporates geometric attention that conditions on 3D atomic coordinates directly, enabling spatially proximal residues to receive higher attention scores through an invariant geometric attention mechanism.
The critical innovation enabling structure-to-token conversion is a two-stage VQ-VAE (Vector Quantized Variational Autoencoder). Stage one trains a small encoder (6M parameters) with attention restricted to 30 nearest neighbors, learning a codebook of 4096 discrete structure tokens that capture local atomic environments around each residue. Stage two freezes this encoder and trains a larger decoder (300M parameters) based on the ESMFold folding trunk architecture to reconstruct full 3D structures including side chains. This tokenization preserves sequence length—each residue receives one structure token—while enabling the transformer to process structure through the same attention mechanisms it uses for sequence.
Training proceeded at unprecedented scale: 1.07 × 10²⁴ FLOPs on 2.78 billion proteins representing 771 billion unique tokens, approximately 25× more compute and 60× more data than ESM-2. The model family spans 1.4B parameters (open-weights), 7B, and a flagship 98B parameter version. Unlike classical masked language modeling with fixed 15% masking, ESM-3 uses variable masking rates during training, supervising predictions across all possible masking levels to enable flexible generation.
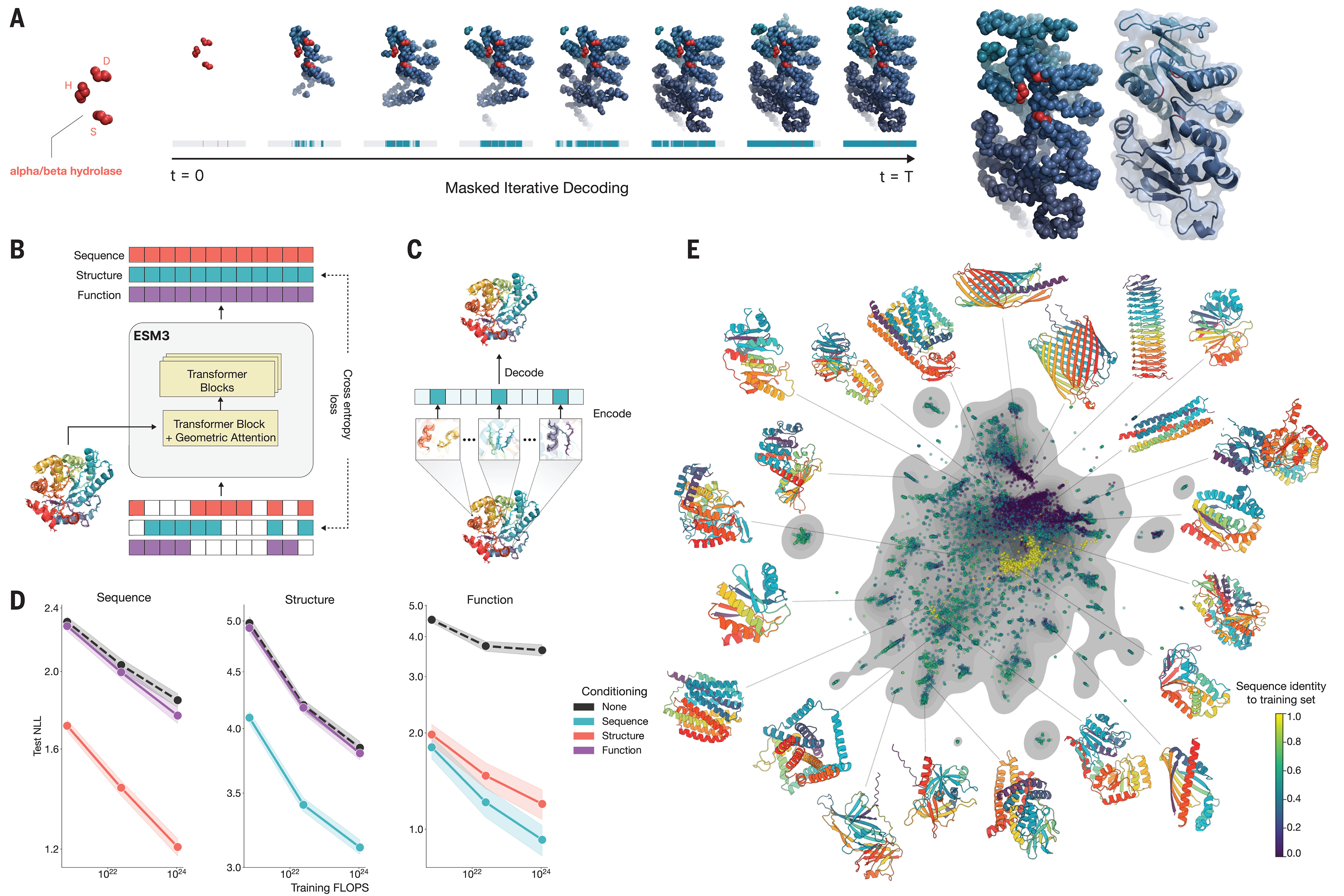
Generative capabilities enable protein design from arbitrary prompts
ESM-3’s training on variable masking enables iterative unmasking generation: starting from fully masked tokens across all modalities, the model progressively predicts and reveals tokens until a complete protein specification emerges. This enables prompting with partial information in any modality.
Motif scaffolding prompts the model with atomic coordinates of key residues—an enzyme’s catalytic triad, a binding site’s contact residues—and generates compatible protein scaffolds. Function-conditioned generation prompts with keywords like “α/β hydrolase” to generate proteins matching functional specifications. Atomic coordination tasks specify residues that must be close in 3D space despite being distant in sequence, testing the model’s ability to design appropriate tertiary structure.
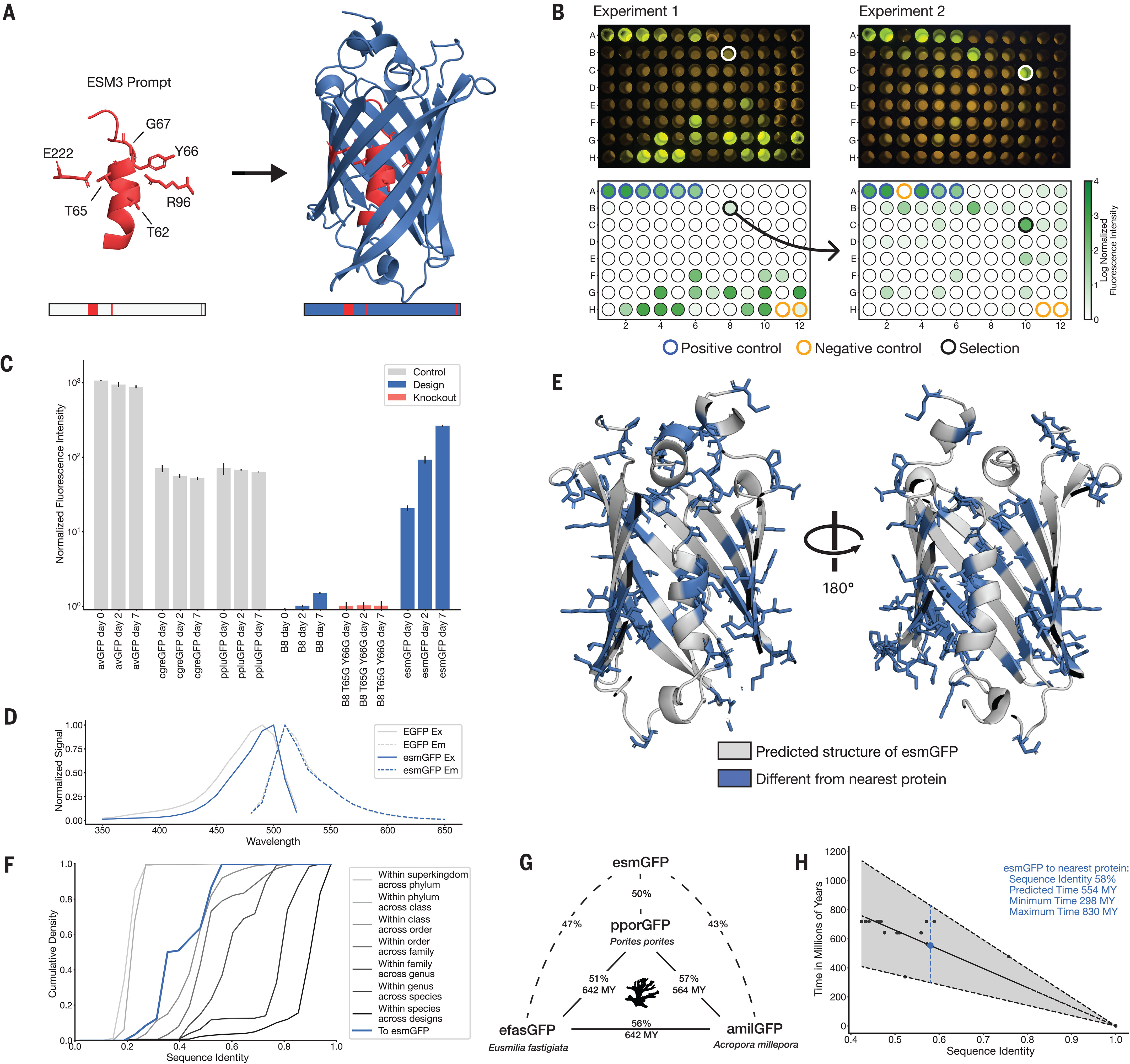
The most striking demonstration is esmGFP, a novel green fluorescent protein generated through chain-of-thought reasoning over sequences and structures. Starting from the chromophore-forming residue positions of natural GFP, ESM-3 generated a protein with 58% sequence identity to the nearest natural fluorescent protein—96 mutations across 229 residues, representing an evolutionary distance equivalent to approximately 500 million years of natural evolution. Critically, esmGFP spontaneously forms a fluorescent chromophore with peak emission at 512nm (identical to natural GFPs) and brightness comparable to wild-type proteins. This validates that ESM-3 captures enough protein biology to generate functional novelty far from its training distribution.
AlphaFold evolved from distance prediction to end-to-end structure generation
Understanding the ESM trajectory requires comparison with AlphaFold’s parallel evolution. AlphaFold 1, presented at CASP13 in 2018, used a deep residual convolutional neural network to predict probability distributions over binned inter-residue distances (64 distance bins producing N×N “distograms”). A separate network predicted backbone torsion angles. Structure assembly then used gradient descent to minimize a potential derived from these distograms—a two-stage approach requiring post-hoc optimization.
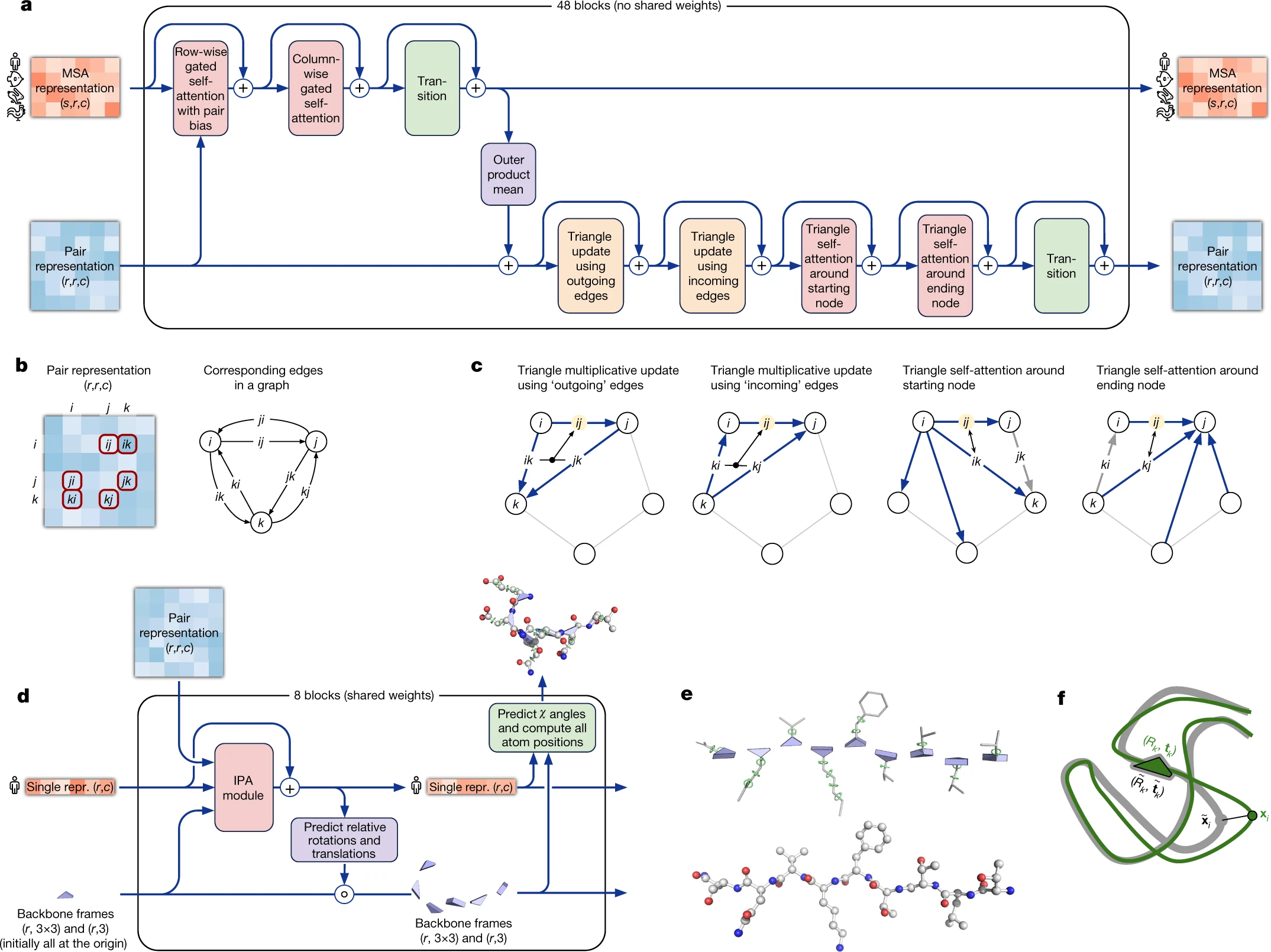
AlphaFold2, the CASP14-dominating system, introduced end-to-end differentiable structure prediction. Its two main components—the Evoformer (48 blocks jointly processing MSA and pair representations) and the Structure Module (converting refined representations to 3D coordinates via IPA)—are trained jointly with gradients flowing from structural loss back through sequence processing.
The Evoformer’s innovations center on co-evolutionary reasoning. Row-wise and column-wise attention processes the MSA representation (sequences × residues). The outer product mean transforms MSA representations into pair representation updates. Triangle multiplicative updates and triangle attention exploit the geometric constraint that pairwise distances must satisfy the triangle inequality—if residue A is close to B, and B is close to C, this constrains the A-C distance. The pair representation is interpreted as edges in a directed graph, with triangle operations enforcing geometric consistency across triplets.
AlphaFold3 adopted diffusion for universal biomolecular modeling
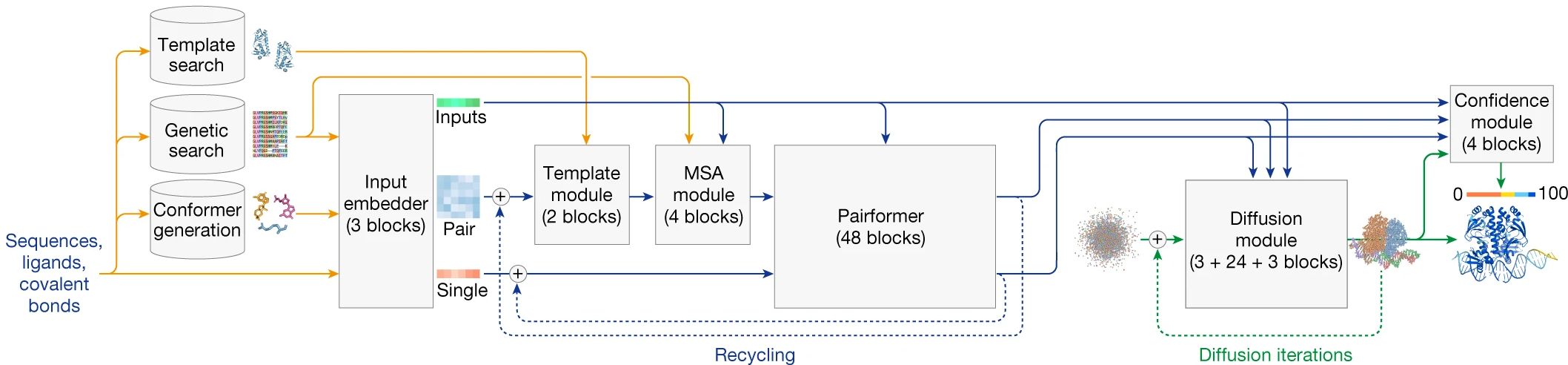
AlphaFold3, released in May 2024, represents another architectural revolution within the AlphaFold lineage. Most dramatically, it replaces the structure module entirely with a diffusion module that operates directly on raw atom coordinates without rigid body frames or torsion angle representations.
The MSA processing simplifies substantially: from 48 Evoformer blocks to just 4 MSA blocks using pair-weighted averaging. The heavy lifting shifts to 48 Pairformer blocks operating only on pair and single representations, eliminating the MSA representation from downstream processing. The diffusion module then generates structures by iteratively denoising from random coordinates to final atomic positions—a generative approach producing distributions over possible structures rather than single predictions.
This architectural shift brings several advantages. Diffusion naturally handles arbitrary chemical components without specialized parameterization, enabling prediction of protein-ligand, protein-DNA/RNA, and protein-protein complexes within a unified framework. The multiscale learning inherent to diffusion—low noise levels learn local stereochemistry while high noise levels learn global fold—eliminates the need for explicit violation losses enforcing bond lengths and angles. On the PoseBusters benchmark for protein-ligand prediction, AlphaFold3 achieves 76.7% success rate (RMSD < 2Å) compared to 52.1% for traditional docking with holo protein structures.
However, the generative nature introduces new failure modes. AlphaFold3 exhibits a 4.4% chirality violation rate on benchmarks—atoms placed in mirror-image configurations. Atomic clashing occurs, particularly in homomeric complexes. And the diffusion process can hallucinate spurious structure in intrinsically disordered regions, though cross-distillation from AlphaFold-Multimer v2.3 predictions (representing disorder as extended loops) mitigates this tendency.
MSA-based and language model approaches encode the same evolutionary information differently
The philosophical distinction between AlphaFold and ESM reduces to how they access evolutionary information. AlphaFold performs explicit co-evolutionary inference: given a query sequence, it searches databases to identify thousands of homologs, constructs an MSA, and extracts residue coupling patterns indicating spatial proximity. This requires 2-3 TB of sequence databases, CPU-intensive homology search (30+ minutes with jackhmmer), and access to sufficient homologous sequences.
ESM-2 stores this co-evolutionary information implicitly in parameters learned during unsupervised training. Recent theoretical analysis (PNAS 2024) demonstrates that ESM-2 stores statistics of co-evolving residue pairs analogously to Markov Random Fields—the same mathematical framework underlying direct coupling analysis of MSAs. The model learns motifs of pairwise contacts extractable through attention patterns. Both approaches ultimately capture that residues in spatial contact show correlated mutations across evolution; they merely differ in whether this extraction occurs at training time (ESM) or inference time (AlphaFold).
This distinction has practical consequences. When deep MSAs are available (>10,000 sequences), AlphaFold2’s explicit co-evolutionary reasoning provides TM-scores of 0.85 versus ESMFold’s 0.68. But for orphan proteins lacking homologs, the advantage reverses slightly—ESMFold achieves comparable or marginally better performance because its pretrained parameters encode evolutionary patterns beyond any single MSA. For de novo designed proteins with no natural homologs, MSA-based methods cannot function while language models retain their learned priors.
| Scenario | AlphaFold2 TM-score | ESMFold TM-score | Winner |
|---|---|---|---|
| Deep MSA (>10K sequences) | 0.86 | 0.79 | AlphaFold2 |
| Shallow MSA (~30 sequences) | 0.75 | 0.72 | AlphaFold2 |
| Orphan proteins | 0.46 | 0.48 | ESMFold |
| De novo designs | N/A | Functional | ESMFold |
The MSA depth threshold matters substantially. AlphaFold2’s accuracy drops sharply with fewer than 30 effective sequences. ColabFold’s MMseqs2-GPU acceleration reduces MSA search from 30+ minutes to ~1.5 minutes total prediction time, partially mitigating AlphaFold2’s speed disadvantage while maintaining accuracy—an important practical compromise for many applications.
How architectural choices reflect different modeling philosophies
AlphaFold’s architecture explicitly encodes geometric priors: the Evoformer’s triangle operations enforce distance consistency, IPA operates in SE(3)-equivariant feature spaces, and the structure module uses rigid body frames respecting protein backbone geometry. These inductive biases help the model learn efficiently from limited structural data by building in physical knowledge.
ESM’s architecture is more general: standard transformers with attention patterns that happen to learn structural information through the pressure of masked prediction. ESM-3’s geometric attention layer is the first explicit geometric induction in the ESM family, and even then, the bulk of the architecture remains modality-agnostic. This generality enables the multimodal flexibility that lets ESM-3 reason jointly across sequence, structure, and function, but may require more data and compute to match architectures with stronger physics priors.
AlphaFold3’s shift to diffusion represents a convergence toward generality—removing equivariant structure modules in favor of standard transformers with diffusion denoising. This architectural choice, while sacrificing some guaranteed geometric properties (hence the chirality violations), enables the universal biomolecular modeling that handles proteins, nucleic acids, ligands, ions, and modifications within a single framework.
Practical implications for choosing structure prediction methods
For computational structural biologists, method selection depends on the specific application requirements.
Choose ESMFold when conducting high-throughput screening of millions of sequences, predicting structures for metagenomics projects, rapidly assessing structural viability during protein engineering iterations, working with de novo designed proteins lacking evolutionary history, or operating under computational resource constraints that preclude large database storage and MSA search.
Choose AlphaFold2 when maximum accuracy justifies longer compute times, well-characterized protein families provide deep MSAs, final validation before experimental work demands the highest confidence, or stereochemical quality (Ramachandran distributions, MolProbity scores) is critical for downstream molecular dynamics or docking.
Choose AlphaFold3 when predicting protein-ligand or protein-nucleic acid complexes, working with modified residues, glycosylation, or covalent modifications, or needing unified treatment of diverse biomolecular interactions. Accept that chirality violations and occasional clashes require post-prediction quality checking.
Choose ESM-3 when generating novel proteins from structural or functional constraints, performing motif scaffolding from active site coordinates, exploring sequence-structure-function relationships jointly, or designing proteins for specified functions without natural templates.
A hybrid workflow increasingly represents best practice: use ESMFold for initial filtering at scale, follow with AlphaFold2 or AlphaFold3 on top candidates for final validation, and turn to ESM-3 for generative design tasks.
Conclusions and the path forward
The ESM trajectory—from masked language model demonstrating emergent structural understanding, through single-sequence structure prediction enabling metagenomic-scale surveys, to multimodal generative models producing functional novel proteins—represents a fundamental expansion in what computational methods can achieve for protein biology. The AlphaFold trajectory pursued a different path, from explicit distogram prediction through end-to-end differentiable structure generation to universal diffusion-based biomolecular modeling.
Neither approach has obsoleted the other. AlphaFold2 remains the accuracy leader for structure prediction when MSAs are available and compute time is not limiting. ESMFold enables applications at scales that MSA-dependent methods cannot reach. ESM-3 opens generative capabilities that discriminative structure prediction cannot address, while AlphaFold3 handles biomolecular complexity—complexes, modifications, diverse chemical entities—that protein-only models cannot touch.
The field increasingly recognizes that these approaches encode the same underlying biological information through different computational strategies. Future methods will likely combine explicit evolutionary inference with learned implicit representations, geometric priors with generative flexibility, and discriminative accuracy with generative creativity. For now, understanding the strengths, limitations, and appropriate applications of each approach allows computational biologists to select the right tool for each scientific question.
References
-
Devlin et al., BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding, Arxiv 2019. https://doi.org/10.48550/arXiv.1810.04805
-
Rao et al., Transformer protein language models are unsupervised structure learners, BioRxiv 2020. https://doi.org/10.1101/2020.12.15.422761
-
Rives et al., Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences, PNAS 2021. https://doi.org/10.1073/pnas.2016239118
-
Lin at al., Evolutionary-scale prediction of atomic-level protein structure with a language model, Science 2023. https://doi.org/10.1126/science.ade2574
-
Thomas Hayes et al., Simulating 500 million years of evolution with a language model, Science 2025. https://doi.org/10.1126/science.ads0018
-
Jumper at al., Highly accurate protein structure prediction with AlphaFold, Nature 2021. https://doi.org/10.1038/s41586-021-03819-2
-
Abramson et al., Accurate structure prediction of biomolecular interactions with AlphaFold 3, Nature 2024. https://doi.org/10.1038/s41586-024-07487-w
Enjoy Reading This Article?
Here are some more articles you might like to read next: