SE(3) and E(3) Equivariance in Computational Structural Biology
Symmetry as a Design Principle
The introduction of $\text{SE(3)}$-equivariant neural networks has transformed computational structural biology, enabling algorithms that inherently respect the rotational and translational symmetries of three-dimensional molecular structures. This architectural innovation—pioneered through methods like Invariant Point Attention in AlphaFold2 and extended across protein design tools like RFdiffusion—delivers up to 1000-fold improvements in data efficiency while achieving unprecedented accuracy in structure prediction and design. The shift from invariant to equivariant representations marks a fundamental reconceptualization of how we encode geometric priors into machine learning systems for biomolecular modeling.
The mathematical language of molecular symmetry
Group theory foundations: why $\text{E(3)}$ and $\text{SE(3)}$ matter
Physical molecules exist in three-dimensional Euclidean space, and their properties—binding energies, conformational preferences, catalytic activities—remain unchanged when we rotate or translate them in space. This fundamental physical reality is formalized through group theory. The Special Euclidean group $\mathrm{SE(3) = SO(3) ⋊ ℝ^3}$ describes all rigid body motions: rotations (the 3D rotation group $\text{SO(3)}$) combined with translations ($ℝ^3$). The full Euclidean group $\mathrm{E(3) = O(3) ⋊ ℝ^3}$ additionally includes reflections, relevant for molecules where chirality is not a concern.
These groups act on molecular coordinates through well-defined transformations. A rotation $\mathrm{\mathcal{R} ∈ SO(3)}$ applied to atom positions $\mathbf{x}$ yields $\mathcal{R}\mathbf{x}$, while a translation $\mathbf{t}$ yields $\mathbf{x+t}$. The semidirect product structure (⋊) captures how rotations and translations combine non-commutatively: rotating then translating differs from translating then rotating. Understanding this structure is essential because neural network layers must respect these composition rules to maintain equivariance through deep networks.
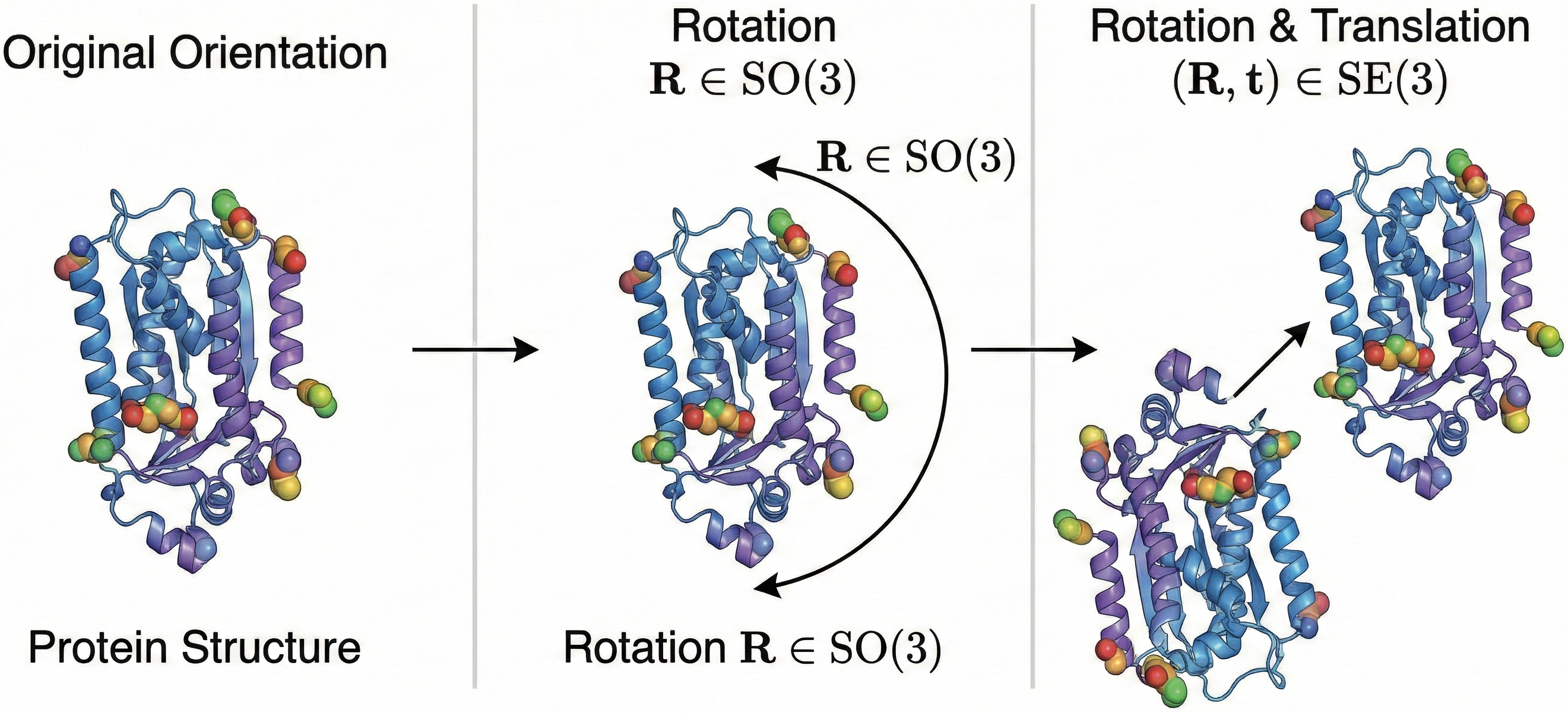
Figure 1: Visualization of $\text{SE(3)}$ group actions on a protein structure. (Left) A protein structure in its original orientation. (Middle) The same structure after rotation $\mathrm{\mathcal{R} ∈ SO(3)}$. (Right) The structure after combined rotation and translation $\mathrm{(\mathcal{R}, t) ∈ SE(3)}$. The physical properties of the molecule remain unchanged under these transformations, motivating $\text{SE(3)}$-equivariant neural network architectures.
Equivariance versus invariance: the mathematical distinction
The formal definition of equivariance captures the core design principle. A function $\mathrm{f: X → Y}$ is G-equivariant if:
\[f(ρ_X(g) · x) = ρ_Y(g) · f(x), \mathrm{∀g ∈ G, x ∈ X}\]where $ρ_X$ and $ρ_Y$ are group representations on the input and output spaces. When $ρ_Y$ is the trivial representation (the identity), equivariance reduces to invariance: $f(ρ_X(g) · x) = f(x)$. For molecular property prediction (like binding affinity), the final output should be invariant—the same molecule rotated in space has the same properties. However, intermediate representations benefit enormously from being equivariant rather than invariant.
The difference is profound. An invariant network discards all directional information immediately, representing molecular geometry only through scalars like pairwise distances. An equivariant network preserves directional information through features that transform predictably under rotations—vectors, tensors, and higher-order geometric objects that maintain their mathematical relationship to the underlying structure. This preserved geometric information enables more expressive representations and more accurate predictions.
Irreducible representations: the building blocks of equivariant features
The mathematical machinery for constructing $\mathrm{SE(3)}$-equivariant neural networks comes from representation theory. Any representation of $\text{SO(3)}$ can be decomposed into a direct sum of irreducible representations (irreps), labeled by non-negative integers $\ell = 0, 1, 2, …$ Each irrep has dimension $(2\ell+1)$ and corresponds to a specific type of geometric object:
- $\ell = 0$: Scalars (1-dimensional), invariant under rotations
- $\ell = 1$: Vectors (3-dimensional), transform like coordinates
- $\ell = 2$: Traceless symmetric 2-tensors (5-dimensional)
- $\ell ≥ 3$: Higher-order tensors
Under rotation $\mathcal{R}$, a type-l feature transforms via the Wigner D-matrix: $f^{\ell} → \mathcal{D}^{\ell}(\mathcal{R}) · f^{\ell}$. The D-matrices are $(2\ell+1) × (2\ell+1)$ unitary matrices that encode how spherical harmonic functions transform. The connection to spherical harmonics $Y_{\ell}^m(θ,φ)$ is fundamental: these functions form a complete orthonormal basis on the sphere and transform equivariantly under rotations:
\[Y_{\ell}^m(\mathcal{R}^{(-1)}x̂) = \sum_{m'} \mathcal{D}^{\ell}_{m'm}(\mathcal{R}) Y_{m'}^{\ell(x̂)}\]
Figure 2: Real spherical harmonics $Y_{\ell m}(\theta,\phi)$.. Rows correspond to the angular momentum degree $\ell = 0, 1, 2, \ldots$, while columns correspond to the order $m = -\ell, \ldots, 0, \ldots, +\ell$. Colors indicate the sign of the harmonic (red: positive; blue: negative), with intensity proportional to the magnitude. The real basis is constructed from linear combinations of the complex spherical harmonics, yielding purely real angular patterns commonly used in physical and chemical applications. Source: e3nn.org.
Tensor products and Clebsch-Gordan coefficients
Combining features of different types requires the Clebsch-Gordan decomposition. When two irreps of degrees $\ell_1$ and $\ell_2$ are multiplied (tensor product), they decompose into a direct sum:
\[\mathcal{D}^{\ell_1} ⊗ \mathcal{D}^{\ell_2} = ⊕_{\ell=|\ell_1-\ell_2|}^{\ell_1+\ell_2} \mathcal{D}^{\ell}\]The Clebsch-Gordan coefficients $\langle \ell_1 m_1 \ell_2 m_2 \vert \ell m \rangle$ provide the precise linear combinations needed to perform this decomposition. These coefficients enable equivariant nonlinearities: while pointwise nonlinearities would break equivariance for $\ell > 0$ features, tensor products followed by CG decomposition provide a mathematically principled way to mix information across feature types while preserving transformation properties.
This framework—irreps, spherical harmonics, Wigner D-matrices, and Clebsch-Gordan coefficients—forms the mathematical foundation for architectures like Tensor Field Networks, $\text{SE(3)}$-Transformers, and Equiformer.
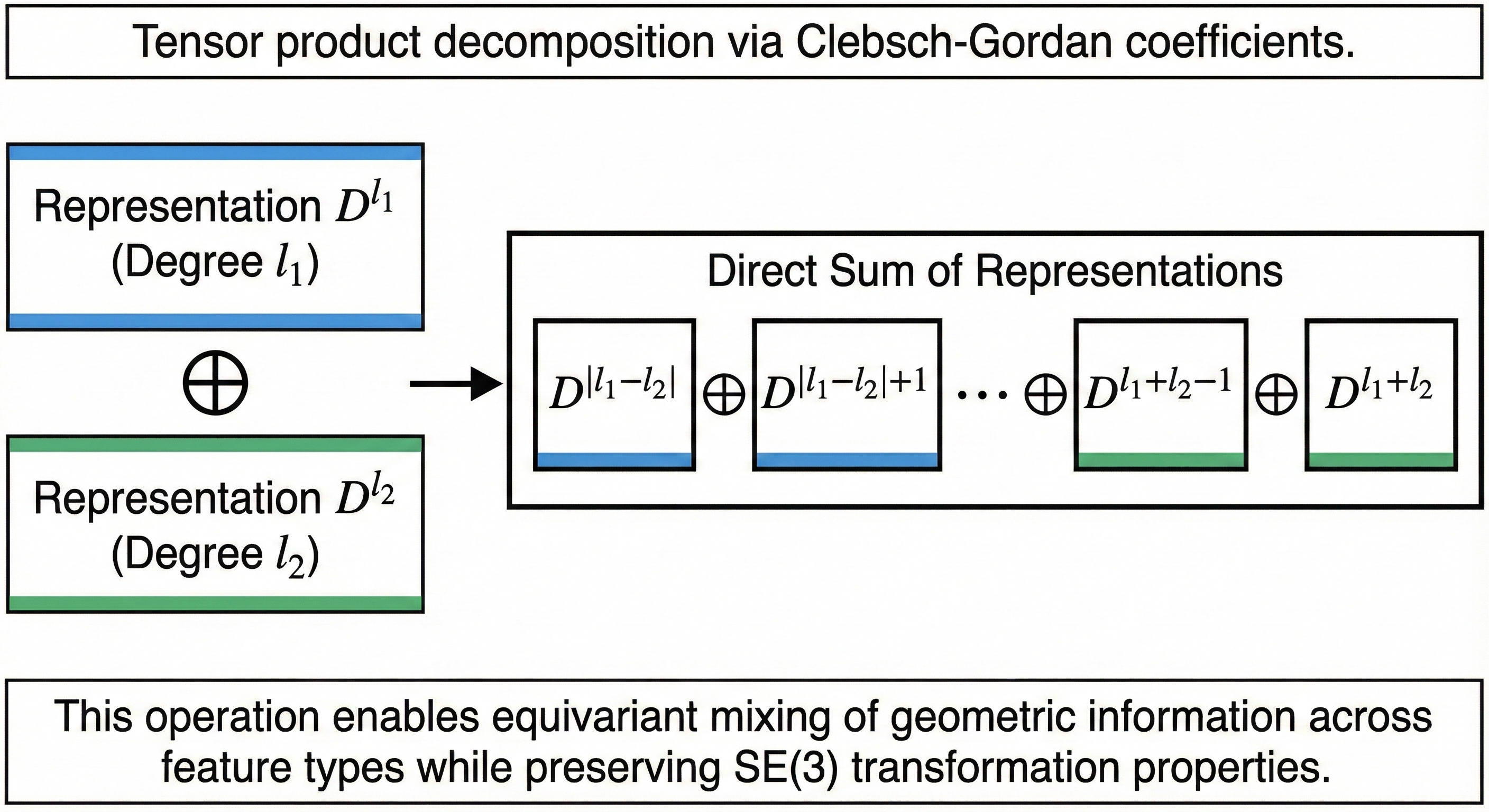
Figure 3: Tensor product decomposition via Clebsch-Gordan coefficients.. When two irreducible representations $\mathcal{D}^{\ell_1}$ and $\mathcal{D}^{\ell_2}$ are combined, they decompose into a direct sum of representations with degrees ranging from $\vert \ell_1 - \ell_2 \vert$ to $\ell_1 + \ell_2$. This operation enables equivariant mixing of geometric information across feature types while preserving $\text{SE(3)}$ transformation properties.
The data efficiency revolution: why symmetry constrains learning
The connection between equivariance and data efficiency is both theoretically grounded and empirically dramatic. The landmark NequIP study (Batzner et al., Nature Communications 2022) demonstrated that $\text{E(3)}$-equivariant networks achieve equivalent accuracy to invariant methods with up to 1000× less training data for molecular potential energy surfaces.
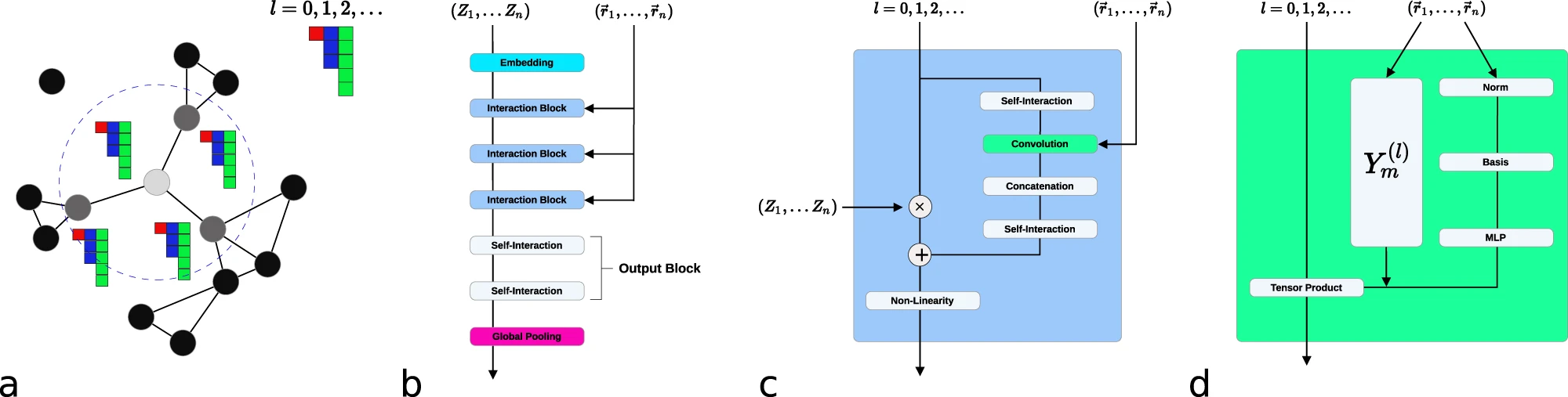
Figure 4: NequIP network architecture for $\text{E(3)}$-equivariant molecular potentials. (a) Atomic system represented as a graph with local neighborhoods. (b) Atomic numbers embedded into $\ell=0$ features, refined through interaction blocks to create scalar and tensor features. (c) The interaction block structure. (d) Convolution operation combining radial functions $\mathcal{R}(r)$ with spherical harmonic projections. Source: Batzner et al., Nature Communications 2022, Figure 1.
This improvement arises from three mechanisms. First, parameter sharing: equivariant constraints reduce the number of free parameters, analogous to how translation equivariance in CNNs shares weights across spatial positions. Second, hypothesis space reduction: constraining the function class to equivariant functions focuses learning on physically meaningful solutions. Third, implicit data augmentation: each training example effectively provides information about all its symmetry-related transformations simultaneously.
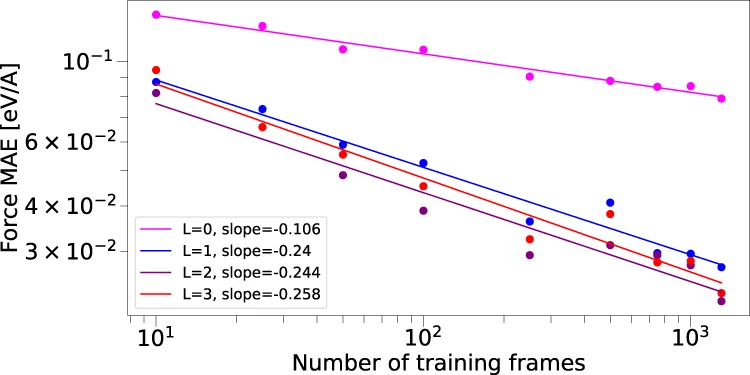
Figure 5: Data efficiency of $\text{E(3)}$-equivariant networks.. Log-log plot of force prediction error (MAE) versus training set size for water molecules, comparing NequIP ($\text{E(3)}$-equivariant) with various $\ell_{max}$ values against non-equivariant baselines. NequIP achieves equivalent accuracy with up to 1000× fewer training examples, demonstrating the dramatic data efficiency gains from incorporating geometric symmetries. Source: Batzner et al., Nature Communications 2022, Figure 5.
Empirically, equivariant networks exhibit qualitatively different learning curves. While invariant models follow standard power-law improvement ($ε ∝ N^b$ for training set size $N$), equivariant models show steeper log-log slopes—they extract more information per additional training example. This finding has profound implications for molecular modeling where generating training data through quantum mechanical calculations is computationally expensive.
Invariant Point Attention: the AlphaFold2 breakthrough
Frame-based equivariance without spherical harmonics
AlphaFold2’s Invariant Point Attention (IPA) represents a conceptually distinct approach to $\text{SE(3)}$ equivariance. Rather than using spherical harmonics and irreducible representations, IPA operates on local coordinate frames attached to each residue, defined by backbone atoms ($\text{N, Cα, C}$).
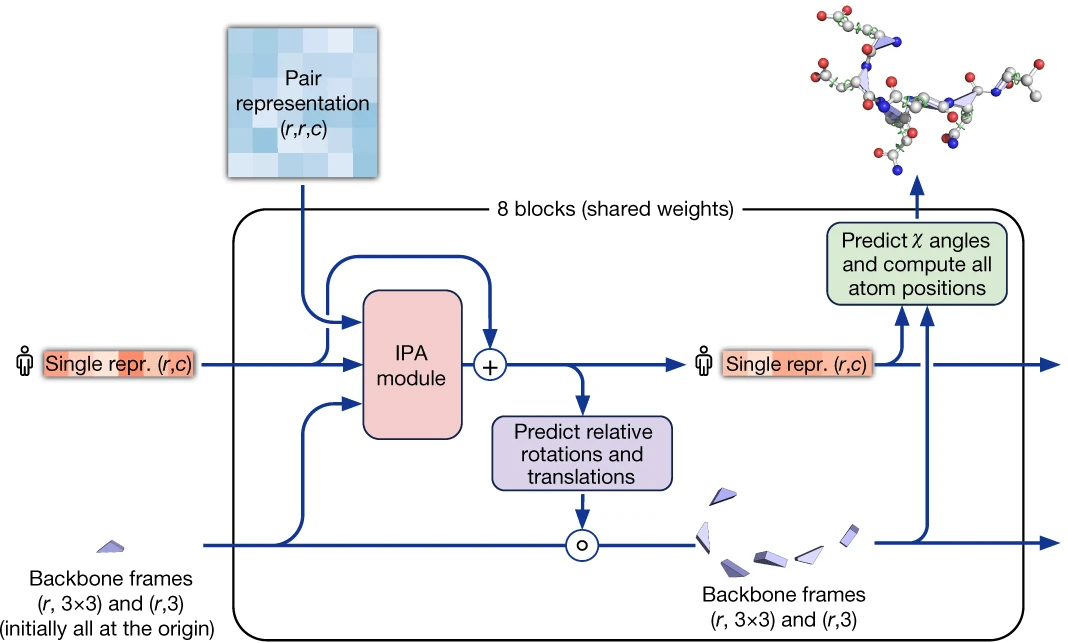
Figure 6: AlphaFold2 structure module architecture. The module processes single representations (extracted from MSA) and pair representations through 8 blocks of Invariant Point Attention (IPA). Each block updates both the residue representations and the backbone frames (rotation and translation for each residue). Blue arrays show pair representation modulation, red arrays show standard attention on abstract features, and green arrays show the invariant point attention mechanism operating on 3D coordinates. Source: Jumper et al., Nature 2021, Figure 3D.
The IPA mechanism combines three attention components. Standard query-key dot products provide sequence-level attention. Pair representation biases incorporate evolutionary and geometric relationships from the pair track. Most innovatively, 3D point queries and keys are generated in local residue frames, transformed to a global frame for comparison, and scored based on their Euclidean distances:
\[a_{ij} \propto \exp(... - (γ/2) · \sum_p \lVert T_i ∘ q̃_i^p - T_j ∘ k̃_j^p \rVert^2)\]Here $\mathrm{T_i ∈ SE(3)}$ is the local frame for residue $i$, and the “spraying” mechanism generates multiple 3D query/key vectors per residue. The crucial insight is that squared Euclidean distances are invariant under global rigid transformations: rotating or translating the entire protein doesn’t change the distances between transformed points.
The IPA design philosophy
The IPA approach offers several advantages over spherical harmonic methods. Implementation is simpler—standard linear algebra rather than spherical tensor machinery. Computational scaling is more favorable, avoiding the $\mathrm{O(L^6)}$ complexity of high-degree spherical tensor products. Local frames are naturally defined for proteins (and other polymers) through backbone geometry.
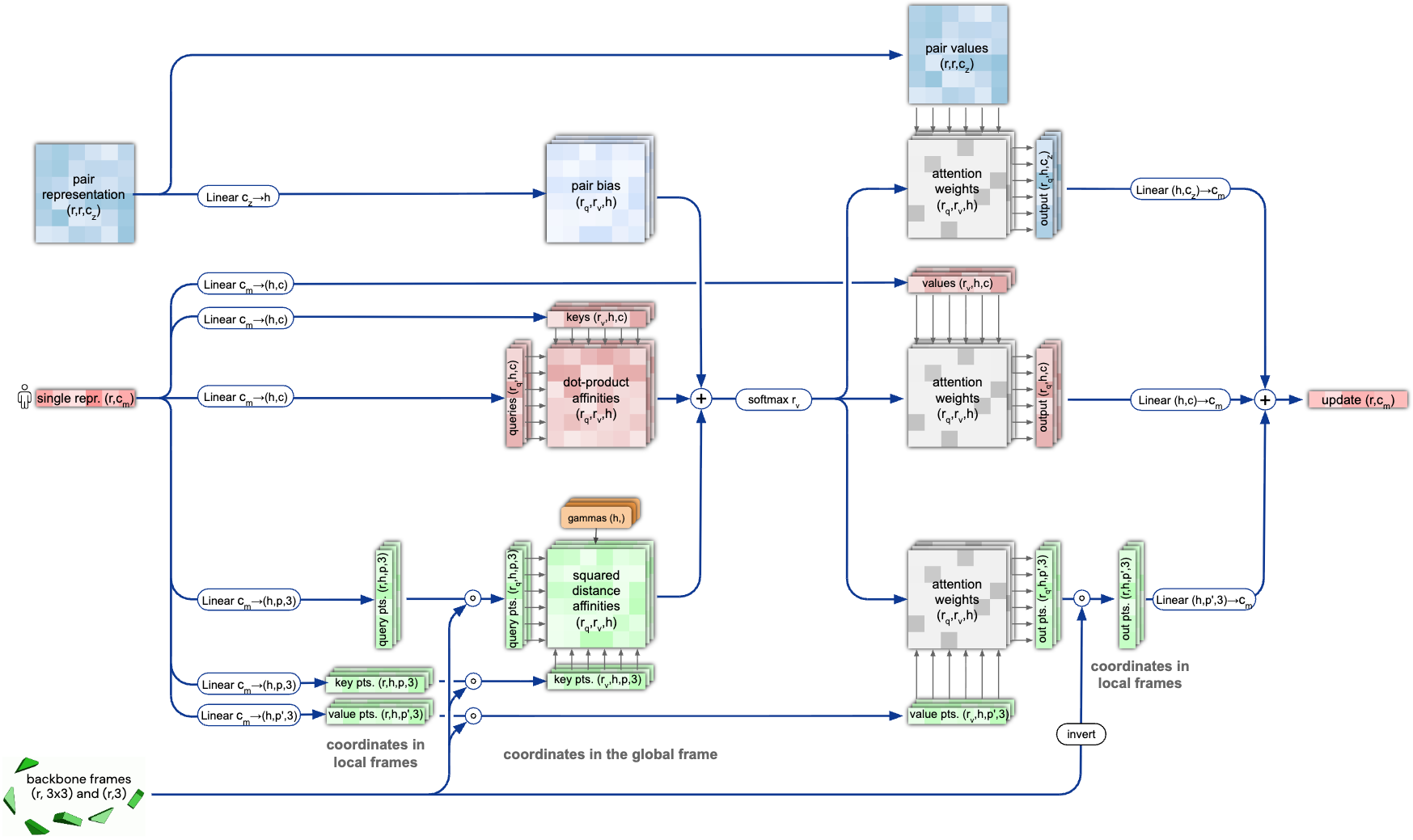
Figure 7: Invariant Point Attention mechanism. (Left) Each residue i has a local coordinate frame $T_i$ defined by its backbone atoms. (Middle) Query and key points ($q̃$ and $k̃$) are generated in local frames. (Right) Points are transformed to a global frame using the residue transformations, and attention weights are computed based on Euclidean distances $\lVert T_i ∘ q̃_i^p - T_j ∘ k̃_j^p \rVert^2$, which are invariant to global rotations and translations of the entire structure. Source: Jumper et al., Nature 2021, Figure S8.
Ablation studies revealed a surprising finding: removing IPA alone only moderately decreases AlphaFold2’s performance, suggesting that much structural information is carried by the recycling mechanism and pair representations. However, IPA becomes critical when combined with recycling, indicating its role in late-stage spatial refinement rather than initial structure determination.
AlphaFold3: the move away from explicit equivariance
AlphaFold3 (Abramson et al., Nature 2024) represents a striking architectural shift. It replaces IPA entirely with a diffusion-based structure prediction module operating on raw atomic coordinates without explicit equivariance constraints. The Evoformer is simplified to a Pairformer, MSA processing is de-emphasized, and the structure module uses denoising diffusion rather than frame-based refinement.
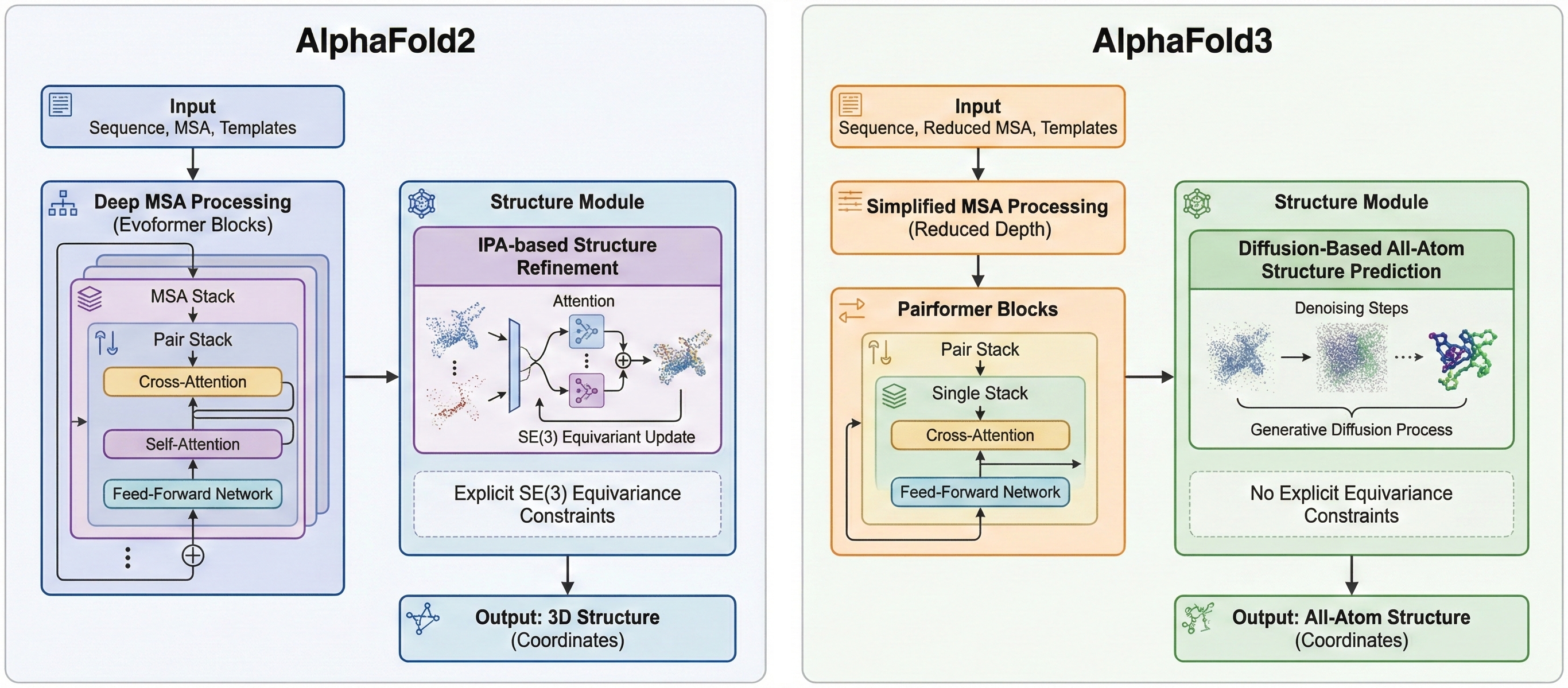
Figure 8: Architectural evolution from AlphaFold2 to AlphaFold3. AlphaFold2 (left) uses deep MSA processing with Evoformer blocks and IPA-based structure refinement with explicit $\text{SE(3)}$ equivariance. AlphaFold3 (right) simplifies to a Pairformer, reduces MSA depth, and replaces IPA with diffusion-based all-atom structure prediction without explicit equivariance constraints.
This change enables AlphaFold3 to model diverse biomolecular complexes—proteins, nucleic acids, small molecules, ions—with a unified architecture. The authors note that “no invariance or equivariance are required after all” when sufficient data and model capacity are available. Whether this represents a fundamental insight or a pragmatic engineering choice for multi-modal modeling remains debated.
Spherical harmonic architectures: TFN, $\text{SE(3)}$-Transformers, and Equiformer
Tensor Field Networks: the foundation
Tensor Field Networks (Thomas et al., 2018) established the spherical harmonic framework for equivariant point cloud processing. Features are organized as spherical tensors of different degrees, and convolutions combine input features with spherical harmonic encodings of relative positions through Clebsch-Gordan tensor products:
\[x_u^{\ell} = \sum_{v∈N(u)} \sum_k \sum_J φ_J^{\ell k}(r) Y^J(r̂) ⊗_\text{CG} x_v^k\]The radial functions $\phi_J^{lk}(r)$ are learnable networks of the scalar distance $\Vert r \Vert$, while the spherical harmonics $Y^{J(r̂)}$ encode directional information. This separation is fundamental: radial functions can be arbitrary neural networks (invariant), while angular dependence comes from the equivariant spherical harmonic basis.
TFN’s computational complexity scales as $O((\ell_{max}+1)^6)$ due to Clebsch-Gordan coefficient calculations, limiting practical use to low degrees (typically $\ell ≤ 3$). Despite this limitation, TFN established the theoretical framework underlying the e3nn library and subsequent architectures.
SE(3)-Transformers: attention meets equivariance
$\text{SE(3)}$-Transformers (Fuchs et al., NeurIPS 2020) combined self-attention with $\text{SE(3)}$ equivariance. Features are organized as “fibers”—concatenations of features across all degrees. Attention weights are computed from type-0 (scalar/invariant) components, ensuring the weighting scheme is $\text{SE(3)}$-invariant. Values span all fiber types, and the weighted combination preserves equivariance.
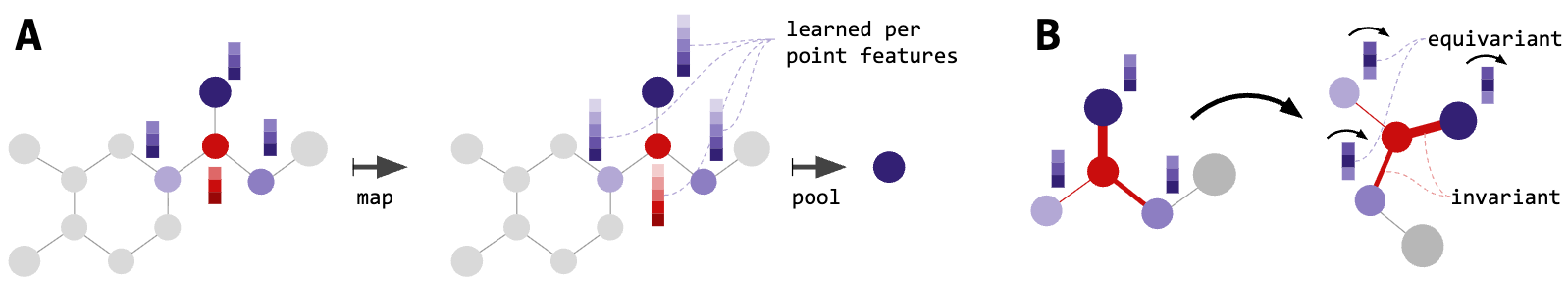
Figure 9: SE(3)-Transformer architecture. Features are organized as 'fibers'—direct sums across all irreducible representation degrees l. Attention weights are computed from type-0 (scalar) features ensuring SE(3)-invariant weighting, while values span all fiber types. The architecture combines self-attention's adaptive filtering with rigorous geometric equivariance. Source: Fuchs et al. "SE(3)-Transformers" NeurIPS (2020), Figure 1.
Key innovations include GPU-accelerated spherical harmonic computation (100-1000× speedup over CPU implementations) and attentive self-interaction layers that increase representational capacity without breaking equivariance. The architecture demonstrated that attention’s adaptive, data-dependent filtering is compatible with geometric equivariance.
Equiformer and EquiformerV2: scaling to higher degrees
Equiformer (Liao & Smidt, ICLR 2023) advanced equivariant transformers with MLP-based attention (more expressive than dot-product), gate activations compatible with equivariance, and careful layer normalization. The architecture achieved state-of-the-art results on QM9 (11/12 molecular property tasks) and OC20 (catalyst discovery).
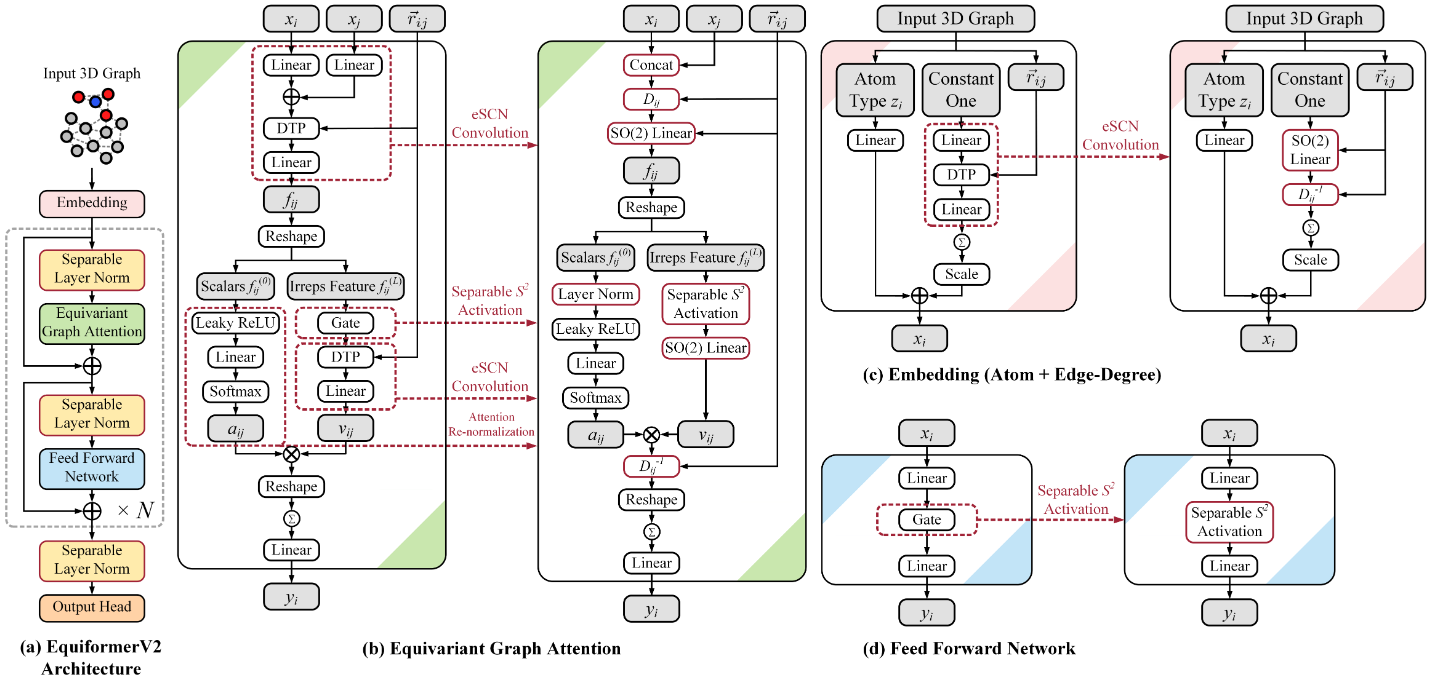
Figure 10: EquiformerV2 architecture improvements. (Top) Network architecture showing eSCN convolutions that replace full $\text{SO(3)}$ tensor products with efficient $\text{SO(2)}$ operations. (Bottom) Computational complexity comparison: traditional spherical tensor methods scale as $\mathrm{O(L^6)}$, while EquiformerV2's eSCN achieves $\mathrm{O(L^3)}$ scaling, enabling practical use of higher degrees ($\ell_{max} = 6-8$) for improved expressivity. Source: Liao et al. "EquiformerV2" ICLR (2024), Figure 1.
The critical bottleneck remained computational scaling with degree. EquiformerV2 (ICLR 2024) addressed this through eSCN convolutions that replace SO(3) tensor products with efficient $\text{SO(2)}$ linear operations, reducing complexity from $\mathrm{O(L^6)}$ to $\mathrm{O(L^3)}$. Additional innovations—attention re-normalization, separable $\mathrm{S^2}$ activations, separable layer normalization—enable scaling to $\ell_\mathrm{max} = 6-8$. On OC20, EquiformerV2 achieves 9% improvement on forces and 4% on energies versus previous state-of-the-art.
Simplified approaches: EGNN and GVP
E(n) Equivariant Graph Neural Networks
EGNN (Satorras et al., ICML 2021) demonstrated that sophisticated equivariance is possible without spherical harmonics or higher-order tensors. The architecture maintains two feature types: invariant node embeddings $h_i$ and equivariant coordinates $x_i$. Message passing updates both:
\[\begin{align} m_{ij} &= φ_e(h_i, h_j, \lVert x_i - x_j \rVert^2, a_{ij}) \\ x_i^{new} &= x_i + \text{C}·\sum_j (x_i - x_j) · φ_x(m_{ij}) \\ h_i^{new} &= φ_h(h_i, \sum_j m_{ij}) \end{align}\](Eq.1) # Invariant messages
(Eq.2) # Coordinate update
(Eq.3) # Node update
The critical insight: coordinate updates use difference vectors ($x_i - x_j$), which transform equivariantly, multiplied by scalar outputs from $φ_x$, which are invariant. This combination preserves equivariance without explicit group representation theory.
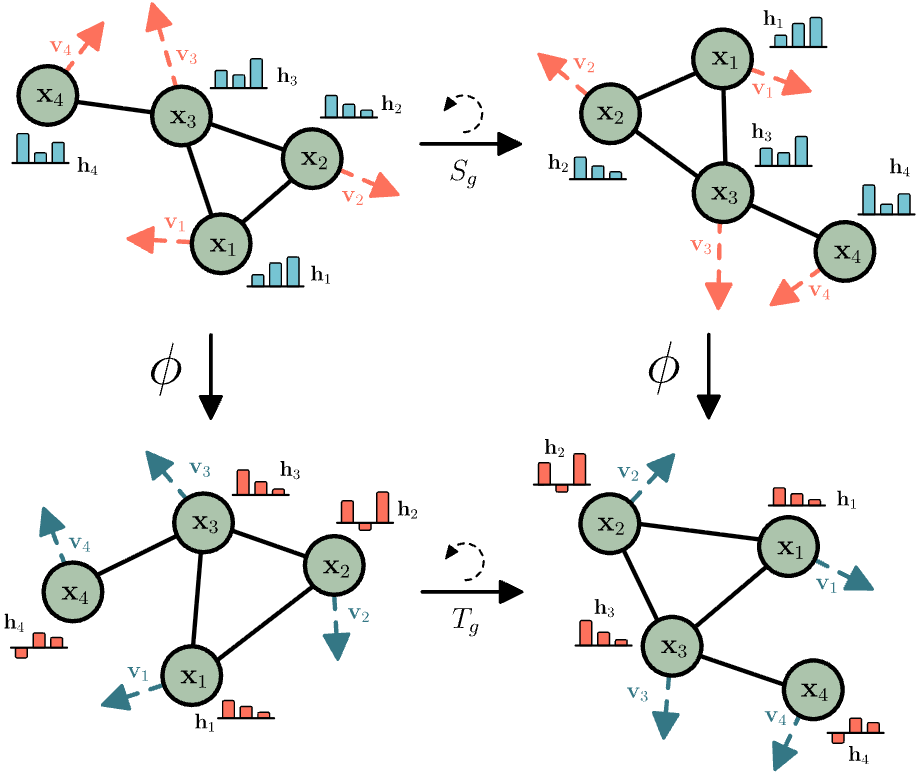
Figure 11: E(n) Equivariant Graph Neural Network (EGNN) architecture. The network maintains two feature types: invariant node embeddings $h_i$ and equivariant coordinates $x_i$. Messages $m_{ij}$ are computed using invariant features (node embeddings and squared distances). Coordinates are updated using difference vectors ($x_i - x_j$) weighted by scalar functions, ensuring equivariance without explicit spherical harmonics or higher-order tensors. Source: Satorras et al. "$\text{E(n)}$ Equivariant Graph Neural Networks" ICML (2021), Figure 1.
EGNN’s simplicity enables efficient scaling to large systems and serves as the backbone for equivariant diffusion models. Limitations include reduced expressivity compared to higher-order methods—EGNN cannot distinguish certain symmetric configurations that spherical tensor methods can.
Geometric Vector Perceptrons
GVP (Jing et al., ICLR 2021) provides another simplified approach, operating on (scalar, vector) feature pairs. The key operation is scalarization: extracting invariant information from vectors via $\text{L2}$ norms, enabling information flow from geometric to scalar features:
\[\begin{align} (s', V') &= \text{GVP}(s, V) \\ V_h &= W_h · V \\ s' &= σ(W_m · [s; \lVert V_h \rVert]) \\ V' &= σ⁺(W_μ · V_h) \end{align}\](Eq.5) # Linear transformation on vectors
(Eq.6) # Scalars updated with vector norms
(Eq.7) # Vector-compatible nonlinearity
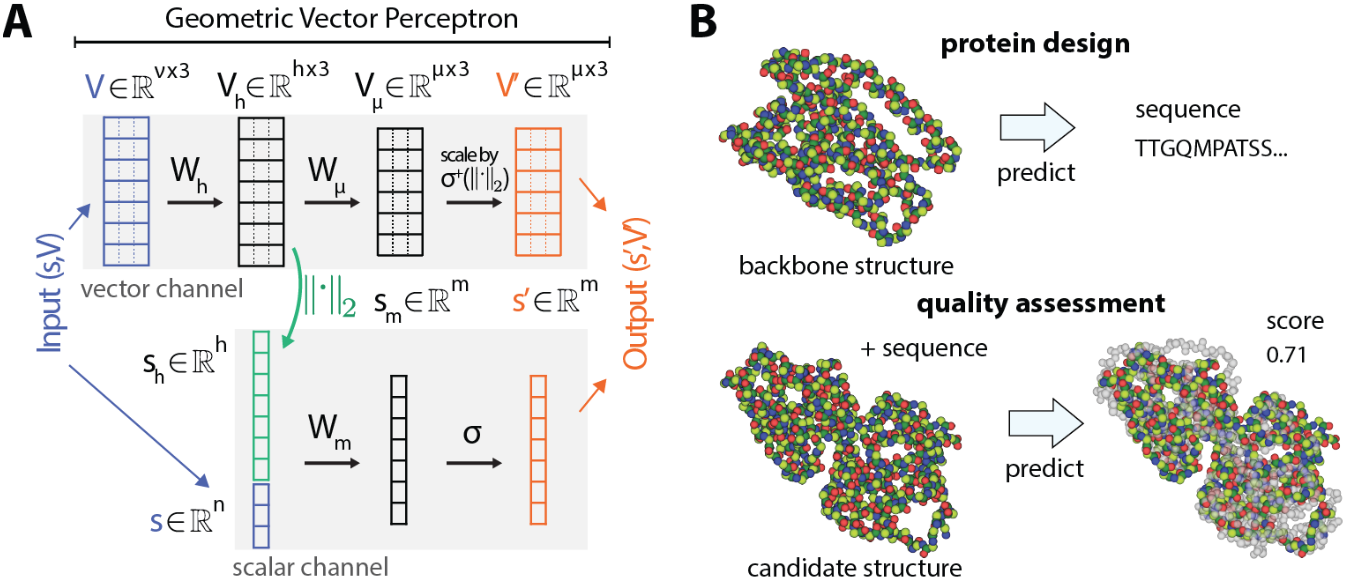
Figure 12: Geometric Vector Perceptron (GVP) layer. The architecture processes (scalar, vector) feature pairs. Key innovation: scalarization extracts invariant information from vector features via $\text{L2}$ norms $\lVert V \rVert$, enabling information flow from geometric to scalar features while maintaining equivariance. GVP provides a computationally efficient alternative to higher-order tensor methods for protein structure tasks. Source: Satorras et al. "$\text{E(n)}$ Equivariant Graph Neural Networks" ICML (2021), Figure 1.
GVP is computationally simple, scales well to large proteins, and integrates easily into GNN frameworks. The GVP-GNN variant serves as the structural encoder for ESM-IF1 and achieves strong results on protein model quality assessment and design benchmarks.
Frame averaging: equivariance without architectural constraints
Frame Averaging (Puny et al., ICLR 2022) offers an alternative paradigm: achieving equivariance through data transformation rather than architecture constraints. Given any backbone network φ, frame averaging constructs an equivariant version:
\[\langle φ \rangle_F(X) = \frac{1}{\vert F(X) \vert} \sum_{g ∈ F(X)} ρ_\mathrm{out}(g) · φ(ρ_\mathrm{in}(g)^{-1} · X)\]For $\text{E(3)}$, frames can be constructed via PCA of coordinates, yielding $\vert F \vert = 8$ distinct frames (from sign ambiguity of eigenvectors). The frame equivariance property $F(gX) = gF(X)$ ensures the averaged output is equivariant.
FAENet (Duval et al., ICML 2023) introduced stochastic frame averaging: sampling one random frame per training batch for approximate equivariance, with full averaging at inference. This achieves 5× speedup versus deterministic approaches while maintaining competitive accuracy on OC20.
The trade-off is clear: frame averaging enables maximal backbone expressivity at the cost of multiple forward passes. Built-in equivariance requires constrained architectures but needs only single passes. The choice depends on application requirements for speed versus flexibility.
Protein design: equivariance enables generation
RFdiffusion: diffusion on $\mathrm{SE(3)^N}$
RFdiffusion (Watson et al., Nature 2023) applies denoising diffusion to protein backbone generation. Each residue is represented as a frame $T_n = (r_n, x_n) ∈ \text{SE(3)}$, and the diffusion process adds noise to both translations (Gaussian noise in $ℝ^3$) and rotations (noise on $\text{SO(3)}$ via $\text{IGSO(3)}$ distribution).
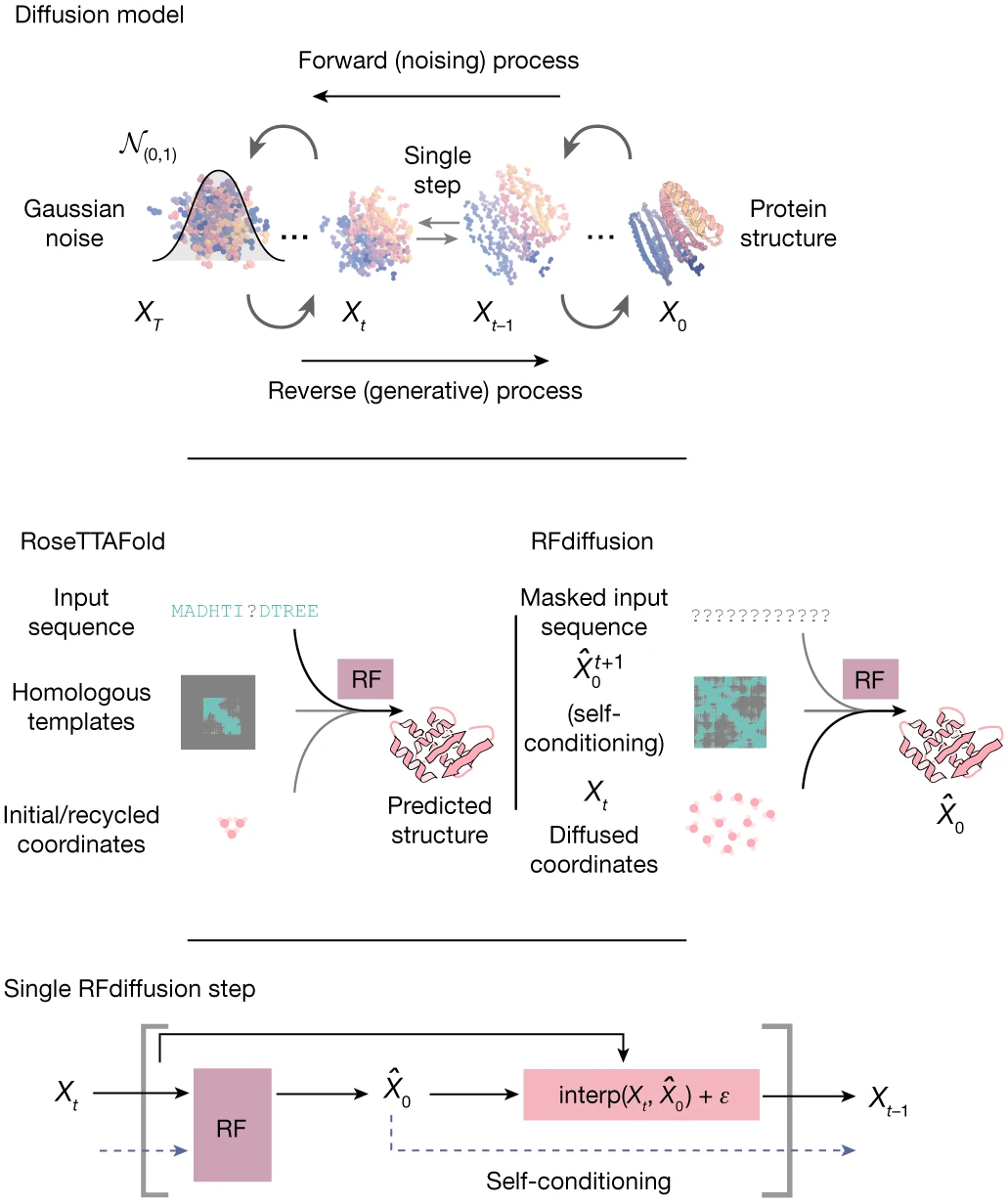
Figure 13: RFdiffusion training and generation framework. (Top) Diffusion models for proteins are trained to recover corrupted structures through iterative denoising. (Middle) The RoseTTAFold structure prediction network is fine-tuned into RFdiffusion with minimal changes—the primary input shifts from sequence to diffused residue frames. (Bottom) Self-conditioning: at each timestep $t$, the model receives its previous prediction $\hat X_0^{(t+1)}$ as template input, improving consistency across the trajectory. Source: Watson et al., Nature 2023, Figure 1a.
The denoiser is a fine-tuned RoseTTAFold structure prediction network, inheriting $\text{SE(3)}$ equivariance through IPA layers. Given noised backbones, it predicts the denoised structure, trained to minimize MSE between predictions and true structures.
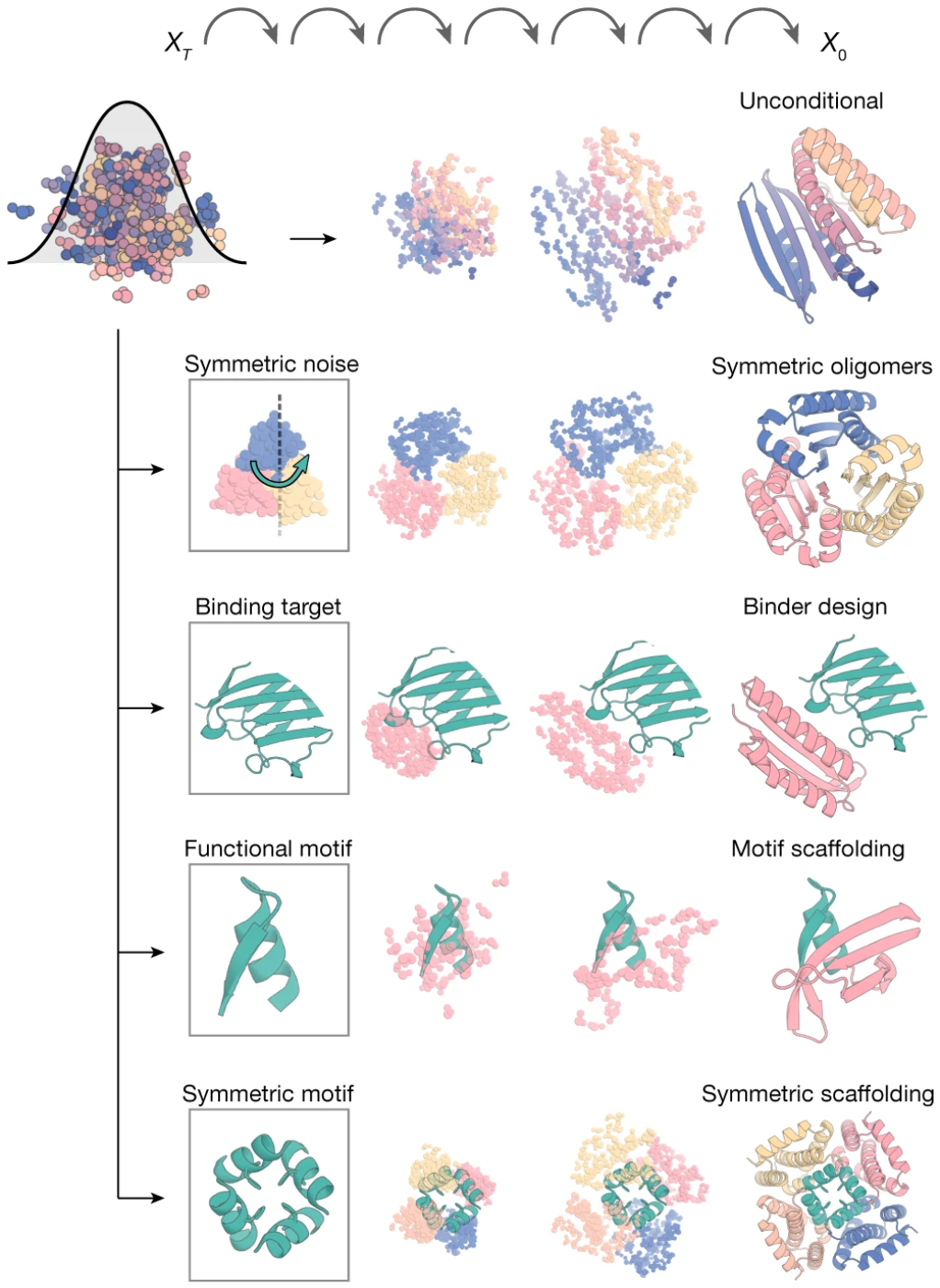
Figure 14: RFdiffusion's broad applicability through conditioning. Random noise is combined with different types of conditioning information: (top to bottom) unconditional generation for novel folds, symmetry specifications for oligomer design, binding targets for binder design, functional motifs for scaffolding, and symmetric motifs for therapeutic applications. In each case, RFdiffusion iteratively refines noise into designed protein structures satisfying the constraints. Source: Watson et al., Nature 2023, Figure 1b.
RFdiffusion’s conditional generation capabilities revolutionized protein design. Motif scaffolding constrains the diffusion to place functional sites in specific geometries. Binder design generated picomolar-affinity binders to targets like influenza hemagglutinin purely computationally. Symmetric design produces cyclic, dihedral, and higher-symmetry assemblies. Experimental validation confirmed designs with backbone RMSDs to predictions under 1Å.
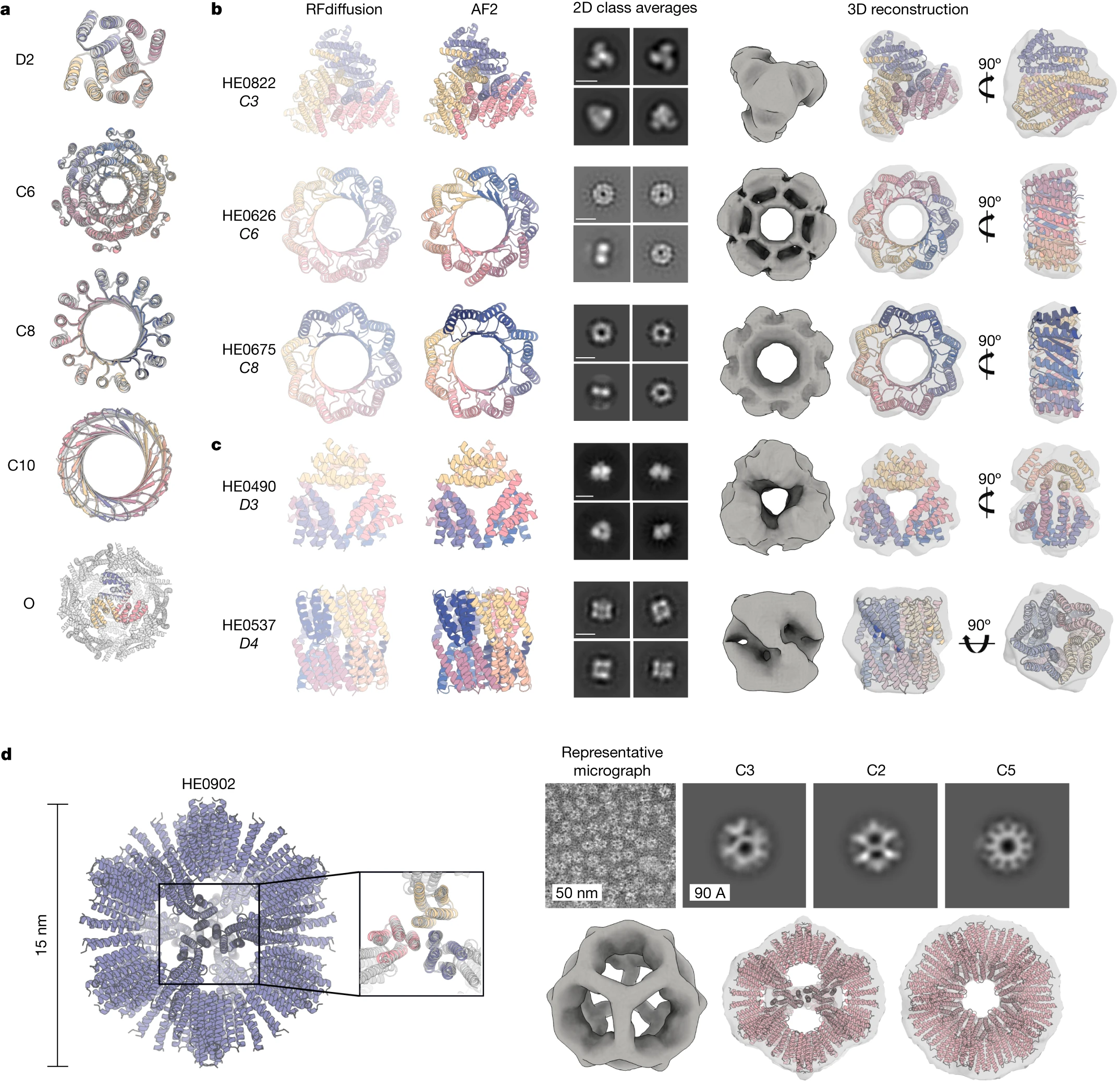
Figure 15: Experimental validation of RFdiffusion designs. (Top) Cryo-EM structure of designed binder in complex with influenza hemagglutinin, showing atomic-level agreement with computational design model (backbone RMSD < 1Å). (Bottom) Symmetric assemblies validated by electron microscopy, including cyclic oligomers with distinctive geometric features matching design predictions. Source: Watson et al., Nature 2023, Figure 3.
ProteinMPNN: invariant inverse folding at scale
ProteinMPNN (Dauparas et al., Science 2022) addresses the inverse folding problem: given a backbone structure, predict amino acid sequences that fold into it. The architecture is a message-passing neural network with 1.66 million parameters—remarkably compact.
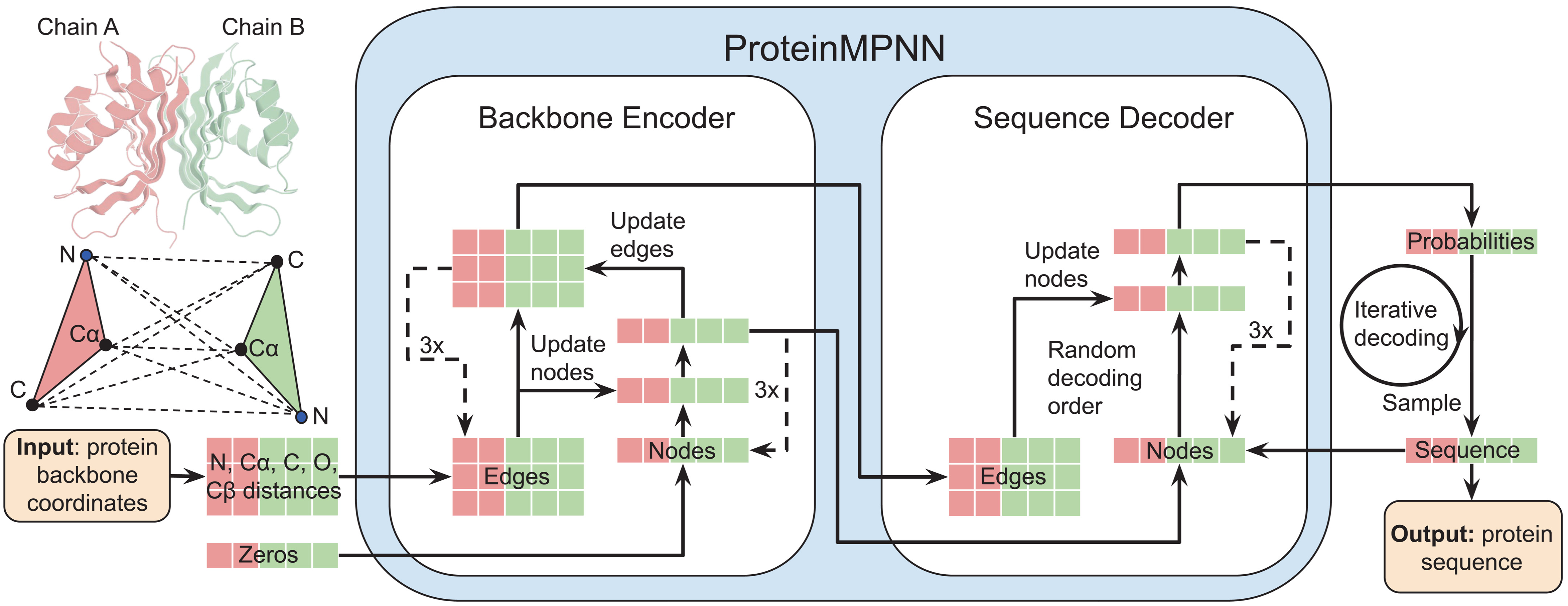
Figure 16: ProteinMPNN architecture for inverse folding. Interatomic distances between $\mathrm{N, Cα, C, O}$, and virtual $\mathrm{Cβ}$ atoms are encoded as edge features and processed through a message-passing neural network (Encoder). The encoded features, combined with partial sequences, generate amino acids iteratively in random decoding order, enabling flexible conditioning on sequence context. Source: Dauparas et al., Science 2022, Figure 1A.
Rather than explicit equivariance, ProteinMPNN uses distance-based invariance: edge features encode 25 interatomic distances per residue pair (between $\mathrm{N, Cα, C, O}$, and virtual $\mathrm{Cβ}$ atoms). These distances are inherently invariant to rotation and translation. Order-agnostic autoregressive decoding, trained with random decoding order permutations, enables flexible design scenarios.
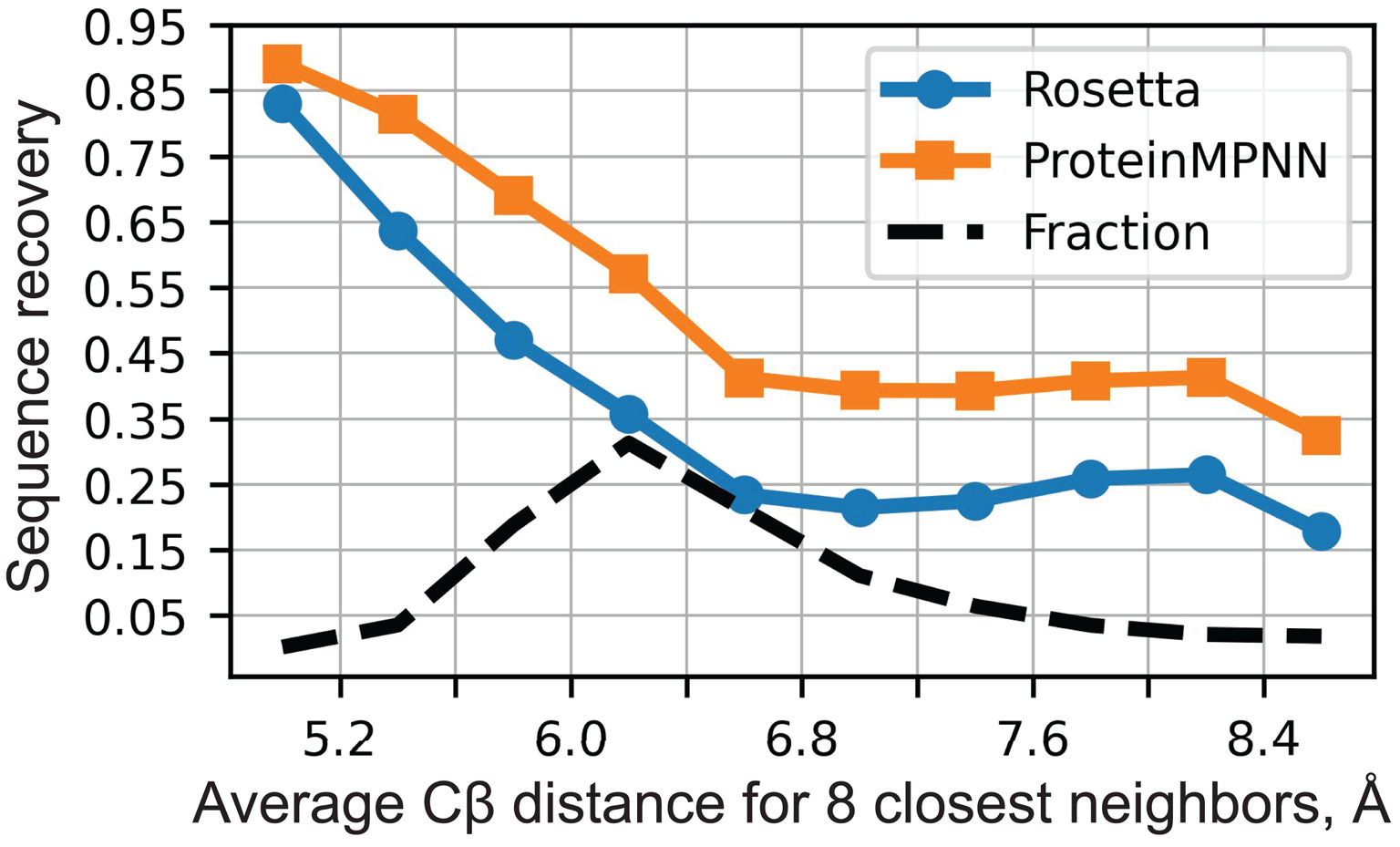
Figure 17: ProteinMPNN native sequence recovery performance. Sequence recovery as a function of residue burial (measured by average $\text{Cβ}$ distance to 8 nearest neighbors). ProteinMPNN achieves 52.4% overall recovery compared to Rosetta's 32.9%, with improvements across all burial levels from protein core to surface. Despite being 200× faster (1.2s vs 258s per 100 residues), ProteinMPNN dramatically outperforms the physics-based baseline. Source: Dauparas et al., Science 2022, Figure 2A.
Performance gains are dramatic: 52.4% sequence recovery versus 32.9% for Rosetta, with inference 200× faster (1.2 seconds per 100 residues versus 258 seconds). Variants include LigandMPNN (2.62M parameters) for ligand-aware design (63.3% recovery near small molecules, 77.5% near metals) and SolubleMPNN for solubility-optimized sequences.
ESM-IF1: language models meet geometric encoders
ESM-IF1 (Hsu et al., ICML 2022) combines a GVP-GNN structural encoder with a Transformer decoder, totaling 142 million parameters—100× larger than ProteinMPNN. Crucially, it was trained on 12 million AlphaFold2-predicted structures, 750× more than the experimental structures in PDB.
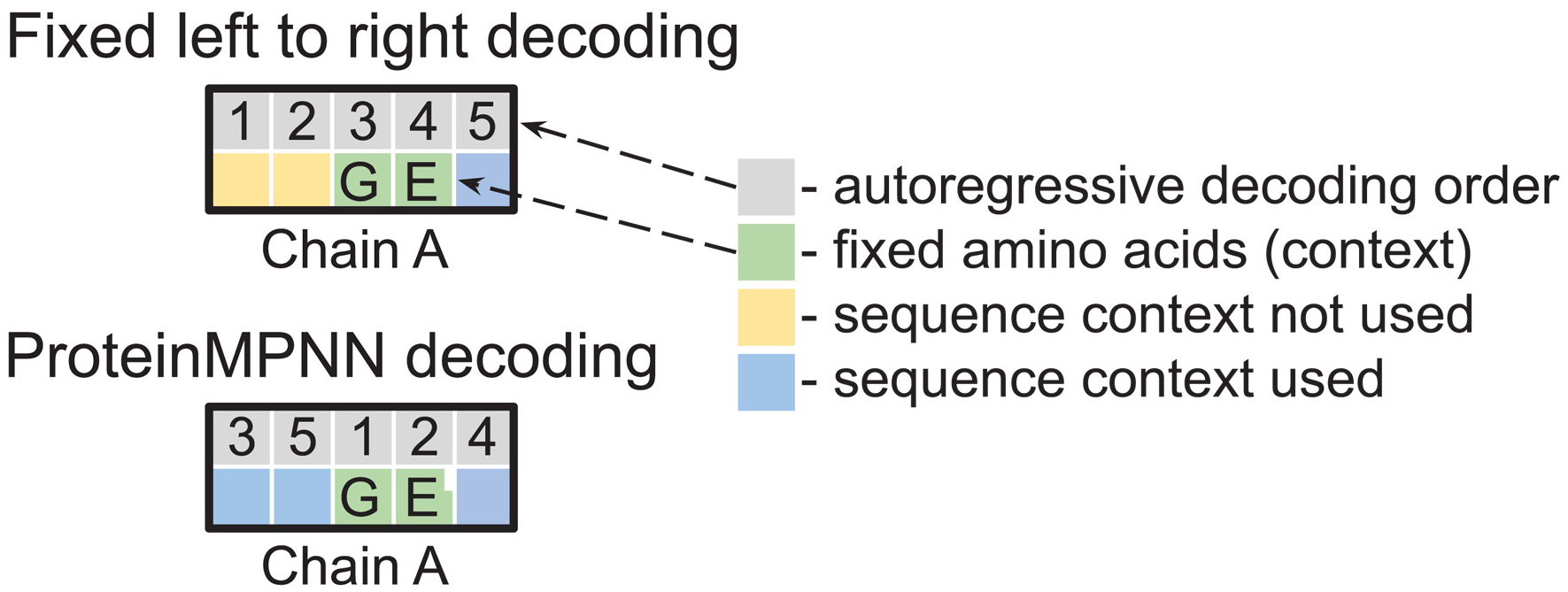
Figure 18: Random decoding order enables flexible conditioning. (Left) Fixed left-to-right decoding cannot use sequence context (green) for earlier positions (yellow). (Right) Training with random decoding orders allows arbitrary inference-time ordering, enabling designs where fixed context regions are decoded first, critical for motif scaffolding and multistate design applications. Source: Dauparas et al., Science 2022, Figure 1B.
The GVP encoder provides rotation-equivariant structural representations, while the Transformer decoder performs autoregressive sequence generation. Performance is comparable to ProteinMPNN (51% sequence recovery) despite different training data and architecture, validating both approaches.
Emerging generative approaches
Chroma (Ingraham et al., Nature 2023) introduced programmable protein generation with composable “conditioners” for constraints: symmetry, substructure, shape, and even natural language prompts via CLIP-like models. The architecture uses equivariant GNN layers with sub-quadratic scaling, enabling generation of 4,000+ residue complexes.
FrameDiff (Yim et al., ICML 2023) and FoldFlow (Bose et al., ICLR 2024) provide principled theoretical frameworks for $\text{SE(3)}$ diffusion and flow matching respectively. FrameDiff’s 17.4M parameter model trains from scratch without pretrained structure prediction, while FoldFlow’s Riemannian optimal transport constructs simpler, more stable flows.
Genie 2 (Lin et al., 2024) scales training to 588,571 AlphaFold Database structures, achieving higher diversity and novelty than RFdiffusion while matching designability. Multi-motif scaffolding enables designing proteins with multiple co-occurring functional sites.
Historical evolution: from constraints to capabilities
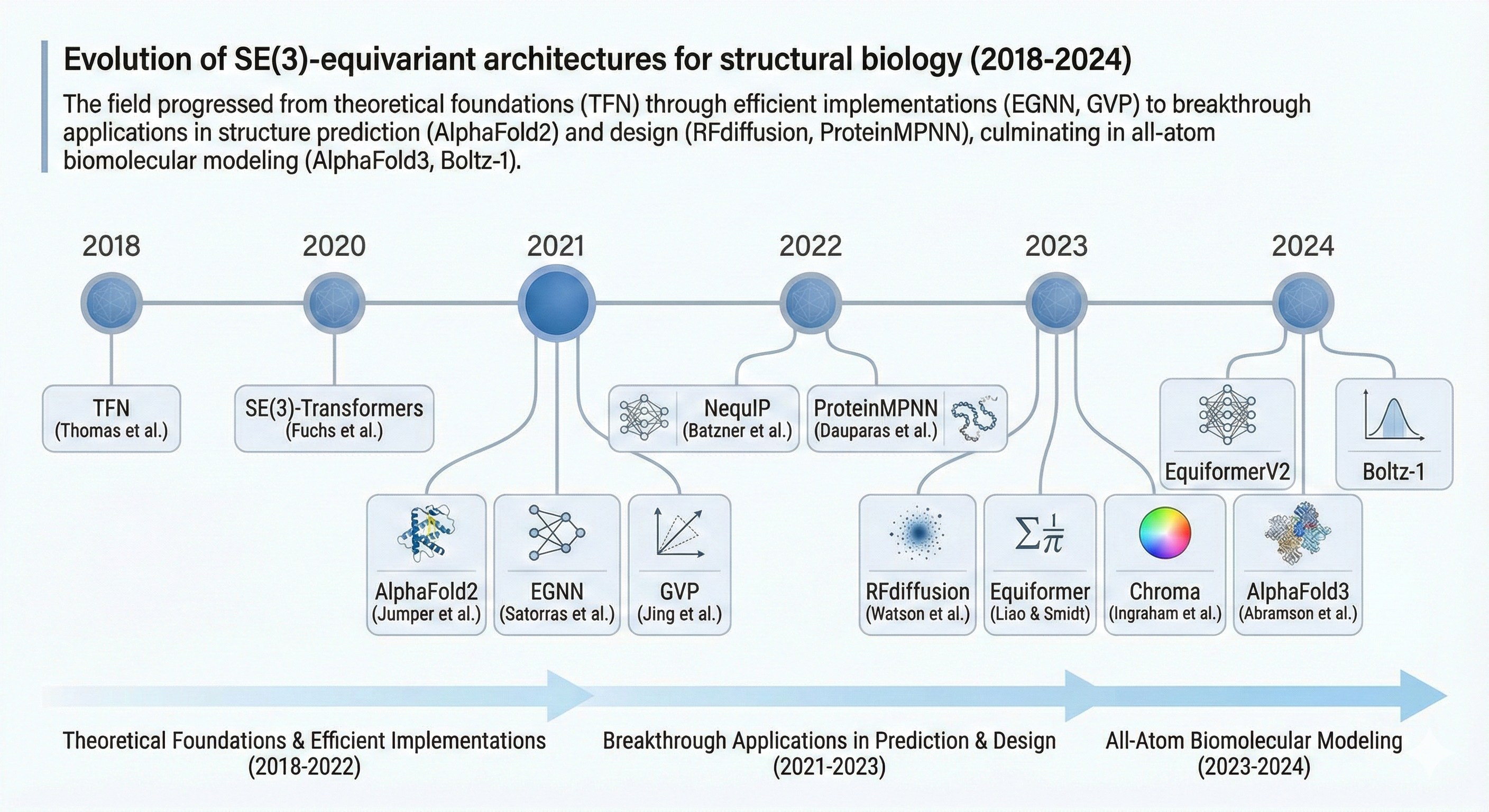
Figure 19: Evolution of $\text{SE(3)}$-equivariant architectures for structural biology (2018-2024). The field progressed from theoretical foundations (TFN) through efficient implementations (EGNN, GVP) to breakthrough applications in structure prediction (AlphaFold2) and design (RFdiffusion, ProteinMPNN), culminating in all-atom biomolecular modeling (AlphaFold3, Boltz-1).
The pre-equivariant era (2017-2019)
Early molecular GNNs like SchNet used continuous-filter convolutions on invariant features (interatomic distances). While successful for molecular property prediction, these approaches discarded directional information—forces, for example, require vector outputs that invariant networks cannot directly produce.
Tensor Field Networks (2018) and Cormorant (2019) established the spherical harmonic framework, demonstrating that neural networks could maintain full geometric information while respecting symmetries. However, computational costs limited practical adoption.
The 2021 inflection point
Three parallel developments transformed the field. AlphaFold2 demonstrated that equivariant attention (IPA) enables atomic-level accuracy in protein structure prediction, achieving median GDT-TS scores above 90 on CASP14 targets—accuracy previously thought decades away.
EGNN showed that simple, efficient equivariance was possible without heavy mathematical machinery, enabling practical adoption across the community. GVP provided protein-specific equivariant layers that integrated naturally into existing pipelines.
The recognition that equivariance provides data efficiency rather than just mathematical elegance—crystallized by NequIP’s 1000× improvement—shifted the field’s understanding of why these constraints matter.
The design era (2023-present)
RFdiffusion’s success demonstrated that equivariant architectures enable not just prediction but generation of novel functional proteins. The shift from understanding natural proteins to designing synthetic ones represents a qualitative capability change enabled by equivariant representations.
AlphaFold3 (2024) and alternatives like Boltz-1 (2024, fully open-source, MIT license) extended prediction to all-atom biomolecular complexes—proteins, nucleic acids, small molecules, ions, modified residues. Interestingly, AlphaFold3’s removal of explicit equivariance in favor of diffusion raises questions about whether architectural constraints or implicit learning from massive data is the path forward.
Practical considerations for implementation
Choosing the right architecture
For protein structure prediction with good MSAs, AlphaFold2/OpenFold remains gold standard. For single-sequence prediction (orphan proteins, high-throughput screening), ESMFold provides 60× faster inference. AlphaFold3 or Boltz-1 is necessary for protein-ligand complexes or nucleic acid interactions.
For molecular property prediction, EquiformerV2 provides state-of-the-art accuracy on catalyst and materials benchmarks. MACE offers excellent accuracy with efficient two-layer architecture. For simpler applications or prototyping, EGNN provides strong baseline performance with minimal complexity.
For protein design, RFdiffusion handles unconditional generation, motif scaffolding, and binder design. ProteinMPNN (or LigandMPNN for ligand contexts) performs inverse folding. Chroma enables text-conditioned and shape-constrained design.
Computational trade-offs
Spherical harmonic methods face $\mathrm{O(L^6)}$ scaling with maximum degree, practically limiting $\mathrm{\ell_{max}}$ to 3-4 without eSCN-style optimizations. Frame-based methods (IPA) have $\mathrm{O(L^2)}$ memory in sequence length, addressable through FlashAttention variants. Frame averaging multiplies computational cost by number of frames (typically 8 for $\mathrm{E(3)}$).
For large-scale deployment, inference latency matters. ESMFold: seconds for typical proteins. AlphaFold2: minutes to hours depending on MSA depth. ProteinMPNN: ~1 second per 100 residues. These timescales enable different screening strategies.
Available libraries and frameworks
e3nn (PyTorch/JAX) provides general $\text{E(3)}$-equivariant neural network primitives—tensor products, spherical harmonics, irreps handling. PyTorch Geometric offers message-passing infrastructure with equivariant layer integration. OpenFold enables AlphaFold2 experimentation and modification. TorchMD-NET provides ready-to-use equivariant transformers for molecular potentials.
Future directions and open challenges
Toward all-atom and multi-modal modeling
The field is moving from coarse-grained (residue-level) to all-atom representations. AlphaFold3, RoseTTAFold All-Atom, and Boltz-2 operate at atomic resolution, enabling modeling of ligands, post-translational modifications, and non-standard residues. Glycosylation—present on 50-70% of human proteins—remains challenging due to flexibility and diversity.
Dynamics and conformational ensembles
Current methods predict static structures, but proteins are dynamic. Boltz-2 incorporates molecular dynamics ensemble data during training, beginning to capture conformational flexibility. True ensemble prediction—representing the Boltzmann distribution over conformations—remains an open challenge with profound implications for understanding function and designing therapeutics.
Scaling laws and the equivariance question
Does explicit equivariance remain necessary at sufficient scale? AlphaFold3’s diffusion-based approach suggests large models may learn symmetries implicitly. However, data efficiency arguments—relevant when training data is expensive to generate—favor explicit equivariance. The field continues to debate whether equivariance is a fundamental design principle or a useful inductive bias that massive data can overcome.
Conclusion: symmetry as scientific insight
The integration of $\text{SE(3)}$ and $\text{E(3)}$ equivariance into neural network architectures for computational structural biology represents more than a technical advance—it embodies a scientific principle. Physical reality respects symmetries, and our computational tools should too.
From AlphaFold2’s frame-based Invariant Point Attention to Equiformer’s spherical harmonic transformers to EGNN’s elegant simplicity, the field has developed a rich toolkit for building symmetry into neural architectures. The practical impact is undeniable: 1000-fold improvements in data efficiency, atomic-level accuracy in structure prediction, and the ability to design functional proteins computationally.
The path forward will likely blend explicit equivariance with learned representations, all-atom modeling with coarse-grained efficiency, and static prediction with dynamic understanding. What remains constant is the insight that encoding physical symmetries into computational architectures—rather than hoping models learn them from data—accelerates scientific discovery in structural biology.
References for Further Reading
Foundational Theory:
- Bronstein, Bruna, Cohen, Veličković. “Geometric Deep Learning: Grids, Groups, Graphs, Geodesics, and Gauges.” arXiv:2104.13478 (2021)
- Cohen & Welling. “Group Equivariant Convolutional Networks.” ICML (2016)
- Thomas et al. “Tensor Field Networks.” arXiv:1802.08219 (2018)
Structure Prediction:
- Jumper et al. “Highly accurate protein structure prediction with AlphaFold.” Nature 596:583-589 (2021)
- Abramson et al. “Accurate structure prediction of biomolecular interactions with AlphaFold 3.” Nature 630:493-500 (2024)
Equivariant Architectures:
- Fuchs et al. “SE(3)-Transformers.” NeurIPS (2020), arXiv:2006.10503
- Satorras et al. “E(n) Equivariant Graph Neural Networks.” ICML (2021), arXiv:2102.09844
- Jing et al. “Learning from Protein Structure with Geometric Vector Perceptrons.” ICLR (2021), arXiv:2009.01411
- Liao & Smidt. “Equiformer.” ICLR (2023), arXiv:2206.11990
- Liao et al. “EquiformerV2.” ICLR (2024), arXiv:2306.12059
Protein Design:
- Watson et al. “De novo design of protein structure and function with RFdiffusion.” Nature 620:1089-1100 (2023)
- Dauparas et al. “Robust deep learning-based protein sequence design using ProteinMPNN.” Science 378:49-56 (2022)
- Ingraham et al. “Illuminating protein space with a programmable generative model.” Nature 623:1070-1078 (2023)
Data Efficiency:
- Batzner et al. “E(3)-equivariant graph neural networks for data-efficient interatomic potentials.” Nature Communications (2022)
Recent Advances:
- Wohlwend, Corso et al. “Boltz-1: Democratizing Biomolecular Interaction Modeling.” arXiv (2024)
- Yim et al. “SE(3) diffusion model with application to protein backbone generation.” ICML (2023), arXiv:2302.02277
- Bose et al. “SE(3)-Stochastic Flow Matching for Protein Backbone Generation.” ICLR (2024), arXiv:2310.02391
Enjoy Reading This Article?
Here are some more articles you might like to read next: