Protein generation with evolutionary diffusion (Evodiff)
How discrete diffusion on sequences unlocks protein design beyond structure
Sequence-based diffusion models can design proteins that structure-based methods fundamentally cannot access. EvoDiff, developed by Alamdari et al. at Microsoft Research, introduces discrete diffusion directly on amino acid sequences, enabling generation of intrinsically disordered regions, achieving better coverage of natural functional space, and providing native support for conditional generation—all without requiring structural information. This represents a paradigm shift from structure-centric approaches like RFdiffusion toward leveraging the full diversity of 42 million evolutionary sequences rather than the ~200,000 structures in the PDB.
The key insight is deceptively simple: every protein is defined first by its sequence. Structure follows, but not every protein folds into a rigid 3D shape. By operating in sequence space with order-agnostic discrete diffusion, EvoDiff accesses design spaces invisible to structure-based methods while capturing evolutionary patterns that encode function.
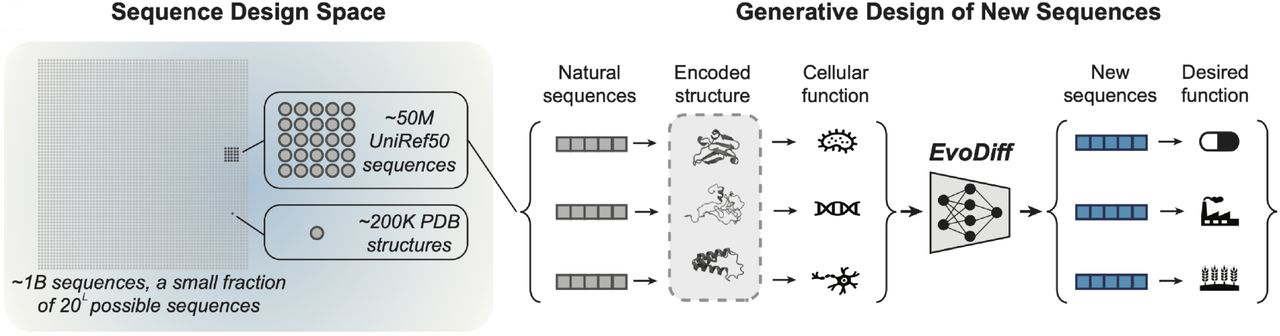
Figure 1: The sequence design space vastly exceeds the structure space. Evolution has sampled ~50 million sequences (UniRef50) from billions of possible sequences, while experimental structures represent only ~200,000 proteins in the PDB. EvoDiff trains on evolutionary-scale sequence data to generate new protein sequences that fold and perform desired functions, accessing proteins that structure-based methods cannot design. Source: Alamdari et al., bioRxiv 2024, Figure 1A.
Discrete diffusion replaces Gaussian noise with token corruption
Structure-based diffusion models like RFdiffusion apply continuous Gaussian noise to 3D coordinates, gradually corrupting atomic positions into random configurations before learning to reverse the process. EvoDiff takes a fundamentally different approach: it operates on discrete amino acid tokens using Markov chains that progressively corrupt sequences through masking or substitution.
The paper introduces two discrete diffusion frameworks. Order-Agnostic Autoregressive Diffusion (OADM) treats each corruption step as masking a single amino acid with a special [MASK] token. After $T = L$ steps (where $L$ is sequence length), the entire sequence becomes masked. The model learns to predict any masked position given arbitrary context, enabling generation in any order—a crucial capability for conditional tasks like motif scaffolding.
The OADM training objective maximizes a reweighted evidence lower bound:
\[\mathcal{L}_t = \frac{1}{L - t + 1} \mathbb{E}_{\sigma \sim \mathcal{U}(S_L)} \sum_{k \in \sigma(\geq t)} \log p(x_k | x_{\sigma(\lt t)})\]This generalizes both left-to-right autoregressive models and masked language models. The timestep is implicitly encoded by the number of masked positions, requiring no explicit timestep embedding.
D3PM (Discrete Denoising Diffusion Probabilistic Models) provides an alternative framework using transition matrices that govern corruption probabilities. The forward process follows:
\[q(x_t | x_{t-1}) = \text{Cat}(x_t; p = x_{t-1} Q_t)\]where $Q$ is a Markov transition matrix. EvoDiff explores two corruption schemes: D3PM-Uniform with a doubly stochastic uniform transition matrix, and D3PM-BLOSUM using biologically-informed transition probabilities derived from BLOSUM62 amino acid substitution frequencies. The BLOSUM variant prioritizes evolutionarily conserved relationships—if histidine mutates, it’s more likely to become asparagine than tryptophan, matching natural evolutionary patterns.
The reverse process combines the learned denoiser with the posterior:
\[p_\theta(x_{t-1} | x_t) \propto \sum_{\tilde{x}_0} q(x_{t-1}, x_t | \tilde{x}_0) \tilde{p}_\theta(\tilde{x}_0 | x_t)\]A hybrid loss $L_λ = L_{vb} + λL_{ce}$ combines the variational lower bound with cross-entropy, though experiments use $λ = 0$ with $T = 500$ timesteps.
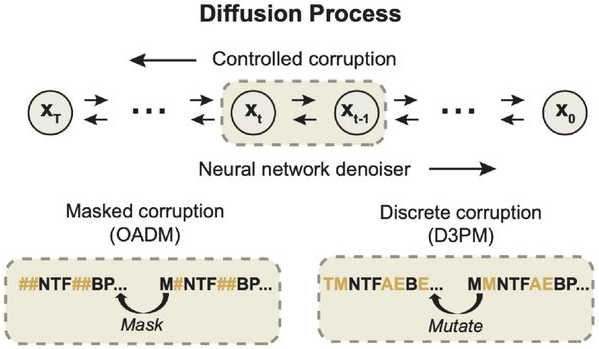
Figure 2: Discrete diffusion processes for protein sequences. In order-agnostic autoregressive diffusion (OADM, left), tokens are progressively masked in random order, with the model learning to predict masked positions given arbitrary context. In discrete denoising diffusion (D3PM, right), amino acids are mutated according to a transition matrix (uniform or BLOSUM62-based), with the model learning to reverse these mutations. Both approaches use neural networks to parameterize the denoising process. Source: Alamdari et al., bioRxiv 2024, Figure 1B.
Architecture builds on convolutional and transformer foundations
EvoDiff deploys two model families targeting different use cases. EvoDiff-Seq models use a ByteNet architecture—a dilated convolutional neural network that scales linearly with sequence length. The 38M parameter small model uses 16 layers with dimension 1020, while the 640M parameter large model expands to 56 layers with dimension 1280. For D3PM variants, 1D sinusoidal positional embeddings encode the diffusion timestep; OADM models encode timestep implicitly through mask count.
EvoDiff-MSA models employ the MSA Transformer architecture with 100M parameters, processing multiple sequence alignments of 512 residues × 64 sequences. Two subsampling strategies are evaluated: “Random” sampling (always including the query) and “MaxHamming” which greedily maximizes diversity via Hamming distance selection.
Training leverages massive evolutionary data. The sequence models train on UniRef50 (~42 million sequences) with maximum length 1024 residues. MSA models use the OpenFold dataset containing 382,296 filtered MSAs derived from 140,000 unique PDB chains and 16 million UniClust30 clusters. Training the 640M sequence model requires 32 V100 GPUs for 12-23 days, processing up to $10^{17}$ tokens over 2 million steps with Adam optimization (learning rate 1e-4, 16,000 warmup steps).
Order-agnostic generation enables natural conditional design
The OADM framework’s most powerful feature is native conditional generation. Because the model trains to predict masked positions given arbitrary context—not just left-to-right—conditioning requires no fine-tuning or architectural changes. For motif scaffolding, simply fix the motif residues as unmasked context and generate the surrounding scaffold. For inpainting disordered regions, mask only those positions and let the model fill them in.
This contrasts sharply with structure-based methods. RFdiffusion requires task-specific fine-tuning on examples mimicking the desired conditioning. ProteinMPNN needs an input backbone structure. EvoDiff’s approach mirrors how BERT-style models handle text infilling, but extends it through the diffusion framework’s iterative refinement.
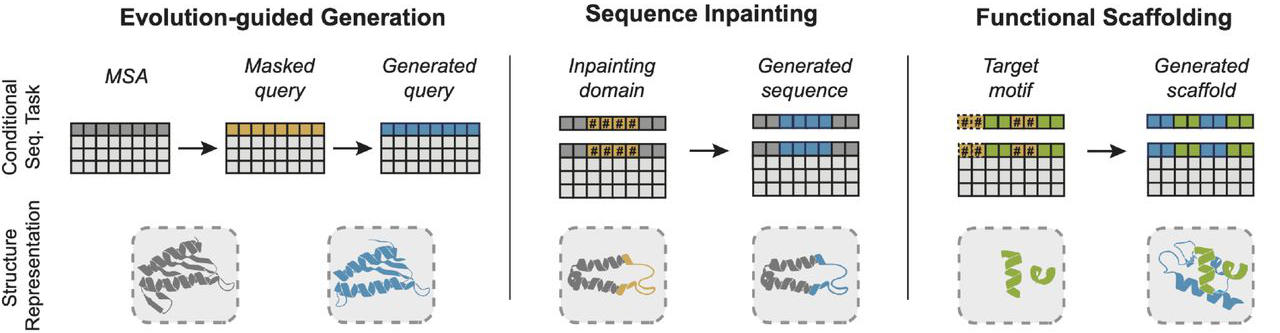
Figure 3: Conditional protein design with EvoDiff via order-agnostic generation. (Left) Evolution-guided generation: Generate novel query sequences conditioned on MSA context, producing family members without task-specific training. (Middle) Sequence inpainting: Fix functional domains (colored) and generate surrounding regions (blue), enabling design of proteins with specified functional modules. (Right) Functional scaffolding: Condition on structural motif sequences (green) to generate scaffolds (blue) without explicit structural information. Source: Alamdari et al., bioRxiv 2024, Figure 1D.
The paper demonstrates three conditioning scenarios:
- Evolution-guided generation: Condition on MSA context to generate novel query sequences maintaining family characteristics
- Inpainting: Fix functional domains or flanking regions, generate the remainder
- Scaffolding: Fix a structural motif’s sequence, generate surrounding scaffold without explicit structure
Intrinsically disordered regions become designable
Perhaps EvoDiff’s most significant capability is generating intrinsically disordered regions (IDRs)—protein segments lacking stable 3D structure that nonetheless perform critical biological functions. Approximately 30% of eukaryotic proteins contain substantial disorder, yet structure-based methods fundamentally cannot design these regions because they have no structure to diffuse toward.
The validation uses 15,996 human IDRs with orthologs from the OMA database. EvoDiff masks IDR regions within their sequence context and generates replacements via inpainting. Evaluation employs DR-BERT, a disorder predictor outputting per-residue scores from 0-1. Generated IDRs achieve disorder score distributions matching natural sequences, demonstrating successful capture of the sequence patterns that encode disorder.
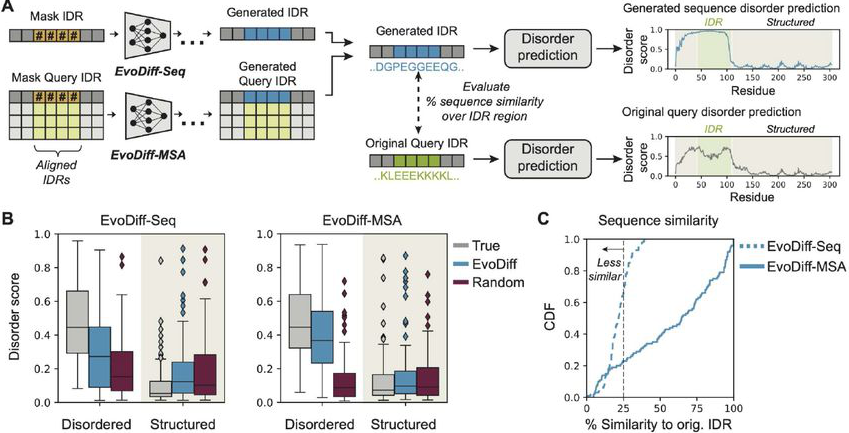
Figure 4: EvoDiff generates functional intrinsically disordered regions. (A) Workflow: Mask IDR positions within their sequence context, generate using EvoDiff-Seq or EvoDiff-MSA, and evaluate disorder scores using DR-BERT. (B) Generated IDRs (blue) match the disorder score distributions of natural sequences (gray) in both disordered and structured regions, while random sequences (red) fail to capture these patterns. EvoDiff successfully generates sequences encoding disorder—a capability inaccessible to structure-based methods. (C) Distribution of sequence similarity relative to the original IDR for generated IDRs from EvoDiff-Seq (blue, dashed) and EvoDiff-MSA (blue, solid) (n=100; dashed line at 25%). Source: Alamdari et al., bioRxiv 2024, Figure 5A-C.
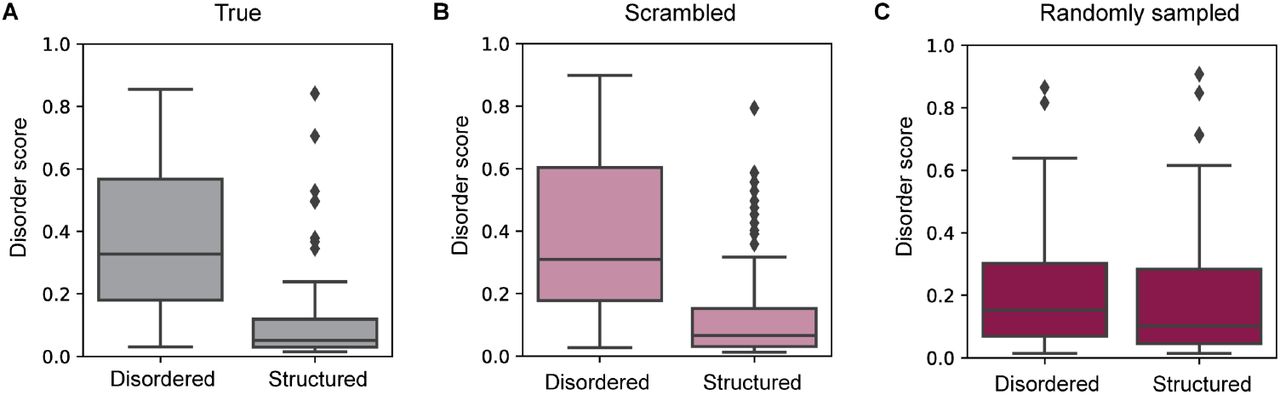
Figure 5: Validation of DR-BERT disorder predictor. The predictor correctly assigns high disorder scores to disordered regions and low scores to structured regions in natural proteins (A), while scrambled (B) and randomly sampled (C) sequences show degraded discrimination. This validates DR-BERT as an appropriate metric for evaluating EvoDiff's generated IDRs. Source: Alamdari et al., bioRxiv 2024, Figure S11.
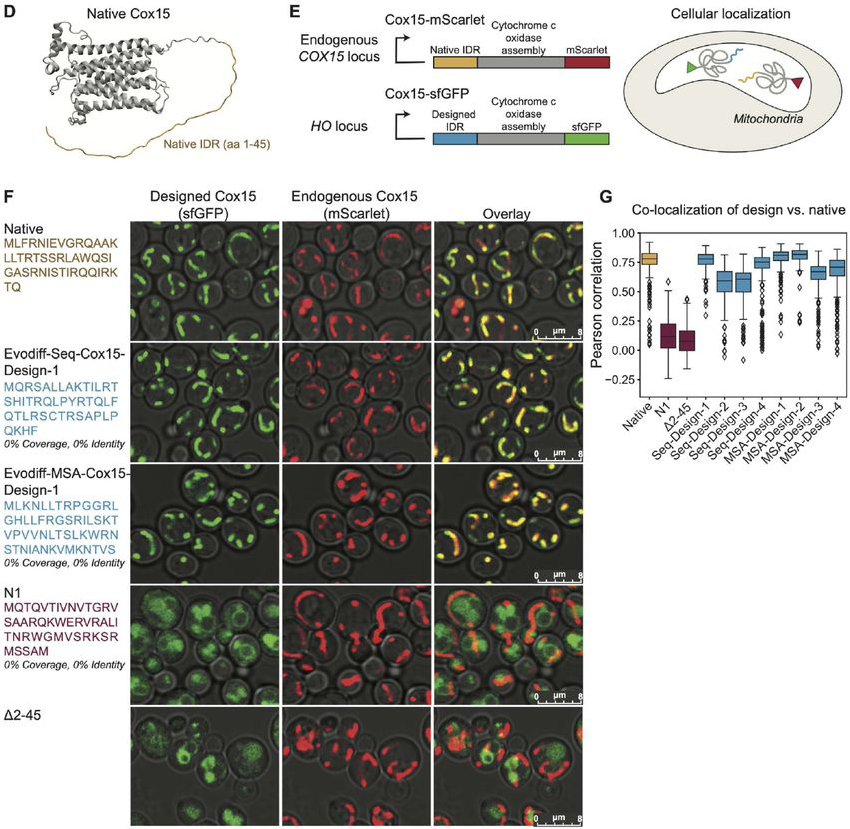
Figure 6: Experimental validation of EvoDiff-designed mitochondrial targeting signals. (D) The yeast Cox15 protein contains an N-terminal disordered IDR (yellow) that targets the protein to mitochondria. (E) Assay design: Cox15 with designed IDRs (sfGFP-tagged) should co-localize with endogenous Cox15 (mScarlet-tagged) if the designed targeting signal functions. (F) Microscopy images show designed IDRs (EvoDiff-Seq-Cox15-Design-1 and EvoDiff-MSA-Cox15-Design-1) achieve mitochondrial localization matching the native sequence. Negative controls (N1, Δ2-45) show diffuse cytoplasmic signal. (G) Quantification confirms all 8 EvoDiff designs achieve co-localization comparable to native Cox15. Source: Alamdari et al., bioRxiv 2024, Figure 5D-G.
This capability has immediate practical applications. Mitochondrial targeting signals, nuclear localization sequences, and many regulatory regions are intrinsically disordered. Structure-based tools simply cannot access this design space.
Benchmarks reveal complementary strengths with structure methods
EvoDiff evaluation spans unconditional generation quality, structural plausibility, functional coverage, and experimental validation.
Perplexity on the test set shows OADM substantially outperforms D3PM variants, and critically, is the only variant where performance scales with model size (640M: 13.05 vs 38M: 14.61). D3PM models show minimal scaling, suggesting the simpler masking approach better captures protein sequence patterns.
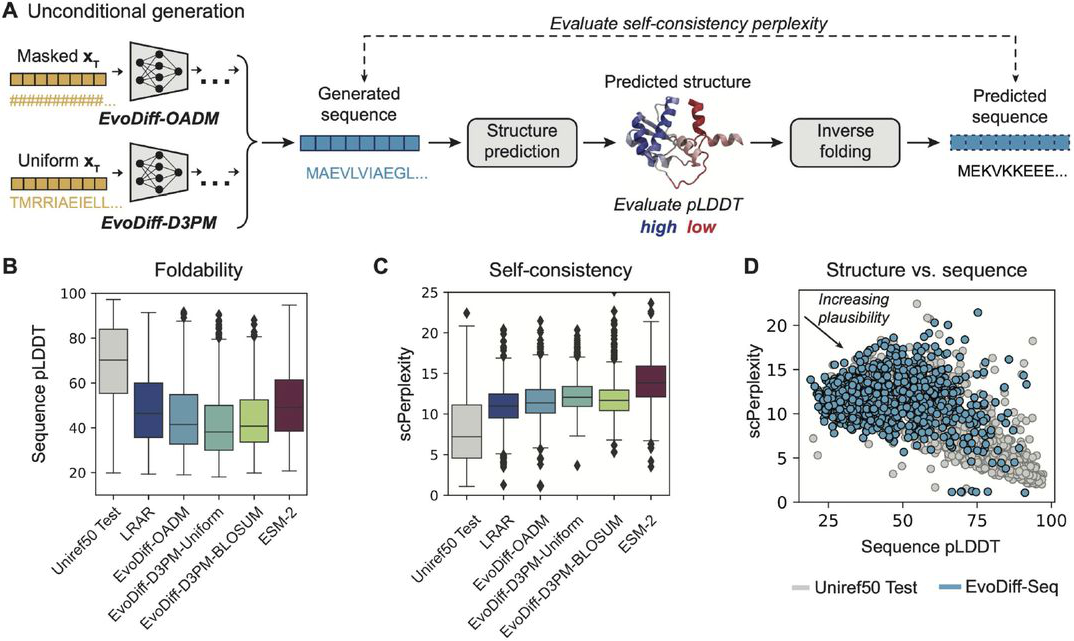
Figure 7: Structural plausibility of EvoDiff-generated sequences. (A) Evaluation pipeline: Generated sequences are folded with OmegaFold (pLDDT measures confidence), then inverse-folded with ESM-IF (scPerplexity measures sequence-structure consistency). (B-C) EvoDiff-OADM models generate more foldable and self-consistent sequences than D3PM variants or baselines. (D) EvoDiff-Seq sequences (blue) show plausible pLDDT-scPerplexity relationships, though not matching natural proteins (gray), indicating they encode foldable structures. Source: Alamdari et al., bioRxiv 2024, Figure 2A-D.
Fréchet ProtT5 Distance (FPD) measures coverage of natural sequence/functional space using ProtT5 embeddings. EvoDiff-Seq achieves FPD = 0.88, substantially outperforming RFdiffusion (1.96) and ESM-2 (2.81). This indicates EvoDiff-generated sequences better span the diversity of natural proteins.
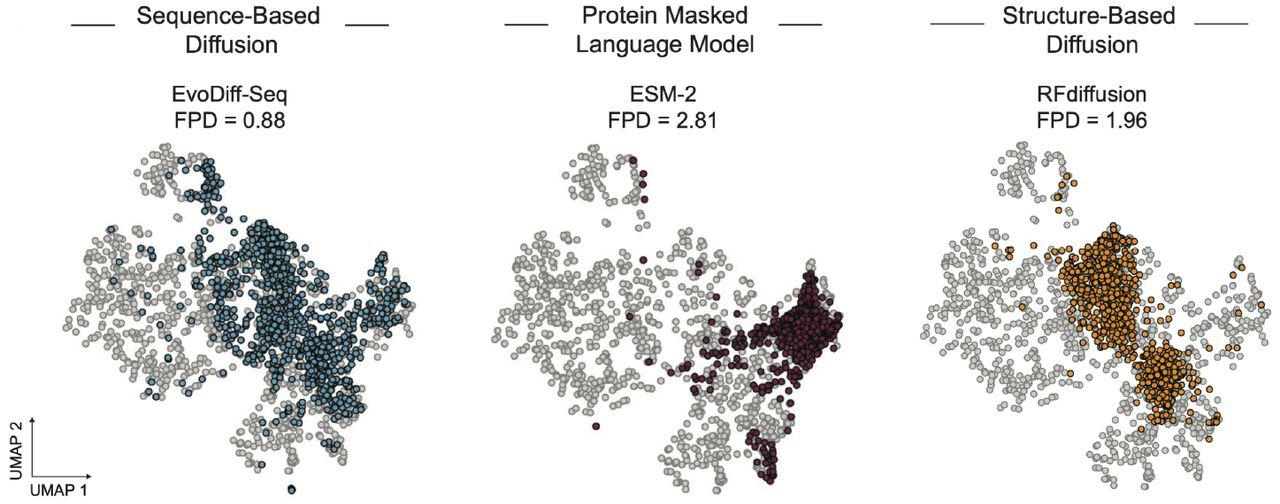
Figure 8: EvoDiff generates sequences covering natural functional space. UMAP visualization of ProtT5 embeddings shows EvoDiff-Seq (blue, FPD=0.88) better spans natural sequence diversity (gray) compared to ESM-2 (FPD=2.81) or RFdiffusion (FPD=1.96). Lower FPD indicates better coverage of the natural sequence and functional landscape captured in protein language model embeddings. Source: Alamdari et al., bioRxiv 2024, Figure 3B.
Structural plausibility uses OmegaFold predictions evaluated by pLDDT (predicted local distance difference test) and self-consistency perplexity via ESM-IF inverse folding. EvoDiff-OADM 640M achieves mean pLDDT of 44.46 with scPerplexity of 11.53—lower than the test set (68.25 pLDDT, 8.04 scPerplexity) but substantially better than random sequences (39.97 pLDDT, 14.68 scPerplexity).
Secondary structure distributions reveal a critical finding. EvoDiff generates proportions of β-strands and disordered regions matching natural proteins (KL divergence
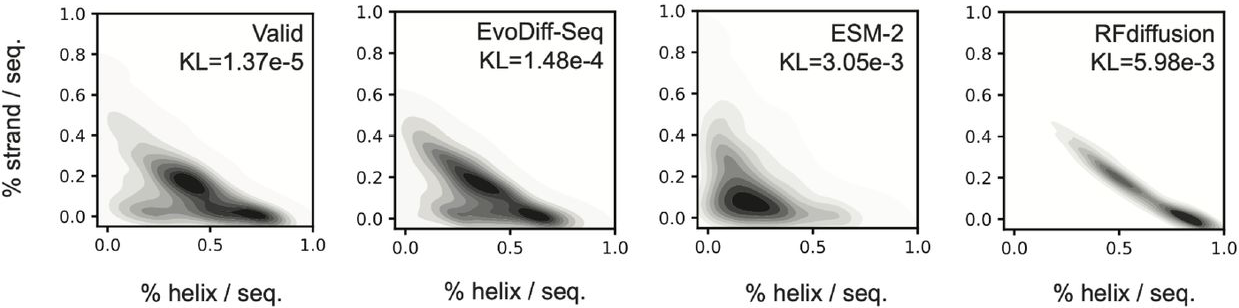
Figure 9: EvoDiff captures natural secondary structure distributions. Predicted secondary structure content (helix vs. strand fractions) shows EvoDiff-Seq (KL=1.48e-4) closely matches natural proteins (Valid), while ESM-2 (KL=3.05e-3) and RFdiffusion (KL=5.98e-3) are biased toward helix-rich structures. This reflects RFdiffusion's training on well-folded PDB structures and ESM-2's exposure primarily to structured protein domains. Source: Alamdari et al., bioRxiv 2024, Figure 3A.
Scaffolding benchmarks across 17 structural motifs (100 trials each) show EvoDiff-MSA achieving 522 successes versus RFdiffusion’s 610, using success criteria of pLDDT ≥ 70 and motif RMSD ≤ 1.0Å. Notably, success rates show almost no correlation between methods—problems easy for RFdiffusion may be hard for EvoDiff and vice versa, suggesting genuinely orthogonal capabilities.
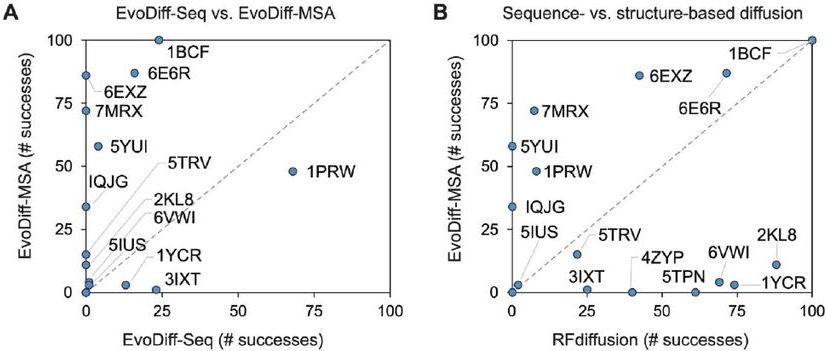
Figure 10: Complementary scaffolding capabilities of sequence- and structure-based methods. (A) EvoDiff-MSA (y-axis) generally outperforms EvoDiff-Seq (x-axis) on scaffolding benchmarks, benefiting from evolutionary information in MSAs. (B) EvoDiff-MSA versus RFdiffusion shows near-zero correlation—problems easy for one method are not necessarily easy for the other, suggesting orthogonal design capabilities. Success defined as pLDDT ≥ 70 and motif RMSD ≤ 1.0 Å over n=100 trials per problem. Source: Alamdari et al., bioRxiv 2024, Figure 6A-B.
Experimental validation confirms functional protein generation
The strongest evidence comes from wet-lab experiments. Of 13 nominated scaffold designs (selected for pLDDT > 70 and motifRMSD < 1Å), 12 expressed successfully—a 92% expression rate demonstrating EvoDiff generates synthesizable proteins.
Calcium binding validation targeted the 1PRW calmodulin binding motif. The design EvoDiff-Seq-1PRW-Design-1 exhibited spectroscopic shift in the presence of Ca²⁺, indicative of successful calcium coordination. CD spectroscopy confirmed generated proteins display expected secondary structure elements, validating OmegaFold predictions.
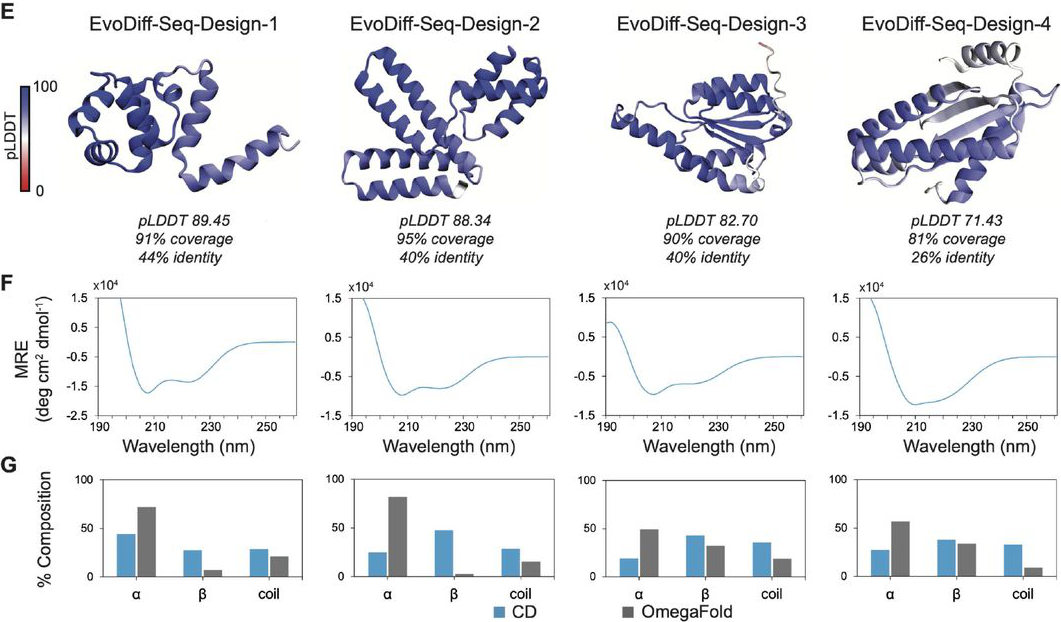
Figure 11: Experimental validation of unconditional EvoDiff designs. (E) Four designs expressed successfully in E. coli or CHO cells, showing diverse predicted structures with varying pLDDT scores and low sequence identity to natural proteins. (F) CD spectra confirm the designs adopt secondary structures with characteristic α-helix and β-sheet signals. (G) Secondary structure composition from CD (blue) generally agrees with OmegaFold predictions (gray), validating computational structure predictions. Source: Alamdari et al., bioRxiv 2024, Figure 2E-G.
MDM2 binding validation:
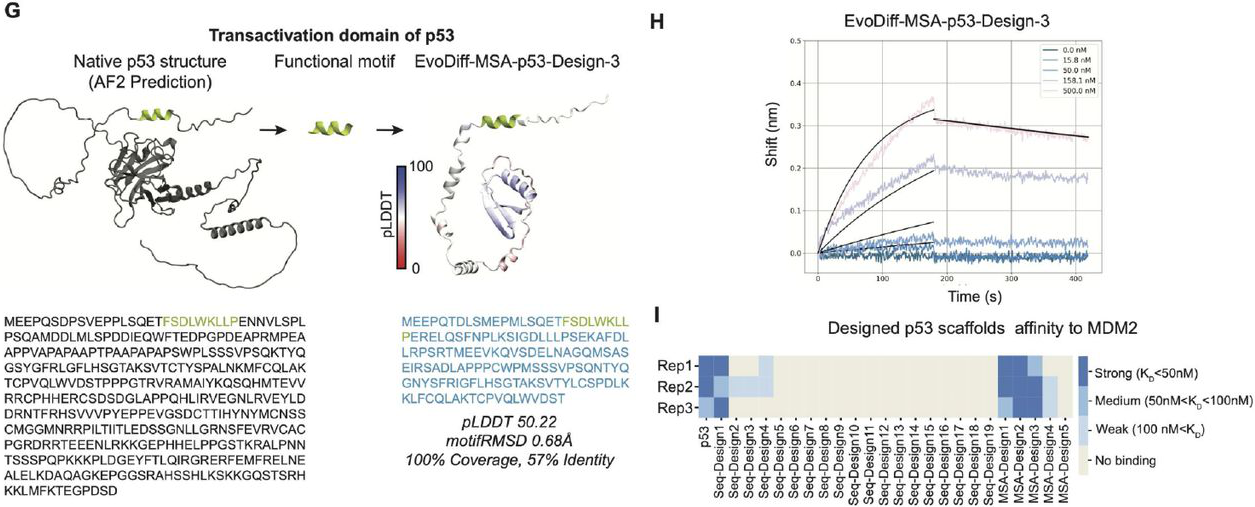
Figure 12: EvoDiff designs functional p53-MDM2 binders without explicit structure. (G) The native p53 transactivation domain contains intrinsic disorder (N-terminal). EvoDiff scaffolds only the binding motif sequence (green) without binder structure information. (H) BLI measurements show EvoDiff-MSA-p53-Design-3 binds MDM2 with KD ~40 nM. (I) Summary of 24 tested designs reveals 4 strong binders (KD < 50 nM) and 8 total designs showing measurable binding, demonstrating EvoDiff's capability for sequence-only functional design. Source: Alamdari et al., bioRxiv 2024, Figure 6G-I.
These results, while preliminary compared to RFdiffusion’s extensive binder validation (\(19%\) experimental success rate, 28 nM HA binders, sub-nanomolar MDM2 binders), demonstrate EvoDiff produces functional proteins rather than merely plausible sequences.
RFdiffusion excels at precise geometric control
RFdiffusion, the leading structure-based diffusion model, operates through $\text{SE(3)}$ diffusion on residue rigid frames. Each residue is represented as a rotation matrix $r ∈ \text{SO(3)}$ plus translation $x ∈ ℝ^3$. The forward process applies Gaussian noise to $\text{Cα}$ coordinates and Brownian motion on $\text{SO(3)}$ for orientations.
Built on fine-tuned RoseTTAFold (~168M parameters), RFdiffusion leverages pretrained structure prediction knowledge through self-conditioning between timesteps. This enables remarkable capabilities: unconditional generation up to 600 residues, motif scaffolding solving 23/25 benchmark problems, and binder design with 19% experimental success rate—two orders of magnitude improvement over previous methods.
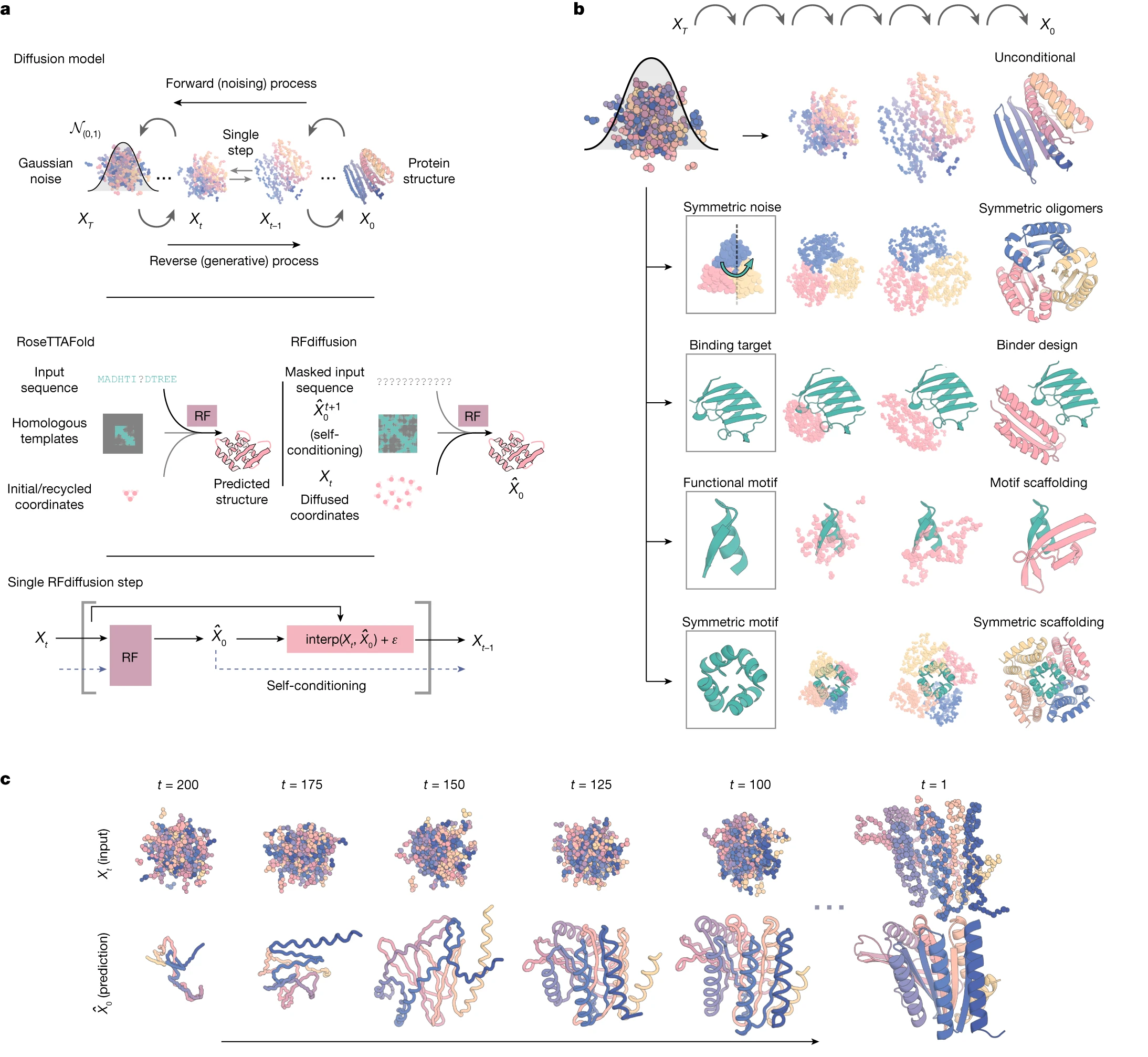
Figure 13: RFdiffusion uses SE(3) diffusion on 3D protein structures. The forward process corrupts residue positions and orientations with Gaussian noise and SO(3) Brownian motion. The reverse process, learned by a fine-tuned RoseTTAFold network, generates protein backbones that can be conditioned on structural motifs, symmetry constraints, or binding targets. This enables precise geometric control but cannot design proteins lacking defined structure. Source: Watson et al., Nature 2023, Figure 1.
However, RFdiffusion’s structure-centric approach creates fundamental limitations:
- Cannot design IDRs: No target structure exists to diffuse toward
- Helical bias: Generated proteins enriched in α-helices versus natural distributions
- Limited training data: ~200K PDB structures versus 42M evolutionary sequences
- Requires ProteinMPNN: Outputs backbones only; sequence design is a separate step
Chroma from Generate Biomedicines offers programmable conditioning through a composable framework enabling symmetry constraints, shape conditioning, and even natural language prompts. Its random graph neural network achieves sub-quadratic scaling, handling proteins exceeding 4,000 residues. However, experimental validation primarily demonstrates structural generation rather than functional design.
Genie 2 uniquely trains on the AlphaFold Database (~588K structures), achieving state-of-the-art unconditional metrics and introducing multi-motif scaffolding—designing proteins with multiple functional sites whose geometric relationships aren’t prespecified.
Protein language models offer complementary capabilities
ESM-2 (8M-15B parameters) uses masked language modeling on ~65M UniRef sequences, producing excellent embeddings for downstream tasks but functioning as a discriminative encoder rather than generative model. Generating sequences requires Gibbs sampling or similar Monte Carlo methods, and like RFdiffusion, produces helix-enriched outputs.
ProGen/ProGen2 (up to 6.4B parameters) employs autoregressive generation with control tags for conditional generation. Generated lysozymes achieved catalytic efficiency comparable to natural enzymes at only 31% sequence identity. However, left-to-right generation limits flexibility for conditional tasks.
ProtGPT2 (738M parameters) adapts GPT-2 for protein sequences, generating 88% globular proteins matching natural amino acid propensities. Like ProGen, its autoregressive nature prevents natural infilling.
CARP (640M parameters) uses ByteNet CNNs like EvoDiff-Seq but with masked language modeling rather than diffusion. It achieves competitive performance with linear complexity, demonstrating CNN architectures effectively capture protein patterns.
EvoDiff’s order-agnostic diffusion distinguishes it from all these models. Unlike autoregressive methods (ProGen, ProtGPT2, RITA), it generates in any order. Unlike masked language models (ESM-2, CARP), it provides a principled generative framework with iterative refinement. Unlike structure diffusion, it operates purely in sequence space, accessing the full evolutionary design landscape.
ProteinMPNN dominates inverse folding but requires structures
ProteinMPNN represents the state-of-the-art for inverse folding—predicting sequences from backbone structures. Its message passing neural network achieves 52.4% native sequence recovery versus 32.9% for Rosetta, while running 200× faster (1.2s versus 258.8s per protein).
The key limitation: ProteinMPNN requires an input structure. It cannot generate de novo; it can only design sequences for existing or predicted backbones. This makes it complementary to both structure-based generators (providing sequences for RFdiffusion backbones) and EvoDiff (validating generated sequences through self-consistency).
FoldingDiff takes another approach: diffusion on internal backbone angles ($φ, ψ, ω$, and bond angles) rather than Cartesian coordinates. This representation provides inherent rotational invariance without equivariant networks. However, it generates backbones only, requiring downstream inverse folding for sequences.
Choosing the right tool depends on the design problem
The protein design landscape now offers genuinely complementary approaches:
| Design Goal | Recommended Tool | Rationale |
|---|---|---|
| Precise binding interface | RFdiffusion + ProteinMPNN | Direct geometric control, validated binder success |
| Disordered functional regions | EvoDiff | Only option for IDRs |
| Diverse sequence exploration | EvoDiff or ProGen | Better functional space coverage |
| Symmetric assemblies | RFdiffusion or Chroma | Native symmetry support |
| Multi-motif scaffolds | Genie 2 or EvoDiff-MSA | Handle complex conditioning |
| Sequence optimization | ProteinMPNN or ESM-IF | Inverse folding from known structure |
An integrated pipeline might use EvoDiff to generate diverse sequences with desired properties, AlphaFold2/ESMFold to validate foldability, ProteinMPNN to optimize sequences for predicted structures, and structure-based methods to refine scaffolds when precise geometry matters.
References
Alamdari S, Thakkar N, van den Berg R, Tenenholtz N, Strome R, Moses AM, Lu AX, Fusi N, Amini AP, Yang KK. Protein generation with evolutionary diffusion: sequence is all you need. Nature Biotechnology (2023). https://doi.org/10.1038/s41587-023-01902-3
Austin J, Johnson DD, Ho J, Tarlow D, van den Berg R. Structured denoising diffusion models in discrete state-spaces. Advances in Neural Information Processing Systems 34 (2021).
Dauparas J, Anishchenko I, Bennett N, Bai H, Ragotte RJ, Milles LF, Wicky BIM, Courber A, de Haas RJ, Bethel N, Leung PJY, Huddy TF, Pellock S, Tischer D, Chan F, Koepnick B, Nguyen H, Kang A, Sankaran B, Bera AK, King NP, Baker D. Robust deep learning–based protein sequence design using ProteinMPNN. Science 378, 49-56 (2022).
Ferruz N, Schmidt S, Höcker B. ProtGPT2 is a deep unsupervised language model for protein design. Nature Communications 13, 4348 (2022).
Hesslow D, Zanichelli N, Notin P, Poli I, Marks D. RITA: a study on scaling up generative protein sequence models. arXiv preprint arXiv:2205.05789 (2022).
Ho J, Jain A, Abbeel P. Denoising diffusion probabilistic models. Advances in Neural Information Processing Systems 33, 6840-6851 (2020).
Hoogeboom E, Nielsen D, Jaini P, Forré P, Welling M. Argmax flows and multinomial diffusion: Learning categorical distributions. Advances in Neural Information Processing Systems 34 (2021).
Hsu C, Verkuil R, Liu J, Lin Z, Hie B, Sercu T, Lerer A, Rives A. Learning inverse folding from millions of predicted structures. ICML (2022).
Ingraham JB, Barber M, Karber M, Tsuboyama K, Doaee A, Metz S, et al. Illuminating protein space with a programmable generative model. Nature 623, 1070-1078 (2023).
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583-589 (2021).
Lin Z, Akin H, Rao R, Hie B, Zhu Z, Lu W, Smetanin N, Verkuil R, Kabeli O, Shmueli Y, dos Santos Costa A, Fazel-Zarandi M, Sercu T, Candido S, Rives A. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123-1130 (2023).
Lin Y, AlQuraishi M. Generating novel, designable, and diverse protein structures by equivariantly diffusing oriented residue clouds. ICML (2023).
Lin Y, AlQuraishi M. Genie 2: Scaling large-scale de novo protein design. arXiv preprint arXiv:2405.15489 (2024).
Lisanza SL, Gershon JM, Tipps SWK, Arnez L, Merle M, Vogt-Maranto L, Chen JD, Pan X, Correia BE, Baker D. Joint generation of protein sequence and structure with RoseTTAFold sequence space diffusion. Nature Biotechnology (2024).
Madani A, Krause B, Greene ER, Subramanian S, Mohr BP, Holton JM, Olmos JL Jr, Xiong C, Sun ZZ, Socher R, Fraser JS, Naik N. Large language models generate functional protein sequences across diverse families. Nature Biotechnology 41, 1099-1106 (2023).
Nijkamp E, Ruffolo JA, Weinstein EN, Naber N, Madani A. ProGen2: Exploring the boundaries of protein language models. Cell Systems 14, 968-978 (2023).
Rao R, Liu J, Verkuil R, Meier J, Canny JF, Abbeel P, Sercu T, Rives A. MSA Transformer. ICML (2021).
Rives A, Meier J, Sercu T, Goyal S, Lin Z, Liu J, Guo D, Ott M, Zitnick CL, Ma J, Fergus R. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences 118, e2016239118 (2021).
Watson JL, Juergens D, Bennett NR, Trippe BL, Yim J, Eisenach HE, Ahern W, Borber AJ, Ragotte RJ, Milles LF, et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089-1100 (2023).
Wu KE, Yang KK, van den Berg R, Alamdari S, Zou JY, Lu AX, Amini AP. Protein structure generation via folding diffusion. Nature Communications 15, 1059 (2024).
Yang KK, Fusi N, Lu AX. Convolutions are competitive with transformers for protein sequence pretraining. Cell Systems 15, 286-294 (2024).
Enjoy Reading This Article?
Here are some more articles you might like to read next: