IgFold - a fast, accurate antibody structure prediction
How deep learning on 558 million antibodies revolutionized structure prediction?
IgFold achieves AlphaFold-level accuracy for antibody structure prediction in under 25 seconds, without requiring multiple sequence alignments. By pre-training a language model on 558 million natural antibody sequences and combining graph transformers with invariant point attention, researchers at Johns Hopkins University created a method that democratizes high-throughput antibody structure prediction. The tool enabled prediction of 1.4 million antibody structures—a 500-fold expansion over experimentally determined structures—opening new possibilities for therapeutic antibody development and large-scale repertoire analysis.
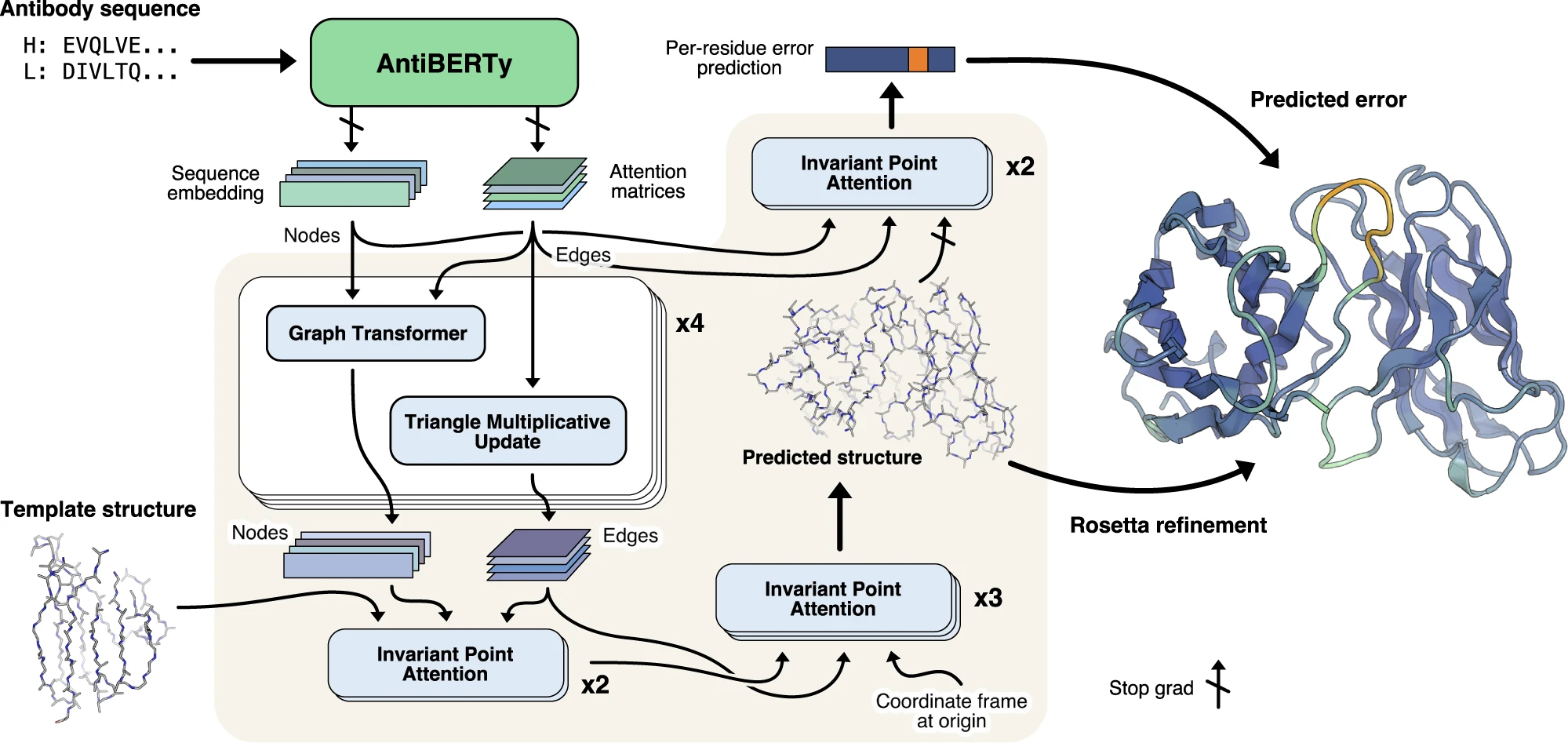
Figure 1: IgFold pipeline for end-to-end antibody structure prediction. IgFold combines pre-trained language model embeddings with graph neural networks and geometric attention for direct coordinate prediction. Antibody sequences (heavy and light chains) are encoded by AntiBERTy, a BERT-based transformer pre-trained on 558 million natural antibody sequences. Node features are extracted from final hidden layer embeddings, while edge features are derived from attention matrices across all transformer layers. Four graph transformer layers refine these representations through edge-biased attention and triangle multiplicative updates. Template structures (optional) are incorporated via invariant point attention (IPA) with fixed coordinates, enabling structure-aware self-attention. Three additional IPA layers predict final 3D backbone atom coordinates by iteratively translating and rotating each residue from an initialized "residue gas" configuration. A separate error estimation module predicts per-residue accuracy using two IPA layers operating on the final structure. Rosetta refinement adds side chains and corrects geometric abnormalities to produce the final full-atom structure. Source: Ruffolo et al. 2023, Figure 1.
Antibodies remain among the most important therapeutic molecules, yet structure prediction has historically been a bottleneck. While AlphaFold revolutionized general protein structure prediction, antibodies present unique challenges: their complementarity-determining regions (CDRs), particularly CDR H3, exhibit extreme sequence diversity and conformational flexibility that general-purpose methods struggle to capture. IgFold addresses this gap by leveraging the massive corpus of sequenced natural antibodies to learn antibody-specific structural patterns, achieving state-of-the-art accuracy with unprecedented speed.
AntiBERTy: learning antibody structure from 558 million sequences
The foundation of IgFold lies in AntiBERTy, a BERT-based language model pre-trained on 558 million natural antibody sequences from the Observed Antibody Space (OAS) database. This pre-training approach fundamentally differs from AlphaFold’s reliance on multiple sequence alignments (MSAs)—a critical distinction that enables IgFold’s dramatic speed advantage.
AntiBERTy uses a compact architecture with 26 million parameters distributed across 8 transformer layers, each containing 8 attention heads. The model produces 512-dimensional embeddings per residue, trained using standard masked language modeling where 15% of amino acids are randomly masked and predicted from surrounding context. This self-supervised objective forces the model to learn co-evolutionary relationships between residues, conservation patterns within framework regions, and the unique grammar of hypervariable CDR loops.
Remarkably, the embeddings encode structural information without any explicit structure supervision during pre-training. When visualized using t-SNE dimensionality reduction, CDR loop embeddings naturally cluster by canonical structural classes—CDR H1 and H2 loops with similar folds group together, while the structurally diverse CDR H3 loops organize roughly by length since they lack canonical forms. This demonstrates that sequence context alone captures nascent structural features, validating the transfer learning hypothesis underlying modern protein language models.
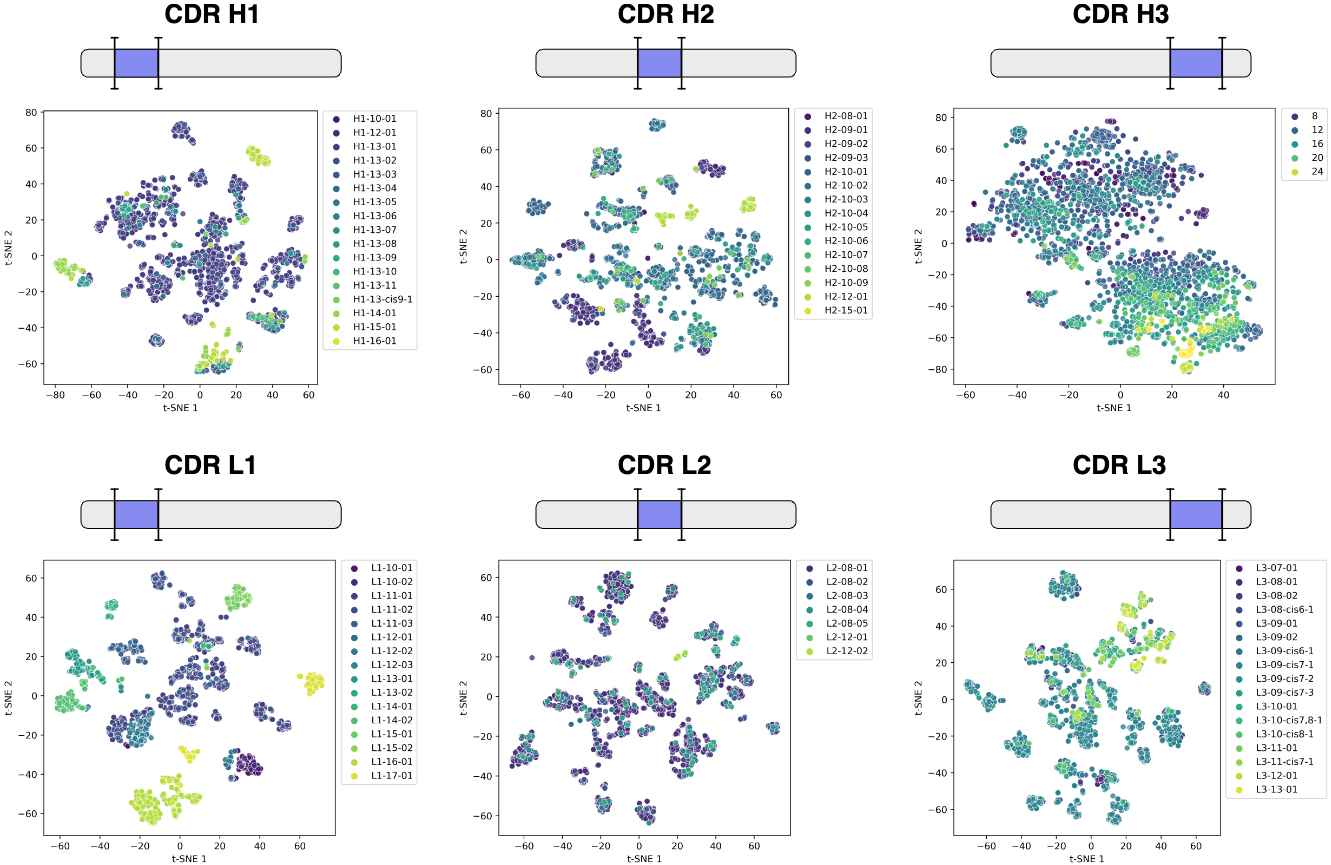
Figure 2: AntiBERTy sequence embeddings encode structural features. Two-dimensional t-SNE visualization of CDR loop embeddings from AntiBERTy for 3,467 paired antibody sequences with experimentally determined structures. For each sequence, embeddings corresponding to each CDR loop are extracted and averaged to create fixed-size representations. Points are colored by canonical structural cluster (CDR H1, H2, L1, L2, L3) or loop length (CDR H3, which lacks canonical classification). CDR H1 shows the clearest structural clustering, with different canonical classes (H1-10-01, H1-12-01, etc.) forming distinct regions in embedding space. CDR L1-L3 also show organization by structural class, though with more overlap. CDR H3 organizes primarily by loop length (colors from yellow=short to purple=long) rather than discrete structural clusters, reflecting the extreme diversity of H3 conformations. This demonstrates that self-supervised pre-training on sequence alone, without structural supervision, learns latent structural features—providing a strong foundation for downstream structure prediction. Source: Ruffolo et al. 2023, Figure S1.
IgFold leverages AntiBERTy’s representations in two ways:
- Node features: Final hidden layer embeddings (L × 512) are projected to per-residue node representations
- Edge features: Attention matrices from all 64 attention heads (8 layers × 8 heads) are stacked into an L × L × 64 tensor, then projected to pairwise edge representations
This dual extraction captures both local residue properties and learned residue-residue interaction patterns—attention weights often correlate with structural contacts, effectively encoding a learned “contact prior” that guides subsequent structure prediction.
Graph transformers translate embeddings into structure
IgFold’s architecture converts language model representations into 3D coordinates through a carefully designed pipeline of graph transformers and geometric attention modules. The complete model contains only 1.6 million trainable parameters—remarkably compact compared to AlphaFold’s ~93 million—yet achieves comparable accuracy on antibody structures.
Graph transformer layers for representation refinement
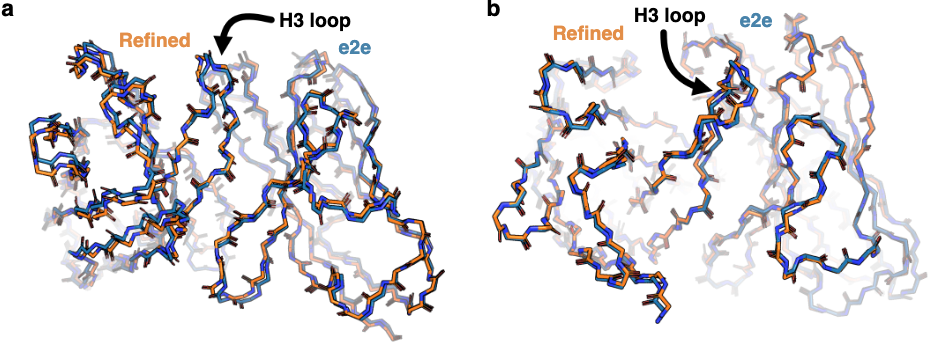
Figure 3: Minimal impact of refinement on prediction accuracy. (a) Comparison of structure prediction RMSD before (IgFold[e2e], orange) and after (IgFold, blue) Rosetta refinement on 197 paired antibody benchmark targets. Median RMSD changes are minimal ($\lt 0.5 Å$) for all structural elements, indicating that direct end-to-end predictions from IgFold's neural network are already geometrically reasonable. The exception is CDR H3, where some long loops show larger refinement effects due to pre-refinement bond length/angle abnormalities. (b) Scatter plots comparing pre- and post-refinement RMSD for each structural element. Points clustered near the diagonal (within gray bands) indicate minimal change. The few outliers represent long CDR loops or unusual structures requiring more extensive energy minimization. Source: Ruffolo et al. 2023, Figure S4a-b.
Four graph transformer layers iteratively refine node and edge representations before coordinate prediction. The attention mechanism incorporates edge information directly:
\[\alpha_{ij} = \frac{\exp(\langle q_i, k_j + e_{ij} \rangle / \sqrt{d})}{\sum_k \exp(\langle q_i, k_k + e_{ik} \rangle / \sqrt{d})}\]This edge-biased attention allows the network to learn which residue pairs should attend strongly to each other, informed by the AntiBERTy-derived edge features. Node updates use gated residual connections where a learned sigmoid gate determines how much new information to incorporate: $h_\text{new} = (1-\beta)h + \beta\hat{h}$.
Edge updates employ triangle multiplicative operations borrowed from AlphaFold2’s Evoformer. These operations enforce geometric consistency by aggregating information from triangles of residues—if residues $i$ and $k$ are close, and $k$ and $j$ are close, then $i$ and $j$ likely interact. Both “outgoing” (aggregating from $i→k$ and $j→k$ edges) and “incoming” (aggregating from $k→i$ and $k→j$ edges) triangle updates are applied, ensuring the edge representation reflects geometric constraints.
Invariant point attention predicts coordinates
The core innovation enabling direct coordinate prediction is Invariant Point Attention (IPA), originally developed for AlphaFold2’s structure module. IPA operates on a “residue gas” representation where each residue is modeled as an independent rigid body defined by its backbone triangle ($\mathrm{N, Cα, C}$ atoms). Each residue maintains a coordinate frame—a 4×4 transformation matrix encoding rotation and translation—that transforms from a canonical reference to the predicted position.
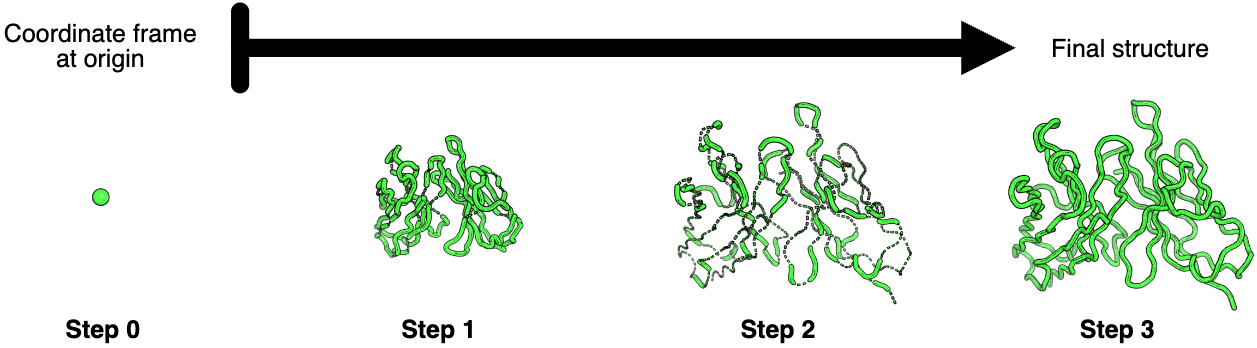
Figure 4: Stepwise coordinate prediction through invariant point attention layers. Progressive refinement of antibody structure through three IPA layers. Starting from initialization with all residue coordinate frames at the origin (Step 0, single green point), the first IPA layer (Step 1) predicts a compact arrangement resembling the final structure but at reduced scale. The second IPA layer (Step 2) expands the structure to approximately correct scale but with numerous chain breaks and abnormal bond geometries (visible as disconnected backbone segments). The third and final IPA layer (Step 3) resolves most geometric abnormalities, producing proper backbone connectivity and realistic bond lengths/angles. This sequential refinement strategy—global arrangement → correct scaling → local geometry—mirrors coarse-to-fine optimization approaches in computational structural biology. Minor remaining geometric imperfections are corrected during subsequent Rosetta refinement. Source: Ruffolo et al. 2023, Figure S2.
IPA achieves $\text{SE(3)}$ invariance (invariance to global rotations and translations) through a clever mathematical construction. In addition to standard query-key-value attention, IPA produces 3D point queries and keys in each residue’s local coordinate frame. These points are projected into the global frame for attention computation, but the attention weights depend only on squared distances between transformed points:
\[\text{attention contribution} \propto \lVert T_i \cdot q_{point} - T_j \cdot k_{point} \rVert^2\]Since the L2-norm is invariant to rotation ($\lVert R \cdot v \rVert = \lVert v \rVert$) and translation ($\lVert v+t - w-t \rVert = \lVert v-w \rVert$), the resulting attention is $\text{SE(3)}$-invariant—the model learns the same representations regardless of how the structure is oriented in space.
IgFold employs IPA in two contexts:
- Template incorporation: 2 IPA layers with fixed template coordinates, acting as structure-aware self-attention
- Structure prediction: 3 IPA layers that iteratively translate and rotate each residue from the origin to its final position
Unlike AlphaFold, IgFold uses separate weights for each IPA layer and allows gradient propagation through rotations, enabling end-to-end training of the coordinate prediction pipeline.
Template incorporation and error estimation
IgFold’s template strategy differs fundamentally from traditional homology modeling. Rather than grafting loop conformations from template structures, IgFold uses templates as attention context through IPA. During training, templates are generated by corrupting true structures: for 50% of training examples, 1-6 consecutive segments of 20 residues are selected and their coordinates moved to the origin (effectively masking them), while remaining residues serve as template context with masked attention preventing information leakage.
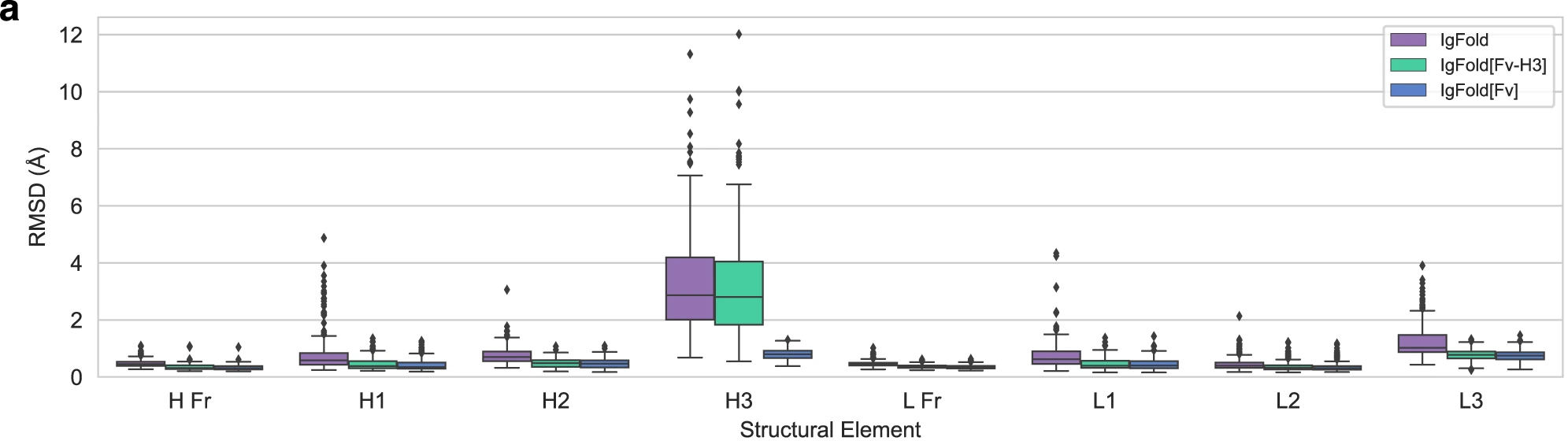
Figure 5: Template incorporation improves CDR H3 prediction for paired antibodies. Effect of template information on structure prediction accuracy. IgFold predictions without templates (orange) compared to predictions with partial templates (IgFold[Fv-H3]: green, providing framework and non-H3 CDR structures) and full templates (IgFold[Fv]: purple, providing complete structure). Benchmark on 197 paired antibodies shows that providing framework templates marginally improves CDR H3 prediction (median RMSD decreases from ~3.3 Å to ~3.0 Å) by establishing correct inter-chain orientation and local structural context. Full template incorporation nearly perfectly recreates the template structure, validating that IPA successfully incorporates 3D coordinate information. Template usage is particularly beneficial for antibodies with unusual frameworks or non-canonical inter-chain orientations. Source: Ruffolo et al. 2023, Figure 4a.
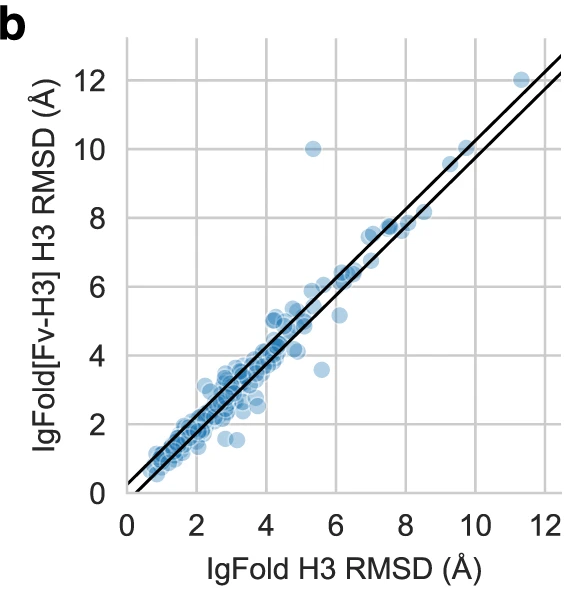
Figure 6: Per-target effect of templates on CDR H3 accuracy. Scatter plot comparing CDR H3 RMSD without templates (IgFold, x-axis) versus with framework templates (IgFold[Fv-H3], y-axis). Most points cluster near the diagonal, indicating templates provide modest improvement for typical cases. Points below diagonal show cases where template context improves H3 prediction; points above diagonal (fewer) indicate template information occasionally constrains the model unfavorably. The overall shift below diagonal demonstrates that structural context generally aids CDR H3 prediction without strongly biasing predictions when templates are suboptimal. Source: Ruffolo et al. 2023, Figure 4b.
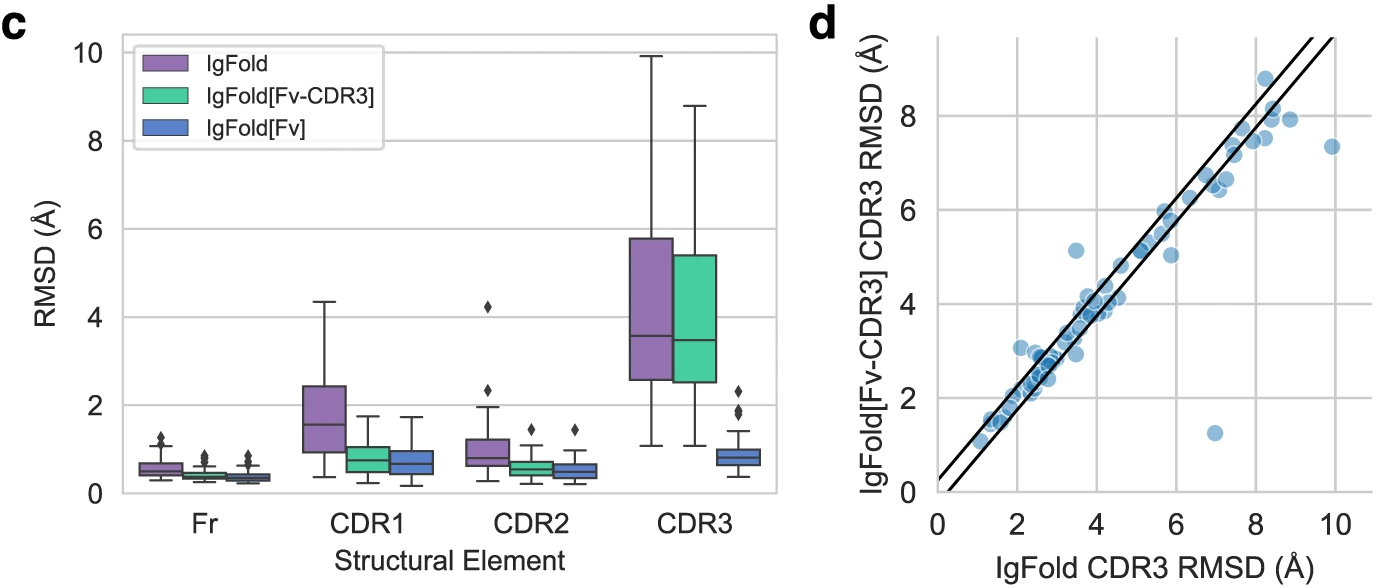
Figure 7: Template effects on nanobody structures. (c) Nanobody benchmark (n=71) shows stronger template effects than paired antibodies. Providing CDR1/CDR2 context (IgFold[Fv-CDR3]) notably improves CDR3 prediction for many targets. (d) Scatter plot reveals several cases (points well below diagonal) where template context substantially improves CDR3 accuracy, particularly for long CDR3 loops that adopt structured conformations. The absence of a light chain in nanobodies makes CDR3 conformation more dependent on framework-CDR interactions, explaining why templates provide greater benefit than for paired antibodies. Source: Ruffolo et al. 2023, Figure 4c-d.
This approach enables flexible template usage at inference time. Users can provide partial experimental structures (e.g., framework regions from a similar antibody) and let IgFold predict unknown regions (e.g., CDR loops) in context. Benchmarks show that providing framework templates (IgFold[Fv-H3]) or full Fv templates (IgFold[Fv]) improves CDR H3 prediction, particularly for challenging cases.
Per-residue error estimation
IgFold provides pLDDT-like confidence scores through a dedicated error estimation module. Two additional IPA layers operate on the final predicted structure using separate node and edge features from AntiBERTy, predicting per-residue $\text{Cα}$ deviation from the true structure. The training loss is the L1 norm between predicted and actual deviation, with structures aligned to the relatively rigid beta barrel framework residues.
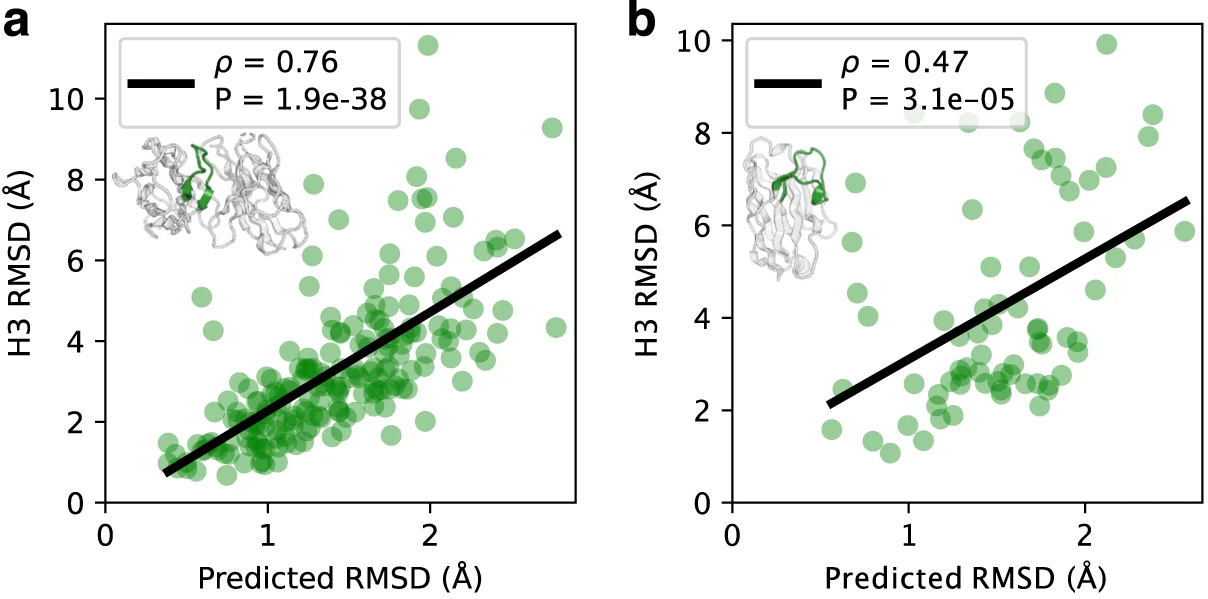
Figure 8: Correlation between predicted and actual CDR loop errors. Validation of IgFold's per-residue error estimation capability. (a) CDR H3 loops from paired antibody benchmark (n=197): predicted RMSD (x-axis) versus actual RMSD from native structure (y-axis) shows strong correlation (Spearman ρ = 0.76, p = 1.9×10⁻³⁸). (b) CDR3 loops from nanobody benchmark (n=71): moderate correlation (Spearman ρ = 0.47, p = 3.1×10⁻⁵). Points colored by predicted error (blue = low, red = high). Gray shaded region represents cumulative average RMSD of predictions. The error estimation model tends to underpredict actual RMSD, particularly for high-error cases, likely due to class imbalance between many low-RMSD framework residues and fewer high-RMSD CDR residues during training. Despite this miscalibration, the model effectively ranks prediction quality, enabling users to identify unreliable structures for experimental validation. Source: Ruffolo et al. 2023, Figure 3a-b.
Critically, gradients are stopped through the error estimation layers—this prevents the model from optimizing for estimated but erroneous structures. The predicted errors correlate well with actual RMSD: Spearman ρ = 0.76 for CDR H3 loops in paired antibodies and ρ = 0.47 for nanobody CDR3. This enables confidence-aware downstream applications; users can trust predictions with low estimated error while flagging uncertain structures for experimental validation or alternative modeling approaches.
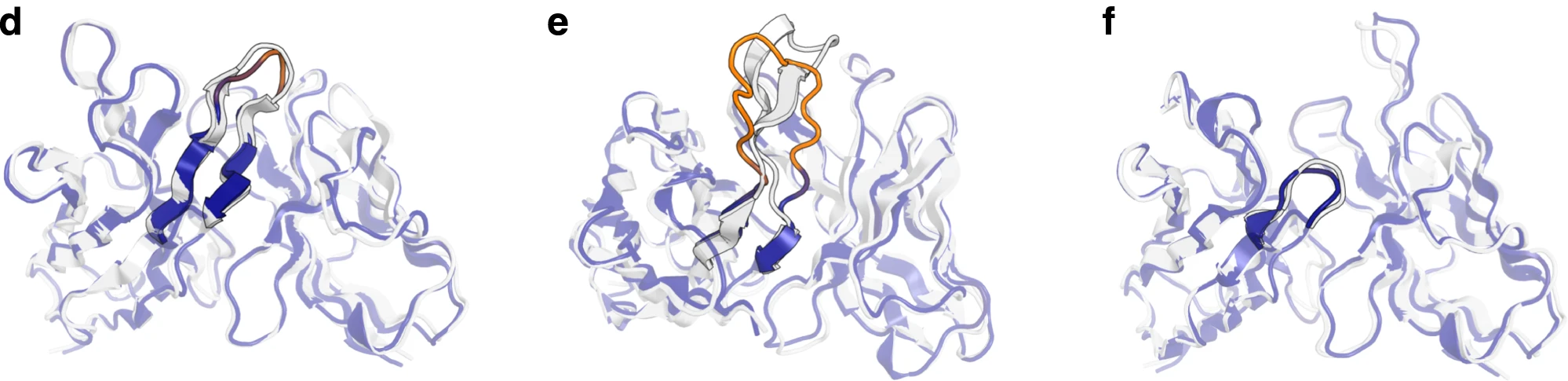
Figure 9: Examples of error-colored structural predictions. Representative structures showing per-residue error estimation. Structures colored by predicted $\text{Cα}$ deviation from native: blue (low error) to red (high error). (d) Benchmark target 7O4Y with accurately predicted 12-residue CDR H3 (RMSD = 1.64 Å)—appropriately assigned low predicted error (blue). (e) Benchmark target 7RKS with challenging 18-residue CDR H3 containing structured beta sheet elements. IgFold struggles with this long loop (RMSD = 6.33 Å, shown in orange/red) and correctly predicts high uncertainty. (f) Target 7O33 with very short 3-residue CDR H3, predicted accurately (RMSD = 1.49 Å) with correspondingly low error estimate (blue). Source: Ruffolo et al. 2023, Figure 3d-f.
Training data augmentation with AlphaFold
IgFold’s training set combines experimental structures with AlphaFold-generated predictions, a pragmatic approach to overcome the limited availability of experimental antibody structures.
| Dataset | Structures | Source |
|---|---|---|
| SAbDab (experimental) | 4,275 | Paired antibodies and nanobodies |
| AlphaFold-augmented (paired) | 16,141 | OAS sequences clustered at 40% identity |
| AlphaFold-augmented (unpaired) | 22,132 | Filtered from 26,971 (pLDDT ≥ 85) |
| Total | ~42,500 | Mixed experimental + predicted |
Structures are sampled with inverse weighting to balance approximately one-third from each source during training. The loss function is mean squared error between predicted and experimental/reference coordinates after Kabsch alignment for optimal superposition.
This augmentation strategy is notably conservative—only high-confidence AlphaFold predictions (pLDDT ≥ 85) are included. The approach acknowledges that while AlphaFold’s antibody predictions are imperfect, they provide valuable additional training signal that improves generalization, particularly for sequence regions underrepresented in experimental databases.
Benchmark performance establishes new state-of-the-art
IgFold was evaluated on temporally separated test sets to ensure fair comparison: 197 paired antibodies and 71 nanobodies deposited after training set compilation.
Paired antibody performance
| Method | CDR H1 (Å) | CDR H2 (Å) | CDR H3 (Å) | Framework (Å) |
|---|---|---|---|---|
| IgFold | 0.77 | 0.73 | 2.73 | 0.47 |
| AlphaFold-Multimer | 0.77 | 0.74 | 2.86 | 0.80 |
| DeepAb | 0.79 | 0.75 | 2.79 | 0.43 |
| ABlooper | 0.80 | 0.73 | 2.94 | 0.53 |
| RepertoireBuilder | 1.06 | 0.90 | 3.53 | 0.49 |
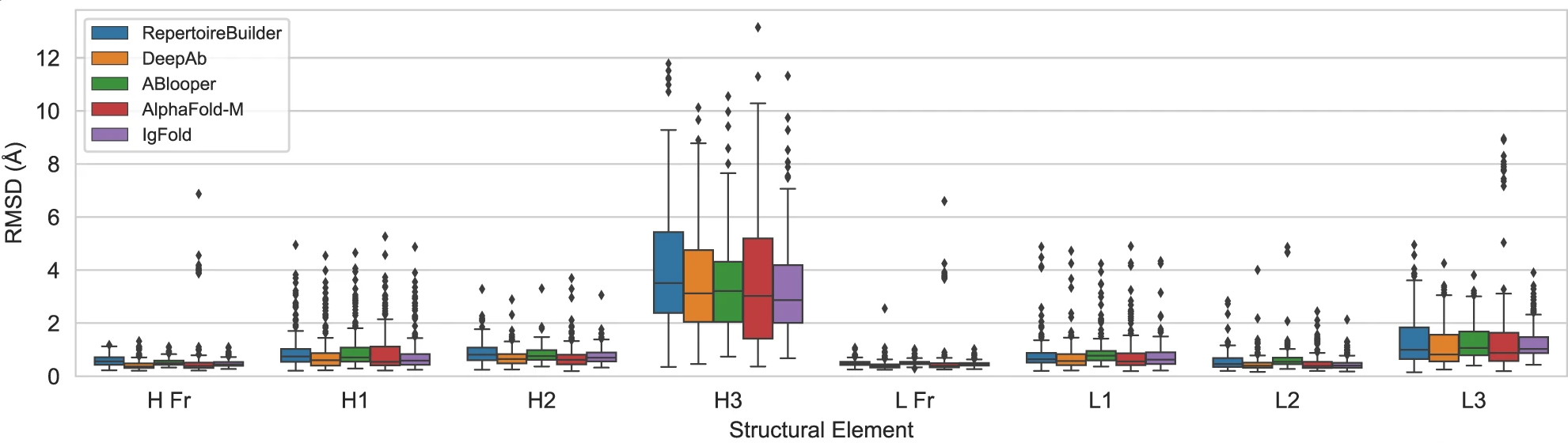
Figure 10: Benchmark performance on paired antibody structure prediction. Comparison of five methods on 197 paired antibody structures deposited after July 2021. Box plots show backbone heavy-atom RMSD ($\text{N, Cα, C, O}$) for framework regions (H Fr, L Fr) and six CDR loops (H1-H3, L1-L3) after alignment of framework residues. All deep learning methods (DeepAb, ABlooper, AlphaFold-Multimer, IgFold) achieve sub-angstrom accuracy on canonical CDR loops and framework regions. The critical differentiator is CDR H3, the most structurally variable region: IgFold achieves the best average performance (3.27 Å RMSD) followed closely by ABlooper (3.54 Å) and AlphaFold-Multimer (3.56 Å). Template-based RepertoireBuilder shows notably worse CDR H3 prediction (4.15 Å RMSD). Box plots show median (center line), interquartile range (box bounds), 1.5×IQR whiskers, and outliers beyond 1.5×IQR. Source: Ruffolo et al. 2023, Figure 2a.
All deep learning methods achieve sub-angstrom accuracy on canonical CDR loops (H1, H2, L1, L2, L3) and framework regions. The critical differentiator is CDR H3—the most variable and structurally challenging region. IgFold achieves the lowest mean RMSD (2.73 Å), though the difference from AlphaFold-Multimer (2.86 Å) is modest. More importantly, the methods appear complementary: approximately 20% of CDR H3 loops differ by >4 Å RMSD between IgFold and AlphaFold-Multimer predictions, suggesting they capture distinct conformational possibilities.
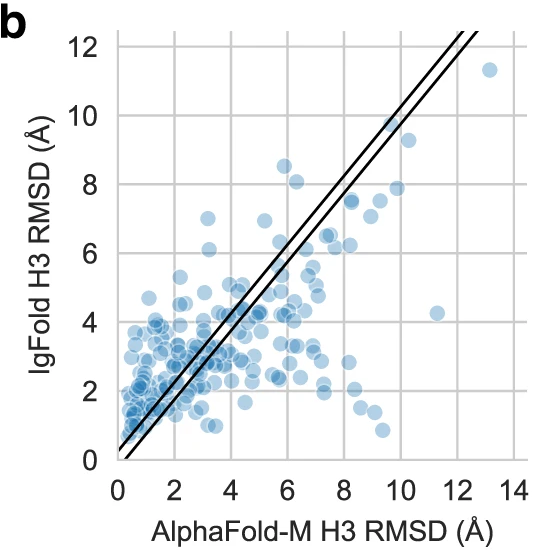
Figure 11: Per-target CDR H3 comparison between IgFold and AlphaFold-Multimer. Scatter plot comparing CDR H3 loop RMSD for IgFold (y-axis) versus AlphaFold-Multimer (x-axis) across 197 paired antibody benchmark targets. Each point represents one antibody structure. Diagonal bands indicate predictions within ±1 Å agreement. Approximately 20% of targets show >4 Å RMSD difference between methods (points far from diagonal), indicating the methods predict distinct CDR H3 conformations for the same sequence. This complementarity suggests ensemble approaches combining both predictions could improve conformational coverage. Points above diagonal represent cases where AlphaFold-Multimer outperforms IgFold; points below diagonal show IgFold superiority. Source: Ruffolo et al. 2023, Figure 2b.
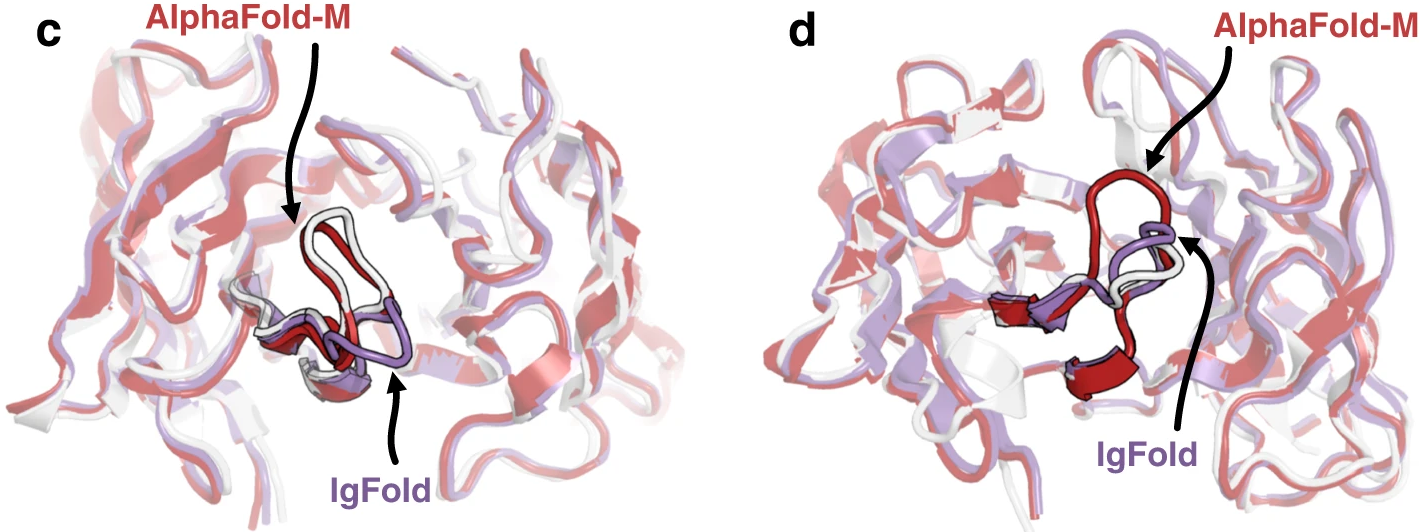
Figure 12: Contrasting CDR H3 predictions for representative targets. Structural comparisons highlighting cases where IgFold and AlphaFold-Multimer predict different CDR H3 conformations. (c) Target 7N3G with 10-residue CDR H3: AlphaFold-Multimer accurately predicts the native structure (RMSD = 0.98 Å, pink) while IgFold predicts an alternative conformation (RMSD = 4.69 Å, dark blue). (d) Target 7RNJ with 9-residue CDR H3: IgFold accurately captures the native "stretched twist" conformation (RMSD = 1.18 Å, dark blue) while AlphaFold-Multimer predicts an extended loop (RMSD = 3.46 Å, pink). Native structure shown in light gray. These examples illustrate that neither method universally outperforms the other—each captures structural features the other misses. Source: Ruffolo et al. 2023, Figure 2c-d.
Nanobody performance reveals different challenges
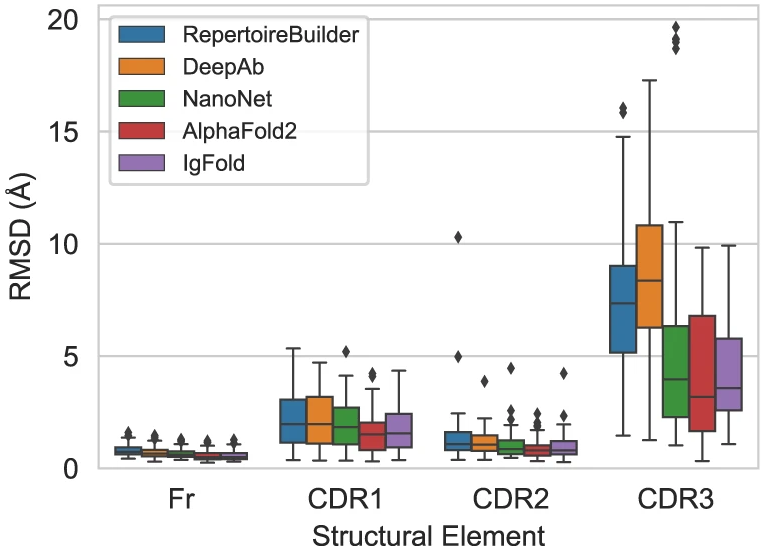
Figure 13: Benchmark performance on nanobody structure prediction. Comparison of five methods on 71 nanobody structures. Single-domain antibodies (nanobodies) present additional challenges compared to paired antibodies due to longer CDR3 loops (median 15 vs 13 residues) and unique conformations enabled by the absence of a light chain. Box plots show RMSD for framework (Fr) and three CDR loops. AlphaFold achieves the best CDR3 performance (4.00 Å RMSD) followed closely by IgFold (4.25 Å). NanoNet, despite being trained specifically for nanobodies, shows worse CDR3 prediction (5.43 Å). DeepAb performs poorly (8.52 Å) as it was trained only on paired antibodies and has not encountered nanobody-specific conformations. All methods achieve $\lt 1 Å$ framework accuracy. Source: Ruffolo et al. 2023, Figure 2e.
| Method | CDR1 (Å) | CDR2 (Å) | CDR3 (Å) | Framework (Å) |
|---|---|---|---|---|
| IgFold | 1.04 | 0.88 | 4.25 | 0.68 |
| AlphaFold | 1.00 | 0.77 | 4.00 | 0.70 |
| NanoNet | 1.19 | 0.85 | 5.43 | 0.70 |
| DeepAb | 1.47 | 1.13 | 8.52 | 0.57 |
| RepertoireBuilder | 1.18 | 1.32 | 7.54 | 0.80 |
Nanobody CDR3 loops present greater challenges than paired antibody CDR H3, with all methods showing higher RMSD values. AlphaFold slightly outperforms IgFold on nanobodies (4.00 Å vs 4.25 Å CDR3 RMSD), while DeepAb—designed primarily for paired antibodies—struggles significantly (8.52 Å). NanoNet, despite being optimized specifically for nanobodies, underperforms both AlphaFold and IgFold, highlighting the value of IgFold’s large-scale pre-training.
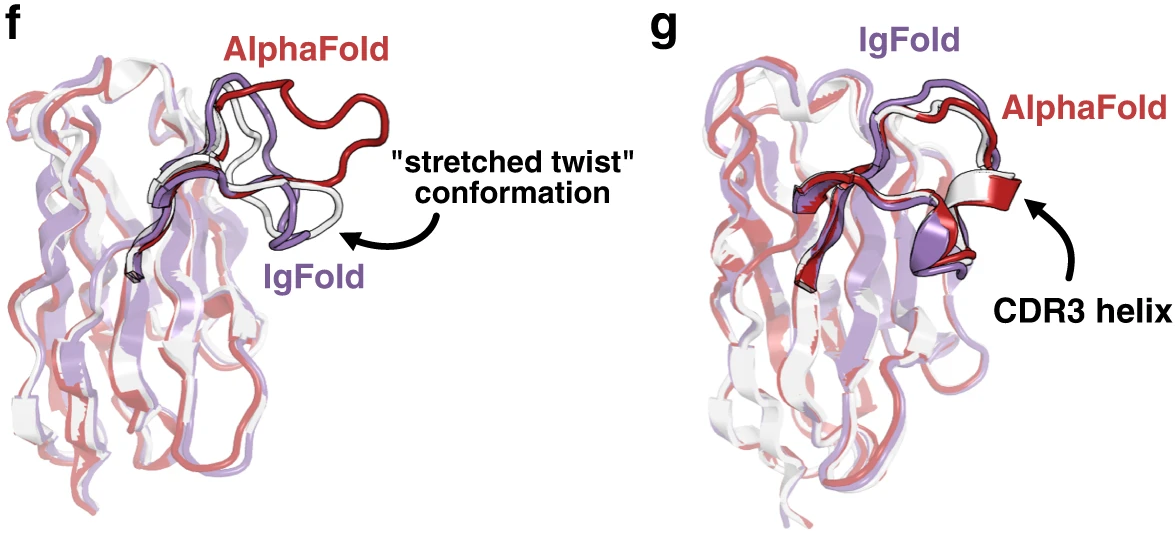
Figure 14: Nanobody CDR3 structural predictions. (f) Target 7AQZ with 15-residue CDR3 adopting the nanobody-specific "stretched-twist" conformation where the CDR3 loop bends to contact framework residues normally occupied by a light chain in paired antibodies. IgFold correctly predicts this nanobody-specific conformation (RMSD = 2.87 Å, purple) while AlphaFold predicts an extended loop inappropriate for single-domain antibodies (RMSD = 7.08 Å, pink). (g) Target 7AR0 with 17-residue CDR3 containing a short helical region. Both methods predict the correct overall loop conformation, but IgFold fails to predict the helical secondary structure (RMSD = 2.34 Å, purple) while AlphaFold accurately captures the helix (RMSD = 0.84 Å, pink). This highlights AlphaFold's advantage from training on diverse protein structures with varied secondary structure elements. Source: Ruffolo et al. 2023, Figure 2f-g.
Runtime comparisons demonstrate practical advantage
| Method | Paired antibody | Nanobody | Speedup vs AlphaFold |
|---|---|---|---|
| IgFold | 23 seconds | 15 seconds | ~19× |
| ABlooper | 174 seconds | N/A | 2.5× |
| AlphaFold-Multimer | 435 seconds | 345 seconds | 1× (baseline) |
| DeepAb | 750 seconds | 224 seconds | 0.5-1.5× |
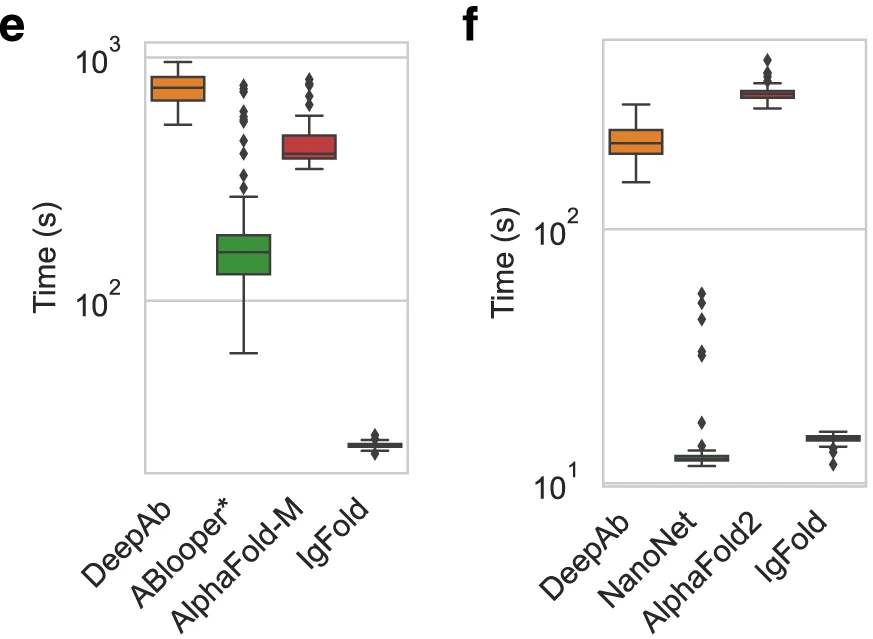
Figure 15: Runtime comparison for paired antibody structure prediction. Wall-clock time from sequence to full-atom structure on identical hardware (12-core CPU, 1× A100 GPU) for 197 paired antibody benchmark targets. IgFold achieves median runtime of 23 seconds—approximately 19× faster than ColabFold's AlphaFold-Multimer implementation (435 seconds) and 33× faster than DeepAb (750 seconds). ABlooper appears faster at 174 seconds but this excludes framework prediction time (provided by IgFold here), making true runtime comparison difficult. IgFold's speed advantage stems from eliminating MSA construction (AlphaFold's primary bottleneck) through pre-trained language model embeddings that encode equivalent evolutionary information. DeepAb's slow runtime reflects expensive Rosetta minimization of predicted geometric constraints. Runtime comparison on 71 nanobody benchmark targets. IgFold requires median 15 seconds—23× faster than AlphaFold (345 seconds). NanoNet achieves similar speed (15 seconds) but with notably worse CDR3 accuracy (5.43 Å vs 4.25 Å). DeepAb averages 224 seconds for nanobodies—faster than its paired antibody runtime due to shorter total sequence length. The modest runtime difference between paired antibodies and nanobodies for each method reflects approximately linear scaling with sequence length. Source: Ruffolo et al. 2023, Figure 4e-f.
IgFold’s ~19× speedup over AlphaFold-Multimer enables applications previously impractical with MSA-based methods. This speed advantage stems from eliminating the MSA construction step—by far the computational bottleneck in AlphaFold—through pre-trained language model embeddings that encode equivalent evolutionary information.
Distinctions from prior antibody-specific methods
IgFold represents a synthesis of ideas from prior work while introducing key innovations. Understanding these differences illuminates the design decisions underlying IgFold’s performance.
DeepAb: Same authors, different philosophy
DeepAb (Ruffolo et al., Patterns 2022), from the same Johns Hopkins group, predicts inter-residue geometric constraints (distances, angles) using a deep residual convolutional network, then uses Rosetta to realize structures from these constraints. This two-stage approach offers interpretability through attention visualization but requires ~12 minutes per structure due to Rosetta minimization.
IgFold’s key advance is end-to-end coordinate prediction—directly outputting backbone atom positions without intermediate constraint representation. This eliminates the structure realization bottleneck and enables gradient-based optimization of the entire pipeline. DeepAb also cannot incorporate template information, while IgFold’s IPA-based template usage provides flexible structural context.
ABlooper: CDR loops only
ABlooper (Abanades et al., Bioinformatics 2022) uses $\text{E(n)}$-Equivariant Graph Neural Networks to predict CDR loop coordinates but requires external framework scaffolds—it cannot predict complete antibody structures independently. ABlooper produces 5 predictions per loop and uses prediction diversity as confidence estimate.
IgFold predicts complete Fv structures end-to-end and provides dedicated error estimation rather than relying on prediction variance. ABlooper also cannot model nanobodies or incorporate loop templates, limiting its applicability for certain use cases.
NanoNet: Fast but limited scope
NanoNet (Cohen et al., Frontiers in Immunology 2022) uses 1D ResNets for extremely fast nanobody structure prediction—generating ~1 million structures in under 4 hours. However, it achieves this speed by predicting only backbone + $\text{Cβ}$ atoms and using simple one-hot encoding without language model pre-training.
IgFold’s 558M-sequence pre-training provides richer sequence representations that improve accuracy, particularly for the challenging CDR3 loops where IgFold achieves 4.25 Å RMSD versus NanoNet’s 5.43 Å. IgFold also handles both nanobodies and paired antibodies with a unified model.
Traditional methods: Template limitations
RosettaAntibody (Weitzner et al., Nature Protocols 2017) and related template-based methods select structural templates via sequence similarity, graft canonical CDR loops, and use de novo sampling for CDR H3. While RosettaAntibody produces physically realistic structures through energy minimization, it requires hours of computation for adequate H3 sampling and is fundamentally limited by template availability for non-canonical sequences.
IgFold’s neural network learns generalizable patterns from 558M sequences, enabling accurate prediction even for sequences dissimilar to any template. The template incorporation mechanism is optional and complementary—useful when partial structural information exists but not required for accurate prediction.
Advantages over AlphaFold and AlphaFold-Multimer
While AlphaFold-Multimer achieves comparable accuracy to IgFold on antibody structures, several factors favor IgFold for antibody-specific applications:
Speed: IgFold’s 19× speedup enables high-throughput applications. Predicting 1 million antibody structures takes ~6 days with IgFold versus ~4 months with AlphaFold-Multimer on equivalent hardware.
No MSA required: AlphaFold’s accuracy depends on MSA quality, but antibodies pose unique challenges—each antibody evolves independently within a single organism, lacking the evolutionary relationships that MSAs capture for other proteins. IgFold’s antibody-specific pre-training captures domain-relevant patterns directly.
No chain-swapping artifacts: AlphaFold-Multimer occasionally predicts C-terminal strand swaps between heavy and light chains—14 cases observed in IgFold’s benchmark set. IgFold’s antibody-specific architecture avoids this failure mode.
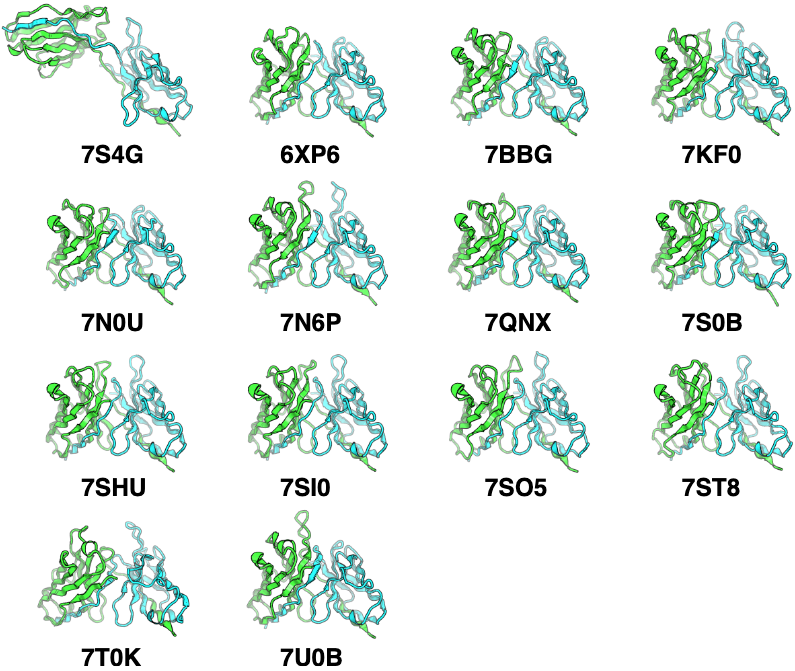
Figure 16: C-terminal strand swapping artifacts in AlphaFold-Multimer predictions. Representative examples of 14 paired antibody benchmark targets where AlphaFold-Multimer incorrectly predicts strand swaps between heavy (green) and light (cyan) chain C-terminal beta strands. In native antibody structures, each chain's C-terminal strand participates in its own immunoglobulin domain beta barrel. AlphaFold-Multimer occasionally predicts swapped architectures where heavy chain C-terminus integrates into light chain beta barrel and vice versa. This systematic error mode occurs in ~7% of benchmark predictions (14/197 targets) and results in elevated framework RMSD despite otherwise accurate structure prediction. IgFold's antibody-specific architecture does not exhibit this failure mode, likely due to training exclusively on correctly folded antibody structures rather than general protein complexes. Source: Ruffolo et al. 2023, Figure S5.
Complementary predictions: The 20% of CDR H3 loops where IgFold and AlphaFold-Multimer disagree by >4 Å suggests ensemble approaches combining both methods could improve coverage of conformational space.
Recent developments build on IgFold’s foundation
Since IgFold’s publication, several methods have advanced antibody structure prediction further:
ImmuneBuilder (Abanades et al., Communications Biology 2023) uses a modified AlphaFold-Multimer architecture achieving CDR H3 RMSD of 2.81 Å—comparable to IgFold—while running 100× faster than AlphaFold-Multimer.
H3-OPT (Chen et al., eLife 2024) combines AlphaFold2 with protein language models through confidence-based model selection, achieving the best reported CDR H3 accuracy of 2.24 Å mean RMSD.
BALMFold (Briefings in Bioinformatics 2024) trains on 336 million antibody sequences and runs 6× faster than IgFold while outperforming it on several benchmarks.
AlphaFold3 (2024) shows improved antibody-antigen docking success rates—up to 60% with extensive sampling—and median unbound CDR H3 RMSD of 2.9 Å for antibodies and 2.2 Å for nanobodies.
These developments validate IgFold’s central insight: antibody-specific pre-training on massive sequence databases, combined with geometric deep learning architectures, provides a powerful foundation for accurate and fast structure prediction.
Large-scale structure predictions expand structural coverage
IgFold’s practical impact is demonstrated through large-scale predictions:
- 104,994 paired antibody structures from OAS (35,731 human, 16,356 mouse, 52,907 rat) clustered at 95% identity
- 1,340,180 human antibody structures from Jaffe et al.’s repertoire sequencing study
These ~1.4 million predicted structures represent a 500-fold expansion over experimentally determined antibody structures. The median predicted CDR H3 RMSD for the 1.3M human antibodies is 1.95 Å—indicating high confidence predictions for most structures.
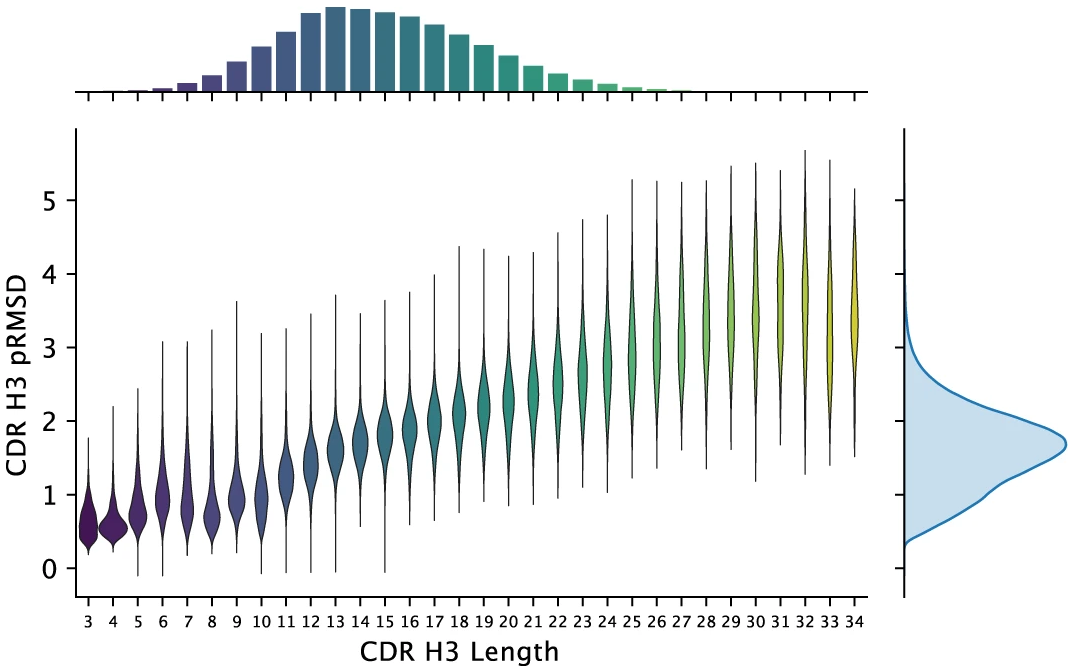
Figure 17: Predicted accuracy distribution for 1.3 million human antibody structures. Distribution of CDR H3 loop lengths (x-axis, histogram at top) and predicted RMSD (y-axis) for 1,340,180 unique paired human antibody sequences from four unrelated individuals. Violin plots show predicted RMSD distribution for each loop length. Median CDR H3 length is 13 residues with median predicted RMSD of 1.95 Å, indicating high-confidence predictions for the majority of structures. Predicted RMSD increases with loop length as expected—short loops (3-8 residues) typically predicted with <1.5 Å error while long loops (>20 residues) show higher uncertainty (>3 Å). This dataset represents a 500-fold expansion over experimentally determined antibody structures (~2,500 in PDB as of 2022), enabling unprecedented large-scale structural analysis of human antibody repertoires. Source: Ruffolo et al. 2023, Figure 4g.
This structural database enables new analyses: clustering antibody repertoires by predicted structure rather than sequence alone, identifying structural convergence in immune responses to specific antigens, and training downstream models for antibody property prediction with structural features.
Applications for antibody engineering and therapeutics
IgFold’s speed and accuracy enable integration into antibody discovery and optimization workflows:
Structure-guided design: Predicted structures inform rational design of CDR mutations, framework humanization, and stability engineering. Confidence scores guide experimental prioritization.
Docking and epitope prediction: Antibody structures from IgFold can seed docking calculations to predict antibody-antigen complexes. Recent studies show that using ensembles from multiple structure predictors improves docking success rates.
Repertoire analysis: Large-scale structure prediction enables structure-based clustering of immune repertoires, potentially revealing structural convergence patterns invisible to sequence-based analysis.
Lab-in-the-loop optimization: Integration with generative models and active learning enables iterative antibody optimization cycles combining computational prediction with experimental validation.
The tFold system (Wu et al., Nat. Commun. 2025) demonstrates end-to-end antibody design: de novo designed antibodies achieved nanomolar binding affinities against PD-1, PD-L1, and SARS-CoV-2 RBD, validating the utility of fast structure prediction in practical antibody engineering.
Conclusion
IgFold demonstrates that antibody-specific deep learning, leveraging massive natural antibody sequence databases and modern geometric attention mechanisms, can match or exceed general-purpose protein structure predictors while running nearly 20 times faster. The key innovations—AntiBERTy pre-training on 558M sequences, graph transformers with triangle multiplicative updates, and IPA-based template incorporation—create a unified framework for accurate, fast antibody structure prediction.
The practical impact extends beyond benchmarks. By enabling structure prediction for millions of antibodies, IgFold transforms structure-based analysis from boutique technique to routine capability. The confidence-aware predictions guide experimental prioritization, while the complementary predictions between IgFold and AlphaFold suggest ensemble strategies for covering conformational space.
Looking forward, the field continues advancing rapidly. Methods like H3-OPT, BALMFold, and AlphaFold3 build on similar principles while pushing accuracy further. The central insight validated by IgFold—that antibody-specific pre-training on massive sequence databases provides essential inductive bias for structure prediction—will likely remain foundational as antibody structure prediction matures into reliable computational infrastructure for therapeutic antibody development.
References
-
Ruffolo JA, Chu LS, Mahajan SP, Gray JJ. Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies. Nature Communications. 2023;14:2389. DOI: 10.1038/s41467-023-38063-x
-
Ruffolo JA, Sulam J, Gray JJ. Antibody structure prediction using interpretable deep learning. Patterns. 2022;3(2):100406. DOI: 10.1016/j.patter.2021.100406
-
Abanades B, Georges G, Bujotzek A, Deane CM. ABlooper: Fast accurate antibody CDR loop structure prediction with accuracy estimation. Bioinformatics. 2022;38(7):1877-1880. DOI: 10.1093/bioinformatics/btac016
-
Cohen T, Halfon M, Schneidman-Duhovny D. NanoNet: Rapid and accurate end-to-end nanobody modeling by deep learning. Frontiers in Immunology. 2022;13:958584. DOI: 10.3389/fimmu.2022.958584
-
Abanades B, Wong WK, Boyles F, Georges G, Bujotzek A, Deane CM. ImmuneBuilder: Deep-learning models for predicting the structures of immune proteins. Communications Biology. 2023;6:575. DOI: 10.1038/s42003-023-04927-7
-
Chen H, et al. H3-OPT: Accurate prediction of CDR-H3 loop structures of antibodies with deep learning. eLife. 2024;13:e91512. DOI: 10.7554/eLife.91512
-
Jumper J, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583-589. DOI: 10.1038/s41586-021-03819-2
-
Lin Z, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023;379:1123-1130. DOI: 10.1126/science.ade2574
-
Weitzner BD, et al. Modeling and docking of antibody structures with Rosetta. Nature Protocols. 2017;12:401-416. DOI: 10.1038/nprot.2016.180
-
Kovaltsuk A, et al. Observed Antibody Space: A resource for data mining next-generation sequencing of antibody repertoires. Journal of Immunology. 2018;201(8):2502-2509. DOI: 10.4049/jimmunol.1800708
-
Olsen TH, Boyles F, Deane CM. Observed Antibody Space: A diverse database of cleaned, annotated, and translated unpaired and paired antibody sequences. Protein Science. 2022;31:141-146. DOI: 10.1002/pro.4205
-
Yin R, Pierce BG. Evaluation of AlphaFold antibody–antigen modeling with implications for improving predictive accuracy. Protein Science. 2024;33:e4865. DOI: 10.1002/pro.4865
-
Chungyoun M, Gray JJ. What does AlphaFold3 learn about antibody and nanobody docking, and what remains unsolved? mAbs. 2025;17:2545601. DOI: 10.1080/19420862.2025.2545601
-
Giulini M, et al. Towards the accurate modelling of antibody-antigen complexes from sequence using machine learning and information-driven docking. Bioinformatics. 2024;40:btae583. DOI: 10.1093/bioinformatics/btae583
-
Elnaggar A, et al. ProtTrans: Towards Cracking the Language of Life’s Code Through Self-Supervised Deep Learning and High Performance Computing. IEEE Transactions on Pattern Analysis and Machine Intelligence. 2022;44:7112-7127. DOI: 10.1109/TPAMI.2021.3095381
Enjoy Reading This Article?
Here are some more articles you might like to read next: