Protein Language Models
A Technical Guide for Enthusiasts
Protein language models (PLMs) have fundamentally transformed computational protein science by learning evolutionary constraints directly from amino acid sequences—without requiring multiple sequence alignments. These deep learning models, trained on hundreds of millions of protein sequences using self-supervised objectives borrowed from natural language processing, now rival or exceed traditional methods for tasks ranging from contact prediction to three-dimensional structure determination. Most remarkably, models like ESMFold can predict atomic-resolution protein structures from single sequences in seconds, achieving accuracy that approaches the gold-standard AlphaFold2 while running 60× faster.
This guide provides computational biologists with the architectural depth and mathematical formulations they need to extend these models, while giving structural biologists the conceptual foundations to understand what PLMs learn and how to apply them effectively.
What makes protein language models different from NLP models
PLMs adapt transformer architectures—the same technology underlying GPT and BERT—to treat amino acid sequences as “sentences” in the language of life. The core insight is that evolutionary pressure constrains which amino acids can appear at each position in a protein: destabilizing mutations at one site must be compensated by changes at contacting positions, creating correlated substitution patterns that PLMs implicitly learn during training.
Unlike natural language models trained to predict words, PLMs are trained on the masked language modeling (MLM) objective: randomly mask 15% of amino acids, then predict the original residue from surrounding context. This forces the model to learn which amino acids are compatible at each position given the sequence context—essentially encoding the rules of protein chemistry and physics into attention patterns.
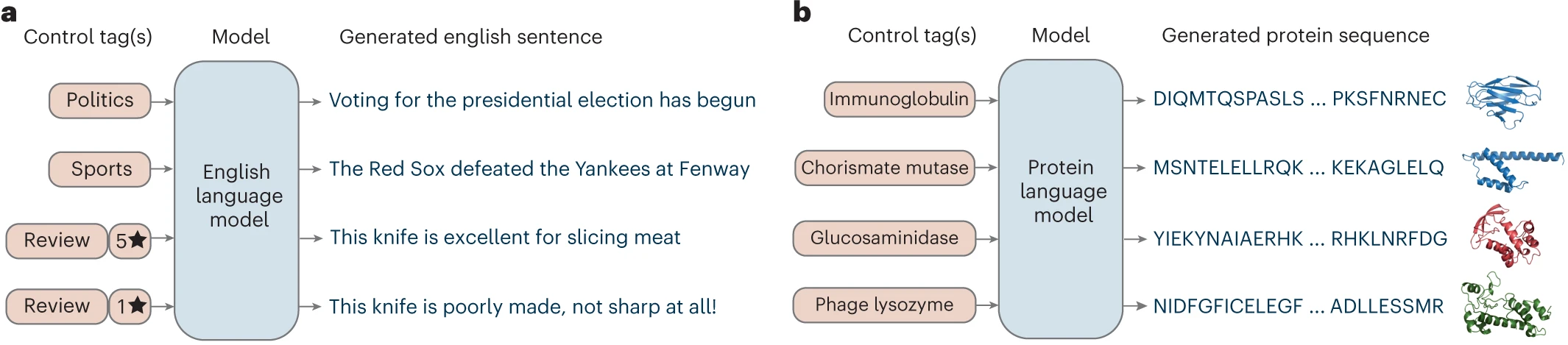
The key distinction from traditional sequence analysis is that PLMs require no explicit evolutionary information at inference time. While methods like PSI-BLAST build position-specific scoring matrices (PSSMs) through iterative database searches and coevolution methods like DCA require deep multiple sequence alignments, PLMs compress evolutionary statistics from millions of training sequences into their parameters. A forward pass through ESM-2 effectively performs an “implicit MSA” lookup instantaneously.
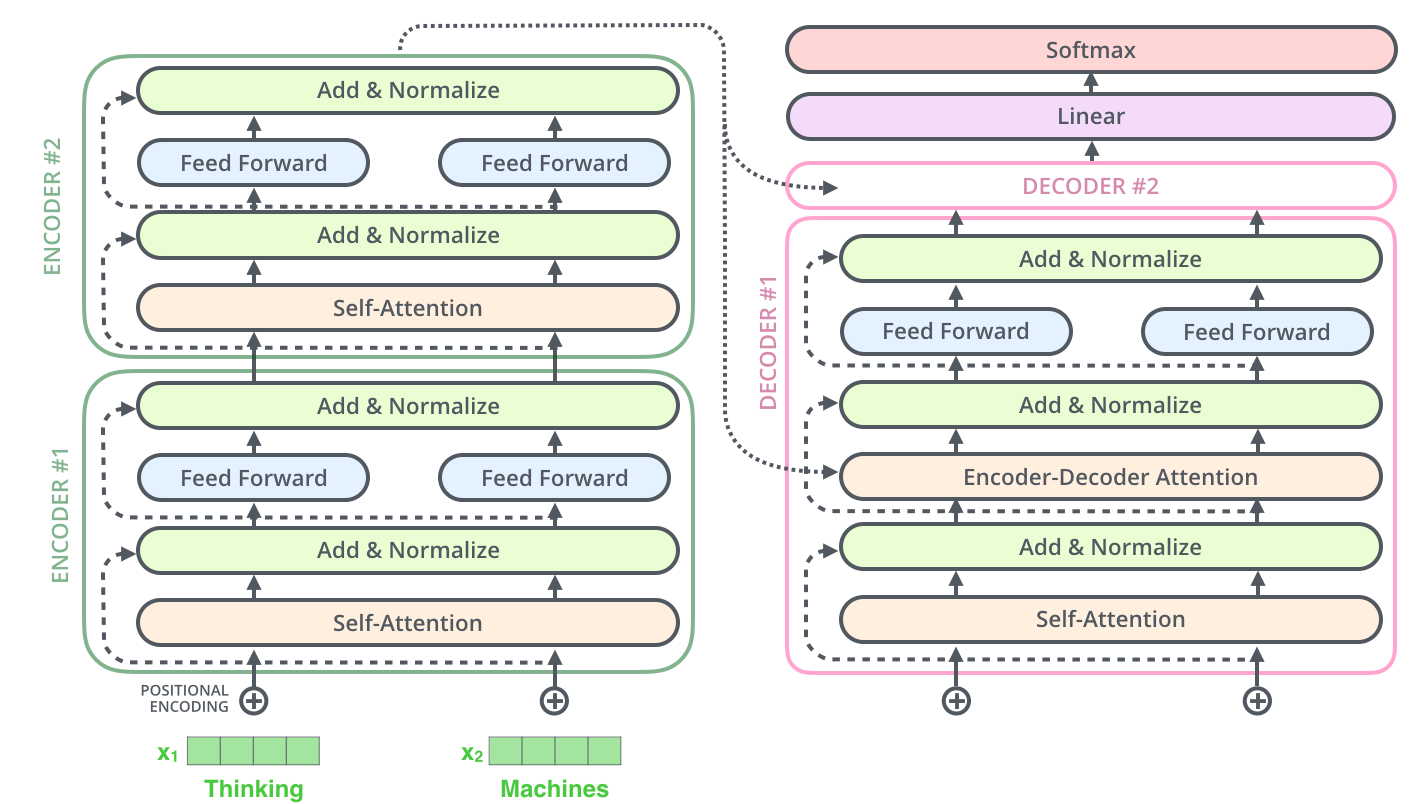
Transformer architecture fundamentals for protein sequences
The transformer’s self-attention mechanism enables PLMs to capture dependencies between amino acids regardless of their sequential distance—critical for proteins where residues far apart in sequence often contact in three-dimensional space. The mathematical formulation of scaled dot-product attention is:
\[\text{Attention}(Q, K, V) = \text{softmax}\left(\frac{QK^T}{\sqrt{d_k}}\right) V\]where Q (queries), K (keys), and V (values) are linear projections of input embeddings. For a protein sequence, each amino acid’s embedding is projected into these three representations. The dot product $QK^T$ measures pairwise compatibility between positions—residue pairs with high attention weights often correspond to structural or functional relationships.
Multi-head attention extends this by running h parallel attention operations with different learned projections:
\[\text{MultiHead}(Q, K, V) = \text{Concat}(\text{head}_1, ..., \text{head}_h) W_O\]ESM-2’s 650M parameter model uses 33 transformer layers with 20 attention heads and 1,280-dimensional embeddings. Different heads specialize for different biological features: some capture local secondary structure motifs, others encode long-range tertiary contacts, and specific heads focus on binding sites and functional regions.
Positional encodings tell the model where each residue sits in the sequence. ESM-1b used learned absolute positional embeddings, limiting sequences to 1,022 residues. ESM-2 switched to Rotary Position Embeddings (RoPE), which encode relative positions through rotations in embedding space:
\[\text{RoPE}(x_{\text{pos}}) = x_{\text{pos}} \cdot \cos(\theta_{\text{pos}}) + \hat{x}_{\text{pos}} \cdot \sin(\theta_{\text{pos}})\]RoPE enables arbitrary-length sequences at inference, a significant advantage for processing large proteins.
The ESM model family sets the benchmark
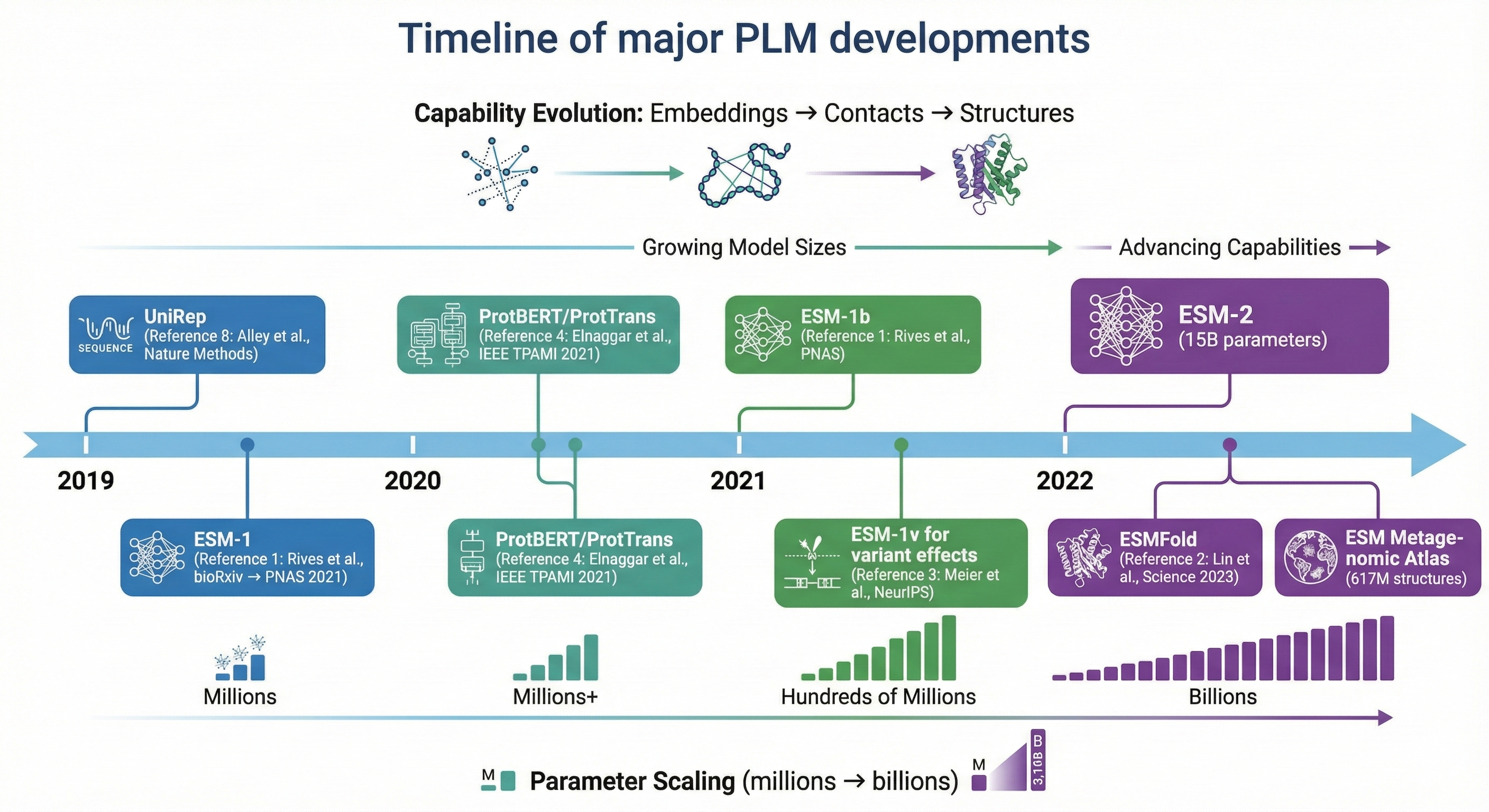
Meta AI’s Evolutionary Scale Modeling (ESM) series represents the most influential PLM family, demonstrating that biological structure and function emerge from scaling unsupervised learning to hundreds of millions of sequences.
ESM-1b (Rives et al., 2021, PNAS) established that a 650M-parameter transformer trained on UniRef50’s 250 million sequences could predict secondary structure at 71.6% accuracy and extract contact information from attention patterns without any structural supervision. The model achieved effective cross-entropy of ~8.46, indicating substantial understanding of amino acid co-occurrence patterns.
ESM-1v (Meier et al., 2021, NeurIPS) specialized for variant effect prediction by training on UniRef90—the higher 90% clustering threshold captures more sequence variation relevant for predicting mutation effects. Using zero-shot log-odds scoring (log P(mutant) - log P(wildtype)), ESM-1v achieved Spearman ρ = 0.52 across 41 deep mutational scanning datasets, matching state-of-the-art methods that require expensive MSA construction.
ESM-2 (Lin et al., 2023, Science) scaled to 15 billion parameters across 48 layers, demonstrating consistent scaling laws where larger models encode increasingly accurate structural information. A key architectural improvement was switching to RoPE positional embeddings and pre-layer normalization. Remarkably, ESM-2’s 150M model outperforms ESM-1b’s 650M model on structure prediction—suggesting architectural improvements matter as much as scale.
| Model | Parameters | Layers | Training Data | Key Innovation |
|---|---|---|---|---|
| ESM-1b | 650M | 33 | UniRef50 (250M seqs) | First large-scale PLM showing structural emergence |
| ESM-1v | 650M | 33 | UniRef90 (98M seqs) | Zero-shot variant effect prediction |
| ESM-2 | 8M–15B | 6–48 | UniRef50 (2021) | RoPE embeddings, scaling to 15B parameters |
ProtTrans family explores diverse architectures
The ProtTrans project (Elnaggar et al., 2021, IEEE TPAMI) systematically evaluated different transformer architectures for proteins, training on unprecedented computational resources including Google TPU Pods with up to 1,024 cores.

ProtBERT-BFD (420M parameters, 30 layers) trained on the Big Fantastic Database containing 2.1 billion metagenomic sequences—8× more proteins than ESM-1b’s training set. This massive sequence diversity improved capture of rare evolutionary patterns, achieving 83% three-state secondary structure accuracy on CB513.
ProtT5-XL-UniRef50 emerged as the best performer across most benchmarks. This 3-billion parameter encoder-decoder model adapts Google’s T5 architecture, initially training on BFD for 1.2 million steps before fine-tuning on UniRef50. For feature extraction, encoder embeddings outperform decoder embeddings, and the model runs efficiently in half-precision (fp16) without accuracy degradation.
Ankh (Elnaggar et al., 2023) demonstrated that protein-specific optimizations outweigh raw scale. By optimizing masking strategies, architecture hyperparameters, and training data selection, Ankh achieves competitive performance with <10% of the training compute of larger models. The key insight: careful engineering matters as much as brute-force scaling for protein applications.
Other architecturally distinct models include UniRep (Alley et al., 2019, Nature Methods), a multiplicative LSTM that pioneered the concept of “evotuning”—fine-tuning on evolutionarily related sequences to boost task-specific performance. ProGen (Madani et al., 2023, Nature Biotechnology) used GPT-style autoregressive training to enable controllable protein generation, producing artificial lysozymes with 73% functionality at sequence identities as low as 31% to natural proteins.
How attention patterns encode structural contacts
Perhaps the most striking finding in PLM research is that transformer attention heads learn to identify residue-residue contacts without any structural supervision. Vig et al. (2020) demonstrated that a single attention head in BERT-based protein models aligns 28% of its attention weight to structural contacts, far exceeding the 1.3% background contact probability.
The mathematical connection runs deep. A factored attention layer is equivalent to a Pairwise Markov Random Field, with Potts models—the standard formulation for Direct Coupling Analysis—as a sparse special case. The attention weight $A_{ij}$ between positions $i$ and $j$ effectively captures the same coevolutionary coupling that DCA extracts from MSAs:
\[\mathrm{P(a_1, ..., a_L)} = \frac{1}{Z} \left(\sum_i h_i(a_i) + \sum_{i\lt j} J_{ij}(a_i, a_j) \right)\]where the coupling matrix $J$ directly predicts contacts. Attention heads learn these couplings implicitly from the masked language modeling objective.
To extract contact predictions from attention:
-
Aggregate attention matrices across heads and layers: $A_{\text{agg}} = \frac{1}{|H||L|}\sum_{h,l} A^{(h,l)}$.
-
Symmetrize: $\hat{A} = \frac{1}{2}(A + A^T)$.
-
Apply Average Product Correction: $\hat A_{ij}^{APC} = A_{ij} - \frac{\bar{A}_i \cdot \bar{A}_j}{\bar{A}}$.
-
Rank pairs and select top L/k predictions.
ESM-1b achieves ~45% precision at L/5 for long-range contacts from attention patterns alone—competitive with CCMpred’s 42% from explicit coevolution analysis on MSAs.
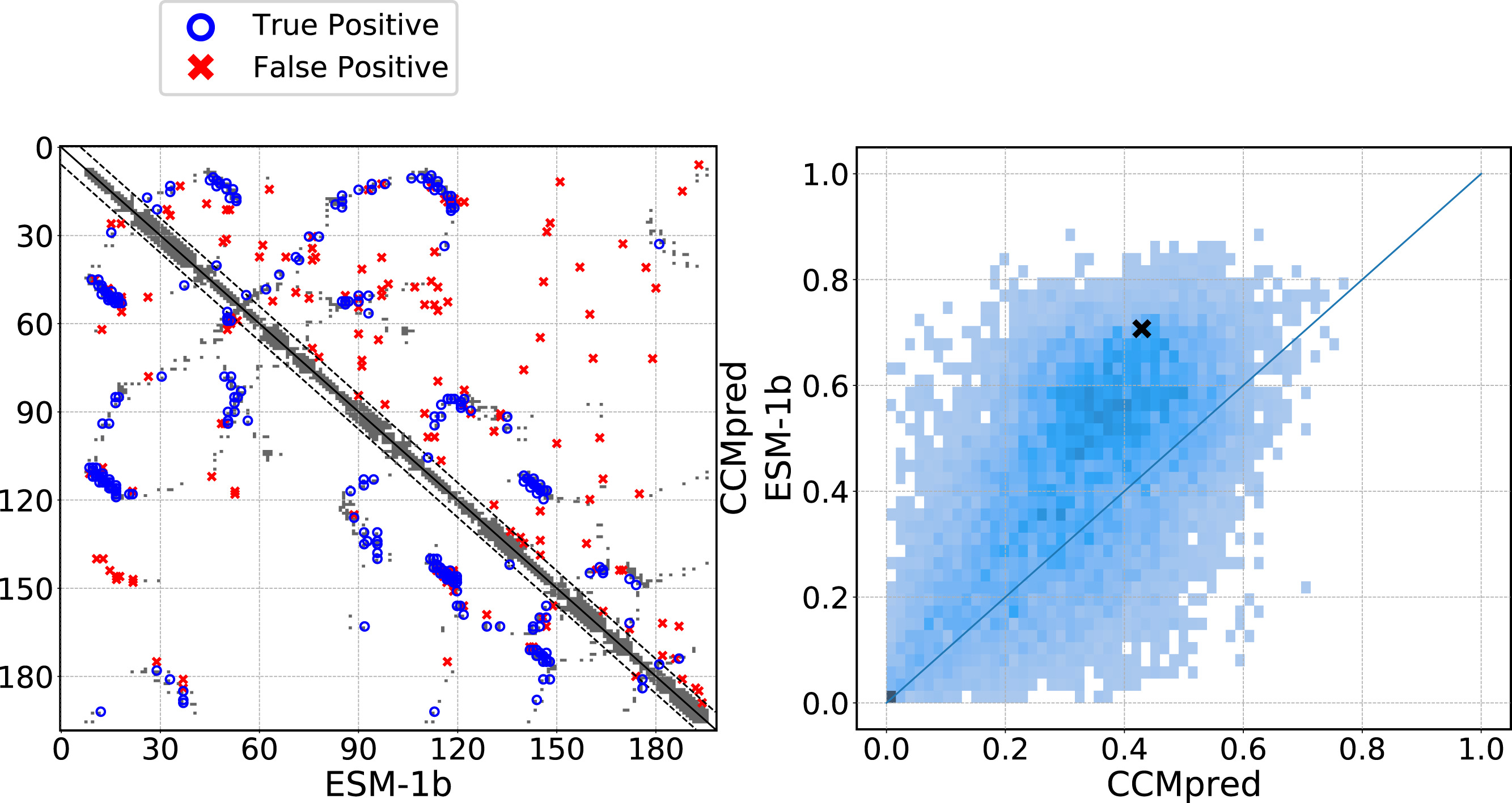
ESMFold predicts structures from single sequences
ESMFold combines ESM-2’s learned representations with a structure prediction module to predict atomic-resolution protein structures from single sequences. The architecture has three components:
Frozen ESM-2 language model (3B parameters) produces per-residue embeddings encoding evolutionary and structural information learned during pretraining. These embeddings replace the MSA processing that AlphaFold2 requires.
Folding trunk (48 blocks) converts ESM-2 embeddings into pair representations using a simplified Evoformer architecture. Unlike AlphaFold2’s full Evoformer, ESMFold removes the expensive axial attention over MSA rows—unnecessary since there’s no MSA to process.
Structure module (8 blocks with shared weights) uses Invariant Point Attention (IPA) to predict 3D coordinates:
\[\text{IPA}(q, k, v, T) = \text{softmax}\left(\frac{1}{\sqrt{c}}\left(q^Tk + w_L \cdot b_{ij} + w_C \sum_p \|T_i \circ \vec{q}_p - T_j \circ \vec{k}_p\|^2\right)\right)v\]where $T_i, T_j$ are residue coordinate frames. IPA’s key property is SE(3) invariance: global rotations and translations leave attention weights unchanged, ensuring predictions don’t depend on arbitrary reference frames.
The model trains on a mixture of 25,000 PDB clusters and 12 million AlphaFold2 predictions as distillation targets, with 75% weight on predicted structures. The Frame Aligned Point Error (FAPE) loss penalizes coordinate deviations under multiple local reference frames, enforcing both global and local structural accuracy.
| Metric | ESMFold | AlphaFold2 (single seq) | AlphaFold2 (full MSA) |
|---|---|---|---|
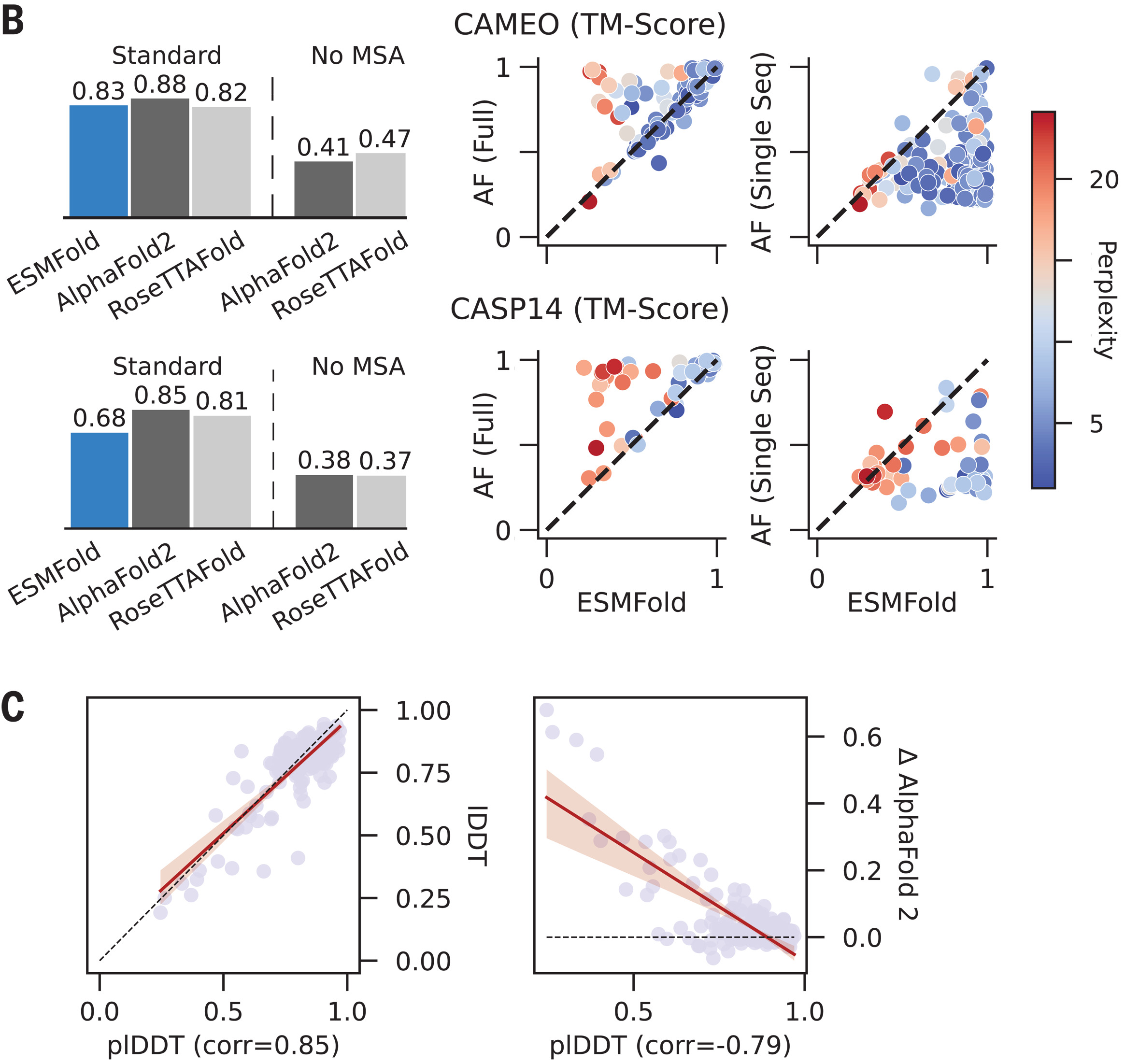
| CAMEO TM-score | 0.83 | 0.73 | 0.89 |
| CASP14 TM-score | 0.68 | 0.37 | 0.85 |
| Inference time (500 residues) | ~8 seconds | ~30+ minutes | ~30+ minutes |
ESMFold excels for orphan proteins lacking homologs, where MSA-based methods fail. On CASP14 targets, ESMFold’s 0.68 TM-score dramatically outperforms AlphaFold2’s 0.37 when MSAs are removed—demonstrating that PLM embeddings effectively encode evolutionary information that would otherwise require explicit MSA construction.
What different transformer layers learn
Probing studies reveal hierarchical learning across PLM layers, with biological complexity emerging progressively:
Early layers (1–10) capture local features: amino acid physicochemical properties, immediate neighbors, and secondary structure elements. Linear classifiers on these representations achieve ~78% secondary structure accuracy.
Middle layers (10–25) encode secondary structure context and medium-range dependencies. Secondary structure prediction accuracy peaks here, reaching 84–86% for ProtT5 and ESM-2.
Deep layers (25–33) integrate long-range tertiary contacts, remote homology relationships, and global fold information. Contact prediction precision rises sharply in final layers, suggesting late integration of three-dimensional structural knowledge.
A remarkable finding from Heinzinger et al. (2021) is that adding explicit MSA information does not improve ProtT5’s secondary structure predictions—the model has already internalized equivalent evolutionary statistics through pretraining on millions of sequences.
Comparison with traditional sequence analysis methods
Understanding when PLMs outperform—and when they don’t—requires comparing against established methods.
Multiple sequence alignments capture evolutionary information through explicit sequence comparison. Tools like MAFFT and HHblits identify conserved regions and variable positions, enabling coevolution analysis. However, MSA quality depends critically on homolog availability: Neff/L > 1 (effective sequences divided by length) is typically required for reliable predictions. For orphan proteins with few homologs, MSA-based methods fail.
PSI-BLAST and profile HMMs build position-specific scoring matrices through iterative database searches. These methods detect ~2× more homologs than pairwise searches below 40% identity but suffer from profile drift (iterative accumulation of false positives) and require substantial computational time for database searches.
Coevolution analysis (DCA, CCMpred, GREMLIN) disentangles direct from indirect correlations in MSAs using maximum entropy models. Performance requires deep MSAs—typically >500 effective sequences—and degradation above ~50,000 sequences due to convergence issues. CCMpred achieves ~42% precision at L/5 for long-range contacts; adding neural networks (MetaPSICOV) improves this to ~56%.
PLMs fundamentally change the tradeoffs:
- Speed: Single forward pass (~seconds) vs. MSA construction (minutes to hours)
- Orphan proteins: PLMs maintain predictive power; MSA-based methods fail
- Interpretability: Traditional methods provide probabilistic scores and E-values; PLM confidence metrics are less principled
- Accuracy ceiling: AlphaFold2 with deep MSAs still outperforms ESMFold on challenging targets (0.85 vs. 0.68 TM-score on CASP14)
The emerging consensus is that PLMs and traditional methods are complementary: PLMs provide speed and handle orphan proteins; MSA-based methods leverage explicit evolutionary signals when available. State-of-the-art pipelines increasingly combine both paradigms.
Computational requirements and practical considerations
Training large PLMs requires substantial resources. ESM-2’s 15B model required massive distributed training across GPU clusters, with exact compute costs comparable to GPT-3 scale (~hundreds of thousands of GPU-hours). Training ProtT5-XL used Google TPU Pods with up to 1,024 cores.
Memory scaling presents the primary bottleneck: standard attention is O(n²) in sequence length. For ESM-1b on a 48GB A100:
- 512 residues: Comparable to CNNs
- 1,024 residues: ~4× more memory than CNN alternatives
- 2,048 residues: Out-of-memory
Optimization techniques for inference include:
- FlashAttention: 4–9× faster inference, 3–14× lower memory through I/O-aware algorithms
- 4-bit quantization: 2–3× memory reduction while preserving variant prediction accuracy
- Sequence packing: Handles 100,000+ tokens without padding waste
For long sequences exceeding context limits, sliding window approaches with overlapping chunks and mean aggregation across chunks provide practical solutions.
The road ahead for protein language models
PLMs have demonstrated that evolutionary information sufficient for structure prediction and function annotation is learnable from sequence data alone—a finding with profound implications for protein science. The ESM Metagenomic Atlas now contains >617 million predicted structures from metagenomic sequences, expanding structural coverage of protein space by orders of magnitude.
Current frontiers include multimodal models combining sequence, structure, and function; protein generation for therapeutic design; and integration with physics-based methods for dynamics and binding prediction. As models continue scaling and architectural innovations emerge, the gap between PLM-based and MSA-based methods continues narrowing—with PLMs offering transformative advantages in speed and applicability to proteins lacking evolutionary context.
References
-
Rives A, Meier J, Sercu T, et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences. 2021;118(15):e2016239118. doi:10.1073/pnas.2016239118
-
Lin Z, Akin H, Rao R, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023;379(6637):1123-1130. doi:10.1126/science.ade2574
-
Meier J, Rao R, Verkuil R, et al. Language models enable zero-shot prediction of the effects of mutations on protein function. Advances in Neural Information Processing Systems. 2021; 34:29287-29303.
-
Elnaggar A, Heinzinger M, Dallago C, et al. ProtTrans: Toward understanding the language of life through self-supervised learning. IEEE Transactions on Pattern Analysis and Machine Intelligence. 2021;44(10):7112-7127. doi:10.1109/TPAMI.2021.3095381
-
Vig J, Madani A, Varber LR, et al. BERTology meets biology: Interpreting attention in protein language models. arXiv preprint. 2020; arXiv:2006.15222.
-
Jumper J, Evans R, Pritzel A, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596(7873):583-589. doi:10.1038/s41586-021-03819-2
-
Rao R, Bhattacharya N, Thomas N, et al. Evaluating protein transfer learning with TAPE. Advances in Neural Information Processing Systems. 2019; 32:9689-9701.
-
Alley EC, Khimulya G, Biswas S, AlQuraishi M, Church GM. Unified rational protein engineering with sequence-based deep representation learning. Nature Methods. 2019;16:1315-1322. doi:10.1038/s41592-019-0598-1
-
Madani A, Krause B, Greene ER, et al. Large language models generate functional protein sequences across diverse families. Nature Biotechnology. 2023;41:1099-1106. doi:10.1038/s41587-022-01618-2
-
Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need. Advances in Neural Information Processing Systems. 2017; 30:5998-6008. doi:10.48550/arXiv.1706.03762
-
Heinzinger M, Elnaggar A, Wang Y, et al. Modeling aspects of the language of life through transfer-learning protein sequences. BMC Bioinformatics. 2019;20:723. doi:10.1186/s12859-019-3220-8
-
Jones DT, Singh T, Kosciolek T, Sheridan S. MetaPSICOV: combining coevolution methods for accurate prediction of contacts and long range hydrogen bonding in proteins. Bioinformatics. 2015;31(7):999-1006. doi:10.1093/bioinformatics/btu791
-
Wu R, Ding F, Wang R, et al. High-resolution de novo structure prediction from primary sequence. bioRxiv. 2022. doi:10.1101/2022.07.21.500999
-
Elnaggar A, Essam H, Salah-Eldin W, et al. Ankh: Optimized protein language model unlocks general-purpose modelling. arXiv preprint. 2023; arXiv:2301.06568
-
Su J, Lu Y, Pan S, Murtadha A, Wen B, Liu Y. RoFormer: Enhanced transformer with rotary position embedding. arXiv preprint. 2021; arXiv:2104.09864
Enjoy Reading This Article?
Here are some more articles you might like to read next: