Inside ProteinMPNN
The neural network transforming computational protein design
Protein sequence design has entered a new era with ProteinMPNN, a Message-Passing Neural Network (MPNN) that solves the inverse folding problem—predicting amino acid sequences that will fold into a target backbone structure—with 52.4% native sequence recovery, dramatically outperforming Rosetta’s 32.9% while running 200 times faster. Published in Science in 2022 by Dauparas et al. from the Baker Lab, ProteinMPNN has become the de facto standard for sequence design in modern computational protein engineering pipelines. The method’s impact extends far beyond benchmarks: it has enabled the design of therapeutic binders, vaccine nanoparticles, and functional enzymes that were previously intractable, fundamentally reshaping how researchers approach de novo protein creation.
The inverse folding problem inverts AlphaFold’s challenge
While AlphaFold2 revolutionized forward folding—predicting three-dimensional structure from amino acid sequence—ProteinMPNN addresses the complementary inverse folding problem: given a desired backbone structure, what sequences will fold into it? This distinction matters profoundly because inverse folding is a one-to-many mapping. A single backbone can accommodate vastly different sequences (for a 100-residue protein, the theoretical sequence space spans $\mathrm{20^{100}}$ possibilities), yet only a tiny fraction will actually fold stably. The design challenge lies in navigating this astronomical space to find the rare sequences with high “designability”—those predicted to reliably adopt the target structure.
The inverse folding problem is fundamental to protein engineering. When researchers design novel protein structures using tools like RFdiffusion or want to stabilize natural proteins, they must ultimately find sequences encoding those structures. Before ProteinMPNN, Rosetta’s physics-based fixed backbone design required hours of computation and extensive expert intervention to achieve modest success rates. ProteinMPNN compressed this to seconds while dramatically improving outcomes.
Graph neural network architecture encodes backbone geometry through pairwise distances
ProteinMPNN represents proteins as sparse graphs where each residue becomes a node and edges connect to the k=48 nearest neighbors based on $\mathrm{C_α-C_α}$ Euclidean distances. This k-nearest neighbor construction scales linearly with protein length and captures the local and medium-range contacts critical for determining compatible sequences. Node embeddings initialize to zeros, with the network learning structural features entirely through message passing rather than explicit input features—a design choice that proved crucial for generalization.
The Backbone Graph.
The model takes the 3D coordinates of the protein backbone (N, C, C, O atoms). It builds a k-Nearest Neighbors (k-NN) graph where:
- Nodes: Represent individual amino acid residues.
- Edges: Connect residues that are close in 3D space (even if they are far apart in the sequence).
- Features: Distances, relative orientations, and dihedral angles.
The edge features encode 25 pairwise distances between backbone heavy atoms ($\mathrm{N, C_α, C, O}$) and a virtual $\mathrm{C_β}$ atom. For each edge connecting residues $i$ and $j$, the model computes all combinations of distances between these five atom types across both residues. These raw distances undergo transformation through radial basis functions (RBFs) using 16 Gaussian basis functions spaced from 0-20 Å, converting continuous distances into rich learnable representations. Additional positional encodings capture sequence-relative position (capped at ±32 residues) and a binary same-chain indicator for multi-chain complexes.
This distance-based representation was a key innovation over previous approaches. The original Structured Transformer from Ingraham et al. (2019) used backbone dihedral angles ($φ, ψ, ω$) and frame orientations. Replacing these with pure pairwise distances improved sequence recovery from 41.2% to 49.0%—a substantial jump attributed to distances providing better inductive bias for residue-residue interactions without the coordinate system dependencies of angles.
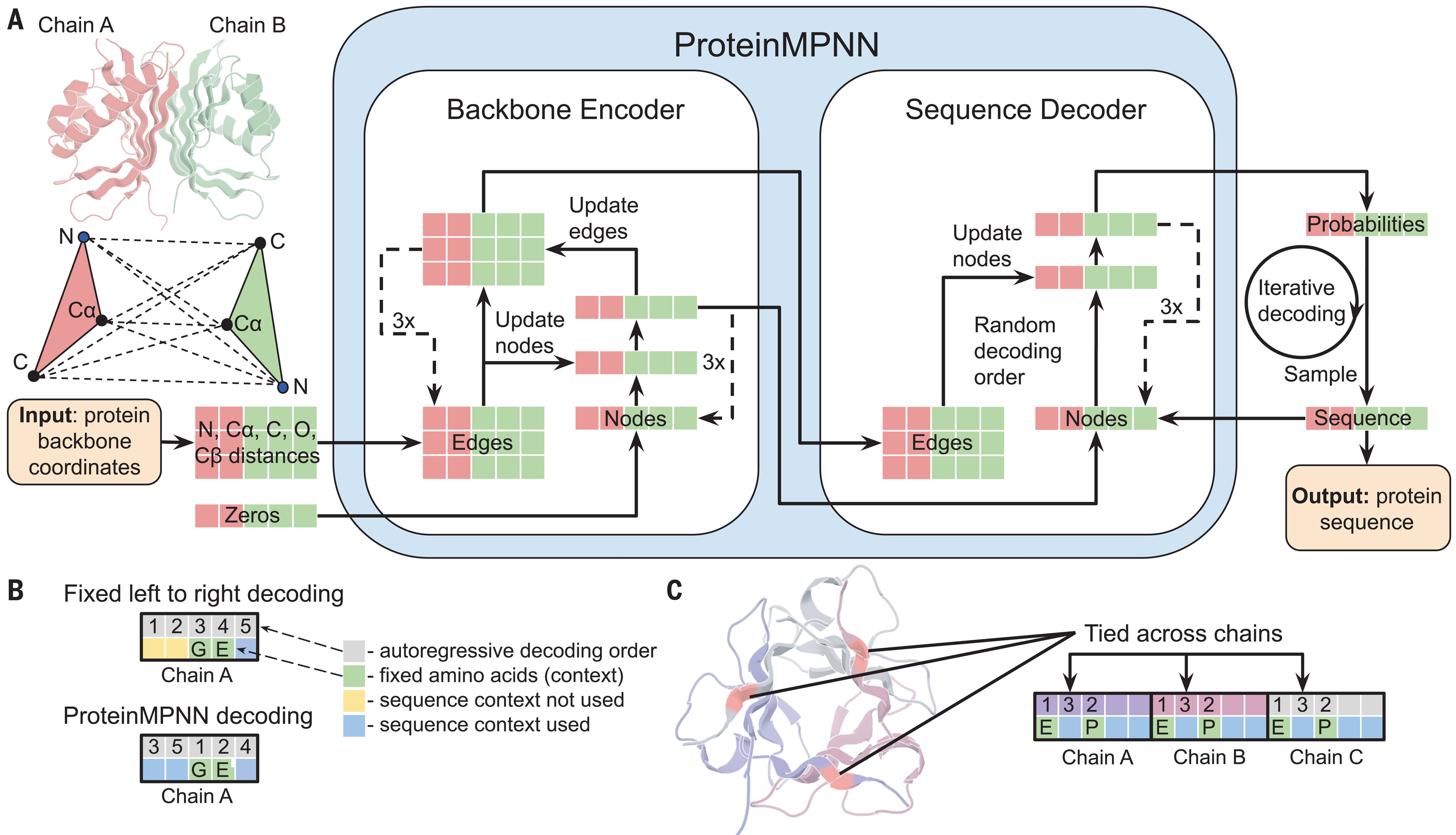
Three encoder layers refine both node and edge representations
The encoder consists of three message-passing layers that iteratively update both node and edge embeddings—another departure from earlier architectures that only updated nodes. Each encoder layer performs two sequential operations. First, messages from neighboring nodes aggregate to update node embeddings through a three-layer MLP operating on concatenated features (source node, target node, edge embedding). The aggregated messages divide by a scaling factor of 30 to stabilize training, followed by a residual connection and layer normalization. Second, an identical operation updates edge embeddings using the newly refined node states.
The mathematical formulation follows:
qij = GELU(Linear(concat(vi, vj, eij))) → MLP with 3n→m→m dimensions
dhi = Σj qij / 30 → Neighbor aggregation with scaling
vi = LayerNorm(vi + Dropout(dhi)) → Residual update with 10% dropout
This dual updating scheme—nodes informing edges and vice versa—contributed approximately 2% improvement in sequence recovery. The model uses GELU activations throughout, 128 hidden dimensions, and produces a compact architecture of only ~1.66 million parameters (compared to ResNet50’s 23 million), yet achieves state-of-the-art performance.
Autoregressive decoding generates sequences in random order
The decoder employs an autoregressive strategy, generating amino acids one position at a time conditioned on both the structural encoder output and previously generated residues. However, unlike language models that decode left-to-right, ProteinMPNN uses order-agnostic autoregressive decoding—the generation order is randomly sampled from all possible permutations during training. This innovation proves essential for practical applications: when designing binders, the target protein sequence is known and should inform the designed binder’s sequence, requiring flexible conditioning that fixed-order decoding cannot provide.
The decoder consists of three layers, each operating similarly to encoder layers but with causal masking ensuring positions can only attend to previously decoded residues in the current permutation. The final hidden state passes through a linear projection to 21 output classes (20 amino acids plus unknown), with softmax producing probability distributions. Temperature-controlled sampling enables trading off sequence diversity against recovery: lower temperatures (T=0.1) yield conservative, high-confidence designs while higher temperatures increase diversity at modest accuracy cost.
For symmetric homo-oligomers, ProteinMPNN introduces tied decoding where equivalent positions across chains share the same amino acid. The model averages logits from all tied positions before sampling, ensuring sequence identity at symmetric sites. This mechanism enabled successful design of cyclic oligomers from C2 to C6 symmetry and even tetrahedral nanoparticles exceeding one megadalton.
Summary: ProteinMPNN architecture.
The primary architecture consists of two main parts:1. Encoder:
- Processes the spatial 3D input structure.
- Encodes the backbone geometry into a contextual representation by using relative distances and orientations between atoms (N, Cα, C, O, and Cβ).
- Uses three layers with 128 hidden dimensions to build a sparse protein graph.
2. Decoder:
- Generates the amino acid sequence iteratively using an Auto-Regressive (AR) approach.
- Supports a freely specified decoding order, allowing users to fix specific parts of the sequence while the model designs the rest.
- Outputs a set of logits for each position, which can be sampled using strategies like greedy decoding, softmax sampling with temperature control, or beam search.
Training with coordinate noise prevents overfitting to crystallographic artifacts
A critical training innovation involved adding Gaussian noise (0.02-0.30 Å standard deviation) to backbone coordinates during training. This regularization addresses a subtle but important problem: crystallographic refinement programs use amino acid identity information when refining local backbone geometry, meaning crystal structures contain sequence-dependent geometric signatures that models can exploit. Without noise augmentation, ProteinMPNN overfit to these artifacts, achieving high recovery on crystal structures but failing on designed or predicted backbones.
The noise-recovery tradeoff proved nuanced. Models trained with 0.02 Å noise achieved highest native sequence recovery on test crystal structures, but models trained with 0.20-0.30 Å noise generated 2-3 times more sequences whose AlphaFold predictions reached 90-95 lDDT-Cα scores. The default recommended model (v_48_020) uses 0.20 Å noise, balancing crystal structure recovery with robustness to the imperfect backbones typical in design applications. This insight—that the best test-set metric doesn’t guarantee the best design outcomes—reflects deeper lessons about machine learning for protein engineering.
The training data comprised PDB structures with X-ray or cryo-EM resolution ≤3.5 Å, clustered at 30% sequence identity into approximately 25,361 clusters. The model trained on single GPUs using Adam optimization with 10% dropout and 10% label smoothing, demonstrating that massive compute is not prerequisite for impactful protein design methods.
Performance benchmarks establish ProteinMPNN as the sequence design standard
On a test set of 402 monomeric proteins, ProteinMPNN achieved 52.4% median sequence recovery compared to Rosetta’s 32.9%—a 59% relative improvement. Recovery correlates strongly with burial: deeply buried core residues approach 90-95% recovery while surface residues hover around 35%, reflecting the greater sequence constraints imposed by the protein interior. Critically, ProteinMPNN improves over Rosetta across all burial levels, not just in specific regimes.
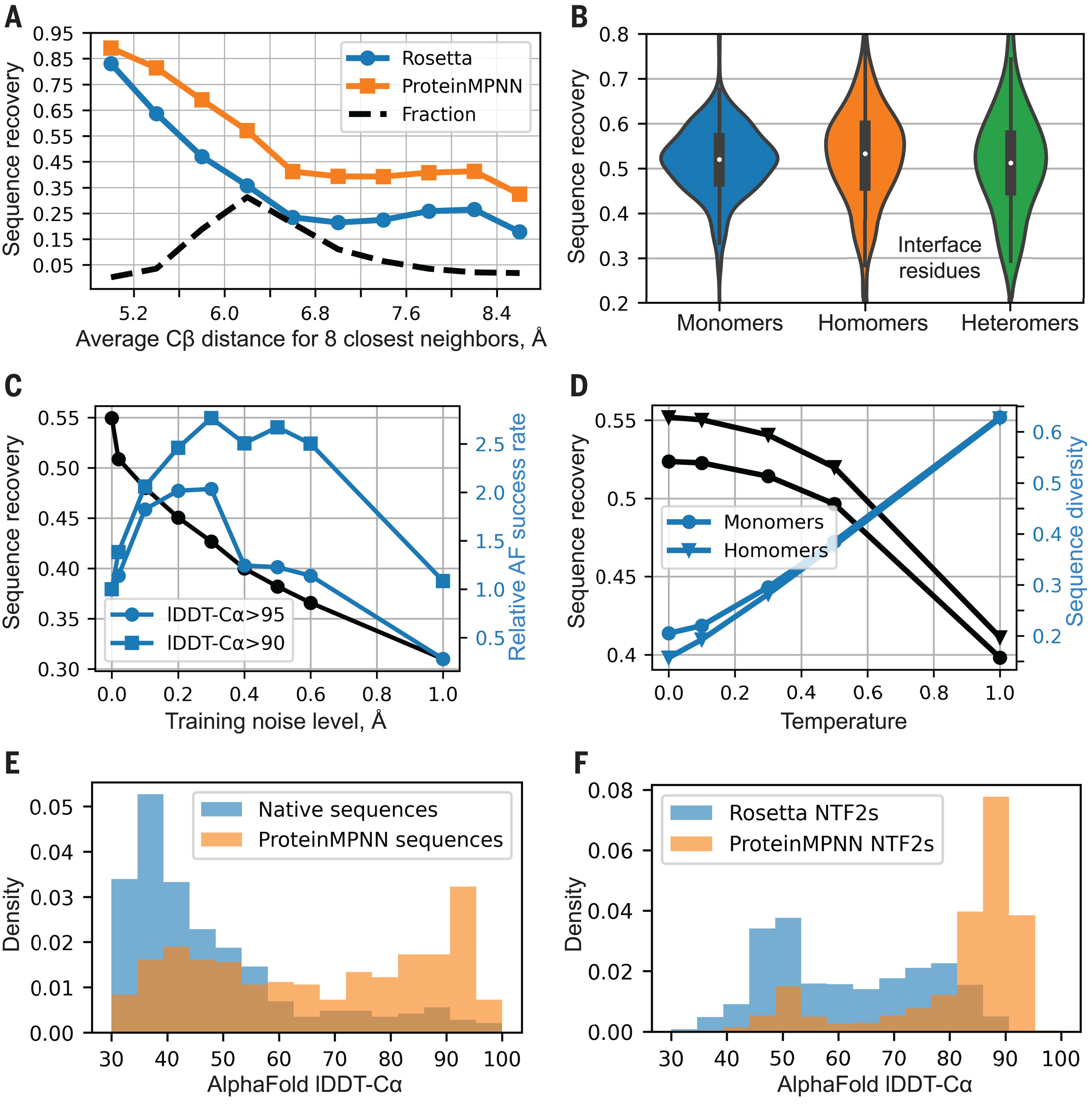
Multi-chain benchmarks demonstrated consistent performance: monomers (52%), homo-oligomers (55%), and hetero-oligomers (51%) all achieved similar recovery across test sets of 690, 732, and 98 structures respectively. The slight improvement for homo-oligomers likely reflects the tied decoding mechanism capturing symmetric constraints. Perhaps most striking was the computation time: 1.2 seconds for a 100-residue protein versus 258.8 seconds (4.3 minutes) for Rosetta—a speedup enabling the evaluation of orders of magnitude more designs.
Beyond static benchmarks, functional validation proved transformative. When researchers had generated novel protein folds through AlphaFold hallucination, the original hallucinated sequences expressed poorly with median soluble yield of just 9 mg/L. Redesigning these backbones with ProteinMPNN boosted median yield to 247 mg/L—a 27-fold improvement—with 73 of 96 designs expressing solubly and 50 achieving correct oligomeric states. For cyclic homo-oligomers, Rosetta sequences achieved 40% soluble expression with 0% correct oligomeric state; ProteinMPNN achieved 88% soluble expression with 27.7% correct assembly.
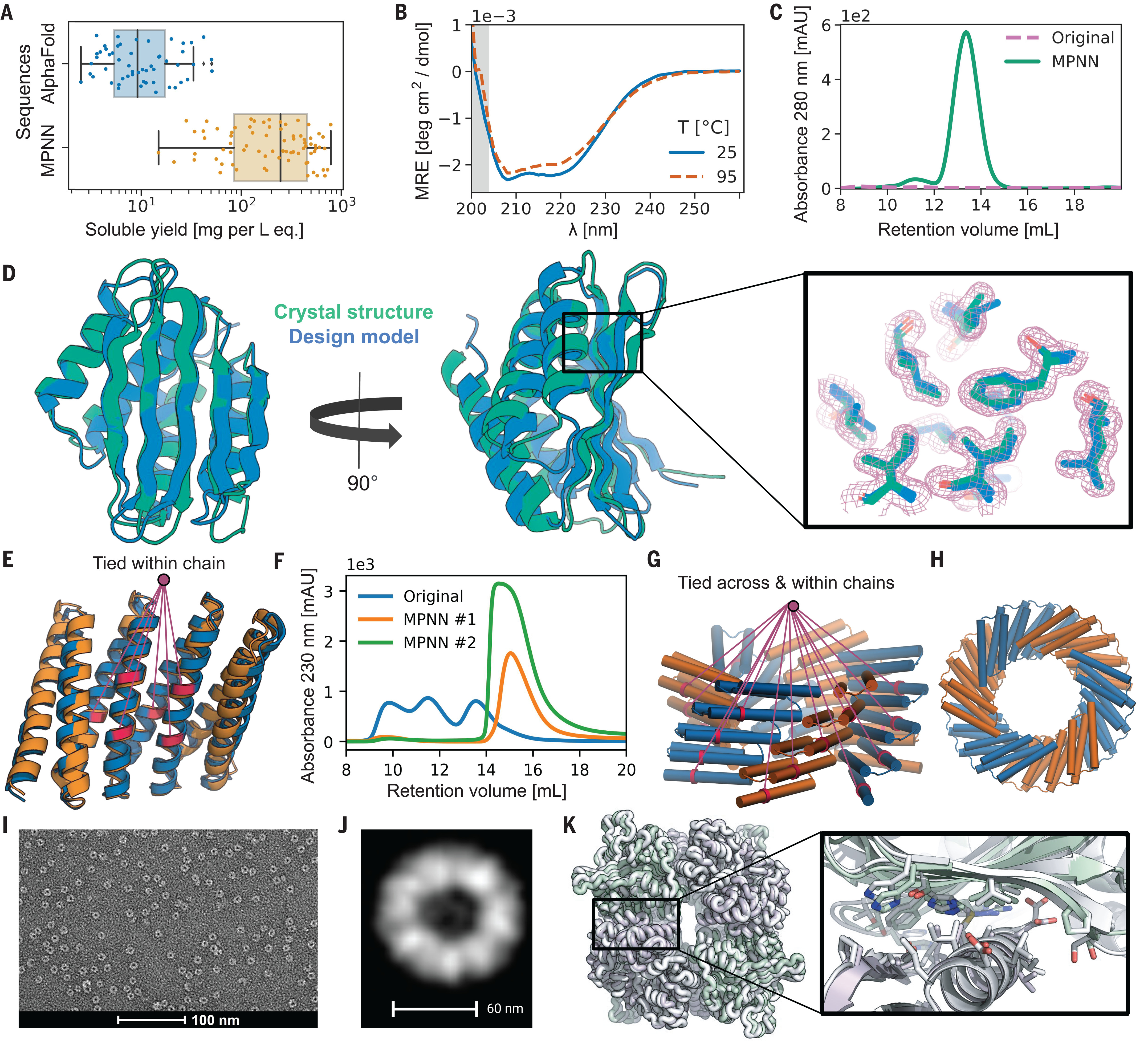
Real-world applications span therapeutics, vaccines, and enzymes
ProteinMPNN has enabled breakthrough applications across protein engineering domains. In vaccine development, tetrahedral nanoparticles designed using ProteinMPNN scaffolds display antigens with precise geometry—two-component nanoparticle vaccines are now in clinical trials for influenza (NCT04896086) and a related icosahedral design (I53-50) underpins SK Bioscience’s licensed SARS-CoV-2 vaccine. The method proved essential for rescuing designs that had failed with Rosetta: a recent PNAS study reported 13 new experimentally confirmed megadalton-scale tetrahedral assemblies designed with “orders of magnitude less computation and no manual refinement.”
For therapeutic binder design, combining RFdiffusion backbone generation with ProteinMPNN sequence design and AlphaFold validation increased success rates nearly 10-fold over previous methods. The RFantibody pipeline has produced de novo designed VHH nanobodies against RSV (1.4 μM affinity), influenza hemagglutinin (78 nM), and SARS-CoV-2 RBD (5.5 μM), with cryo-EM structures confirming designs match predictions to within 0.9-1.45 Å RMSD. For existing therapeutic antibodies, inverse folding-guided optimization achieved up to 26-fold improvement in neutralization and 37-fold improvement in affinity against antibody-escaped SARS-CoV-2 variants.
Enzyme engineering has similarly benefited. Studies combining ProteinMPNN with evolutionary constraints demonstrated improved TEV protease expression (from 1 to 20 mg/L average), elevated melting temperatures, and enhanced catalytic activity. The ability to rapidly generate diverse sequences for experimental testing—rather than spending days on Rosetta calculations—has democratized enzyme optimization workflows.
What types of protein structures can ProteinMPNN design sequences for?
ProteinMPNN is designed to handle a diverse range of 3D protein backbones, from simple monomeric proteins to complex multi-component assemblies. It is primarily a fixed-backbone model, meaning it generates amino acid sequences for a specified set of 3D coordinates provided in a PDB file.
- Monomeric Proteins: Designing sequences for single-chain proteins, including those with entirely novel ("de novo") folds.
- Protein Complexes and Multimers: It can design sequences for assemblies of multiple polypeptide chains, such as cyclic homo-oligomers or heterodimers.
- Symmetrical Assemblies: The model supports symmetry tying, allowing for the design of large, highly symmetrical structures like tetrahedral nanoparticles and other protein nanomaterials.
- Protein-Protein Interfaces: It is widely used to design the interfaces between different protein components to facilitate binding or self-assembly.
- Stability Redesign: Generating stabilizing mutations for existing native proteins to improve their expression or thermal stability.
- Specialized Environmental Variants:
- SolubleMPNN: Fine-tuned specifically for designing soluble proteins.
- Membrane Variants: Supports design with labels for membrane contexts (e.g., transmembrane proteins).
Note on Non-Protein Context: Standard ProteinMPNN only considers protein backbone coordinates; it ignores non-protein atoms like small molecules, DNA, or metal ions. For structures involving these components, the specialized LigandMPNN variant is used.
Limitations reveal where ProteinMPNN struggles
Despite its success, ProteinMPNN has documented limitations. β-sheet rich proteins, including β-barrels and jellyroll folds, show the lowest experimental success rates at approximately 21% compared to 88% for curved β-sheet designs. This stems partly from backbone quality requirements—ProteinMPNN is sensitive to angstrom-level structural accuracy for these topologies—and partly from the intrinsic aggregation tendency of extended β-strands.
Ligand and cofactor blindness represents a fundamental architectural constraint. ProteinMPNN considers only backbone coordinates, ignoring small molecules, metals, and nucleic acids entirely. Performance around ligand-binding sites drops substantially: sequence recovery near small molecules falls to 50.5%, near nucleotides to 34.0%, and near metals to just 40.6%. This limitation necessitated LigandMPNN’s development and means vanilla ProteinMPNN should not be used for enzyme active site or metal-binding protein design.
Solubility biases arise from training data composition. Because training included membrane proteins, ProteinMPNN sometimes places surface hydrophobic residues on soluble designs with membrane-like topologies, causing complete insolubility. The soluble MPNN variant, trained exclusively on soluble proteins, addresses this but requires users to select appropriate models. More broadly, approximately 65% of de novo design failures trace to aggregation or insolubility issues that sequence metrics alone cannot predict.
Sequence quality issues occasionally manifest as low diversity outputs, particularly from imperfect RFdiffusion backbones. Users report excessive glycine and lysine content, repetitive stretches, and very high isoelectric points. These symptoms typically indicate backbone quality problems rather than ProteinMPNN limitations, but distinguishing causes requires experience. Cysteine placement also proves problematic since disulfide bonds aren’t explicitly modeled—biasing against cysteines is common practice.
Comparison with alternative inverse folding methods
ESM-IF1, trained on 12 million AlphaFold-predicted structures, achieves comparable 51% recovery but offers advantages for multi-state design through its ability to condition on multiple conformations simultaneously. Its GVP-GNN encoder captures rotation-equivariant features that generalize better for partially masked backbones, though performance degrades for long masked spans. For antibody CDR design specifically, domain-specialized models like AntiFold outperform both ProteinMPNN and ESM-IF with higher sequence recovery and better binding affinity prediction.
Physics-based Rosetta fixed-backbone design remains valuable when interpretability matters or when designing around specific chemical interactions that benefit from explicit energy function terms. However, the 200-fold speed disadvantage limits throughput, and lower recovery rates reduce overall design campaign success. The emerging consensus reserves Rosetta for refinement stages or specialized chemical contexts while using neural methods for initial sequence generation.
LigandMPNN extends the architecture for atomic context
LigandMPNN, published in Nature Methods in 2025, represents the most significant ProteinMPNN extension. It introduces a three-graph architecture: the original protein backbone graph, an intra-ligand graph with atoms as nodes encoding element types and interatomic distances, and a protein-ligand graph connecting residues to nearby ligand atoms. Two additional message-passing layers process ligand information before the decoder, expanding model size to 2.62 million parameters while maintaining sub-second inference.
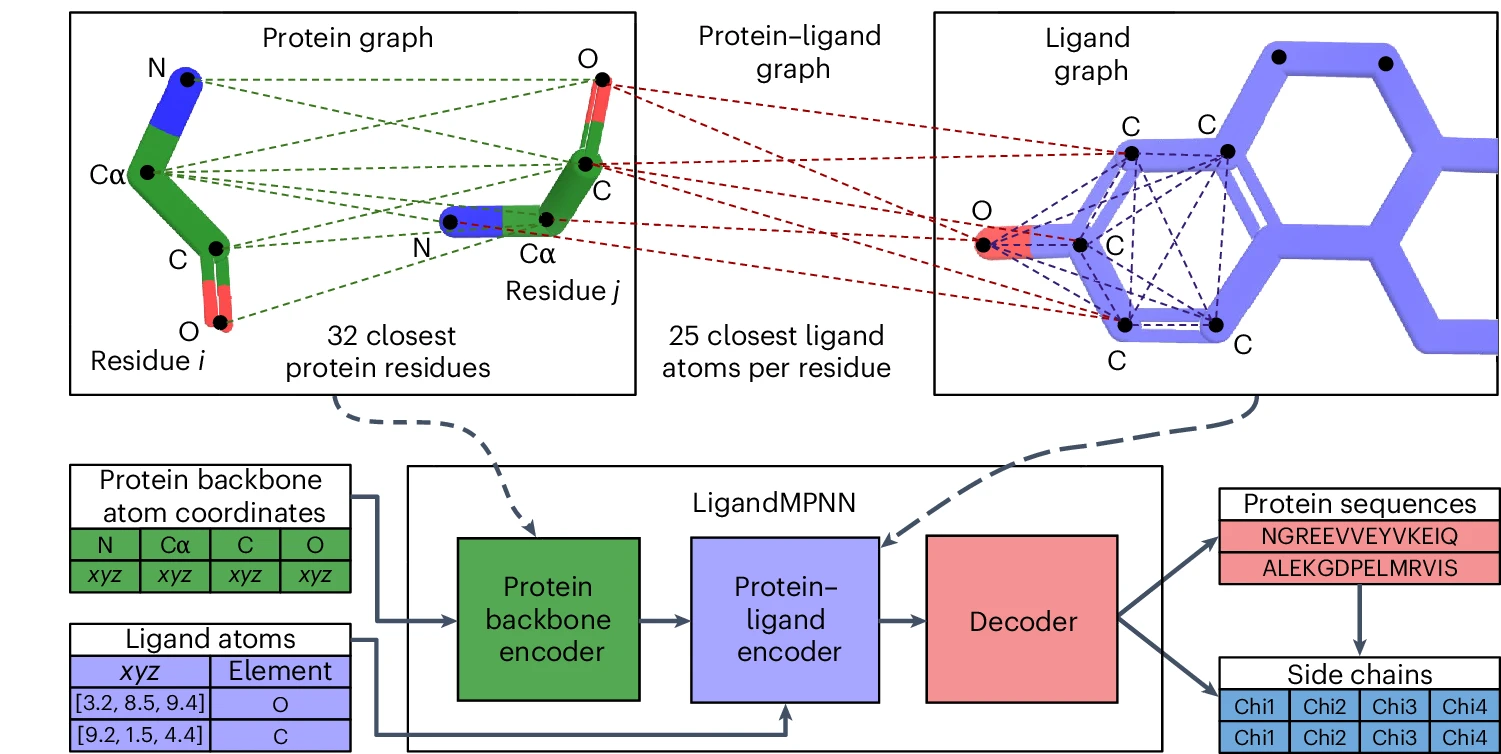
Performance improvements near ligands are dramatic. Sequence recovery around small molecules jumps from 50.5% to 63.3%, around nucleotides from 34.0% to 50.5%, and around metals from 40.6% to 77.5%—nearly doubling accuracy for metal-binding sites. LigandMPNN additionally predicts sidechain torsion angles alongside sequences, enabling evaluation of designed binding poses. Over 100 experimentally validated small-molecule and DNA-binding proteins demonstrate real-world utility, with binding affinity improvements exceeding 100-fold in rescue experiments.
Additional variants address specific needs. SolubleMPNN trains exclusively on soluble proteins, eliminating membrane protein biases for designs with membrane-like topologies. ThermoMPNN uses transfer learning from ProteinMPNN features to predict stability changes (ΔΔG) for point mutations without requiring multiple sequence alignments. DynamicMPNN (2025) explicitly models conformational ensembles, achieving up to 25% improvement on multi-state design problems by training on paired conformational states covering 75% of CATH superfamilies.
Integration with RFdiffusion defines modern design pipelines
The contemporary protein design workflow integrates three complementary tools: RFdiffusion generates backbone structures through diffusion-based denoising, ProteinMPNN designs sequences for those backbones, and AlphaFold2 validates that designed sequences are predicted to fold as intended. This pipeline, established by Watson et al. in Nature 2023, typically generates 10,000+ backbone candidates, designs 2-8 sequences per backbone with ProteinMPNN, then filters by self-consistency metrics (predicted structure within 2 Å RMSD of design model, pLDDT >70).
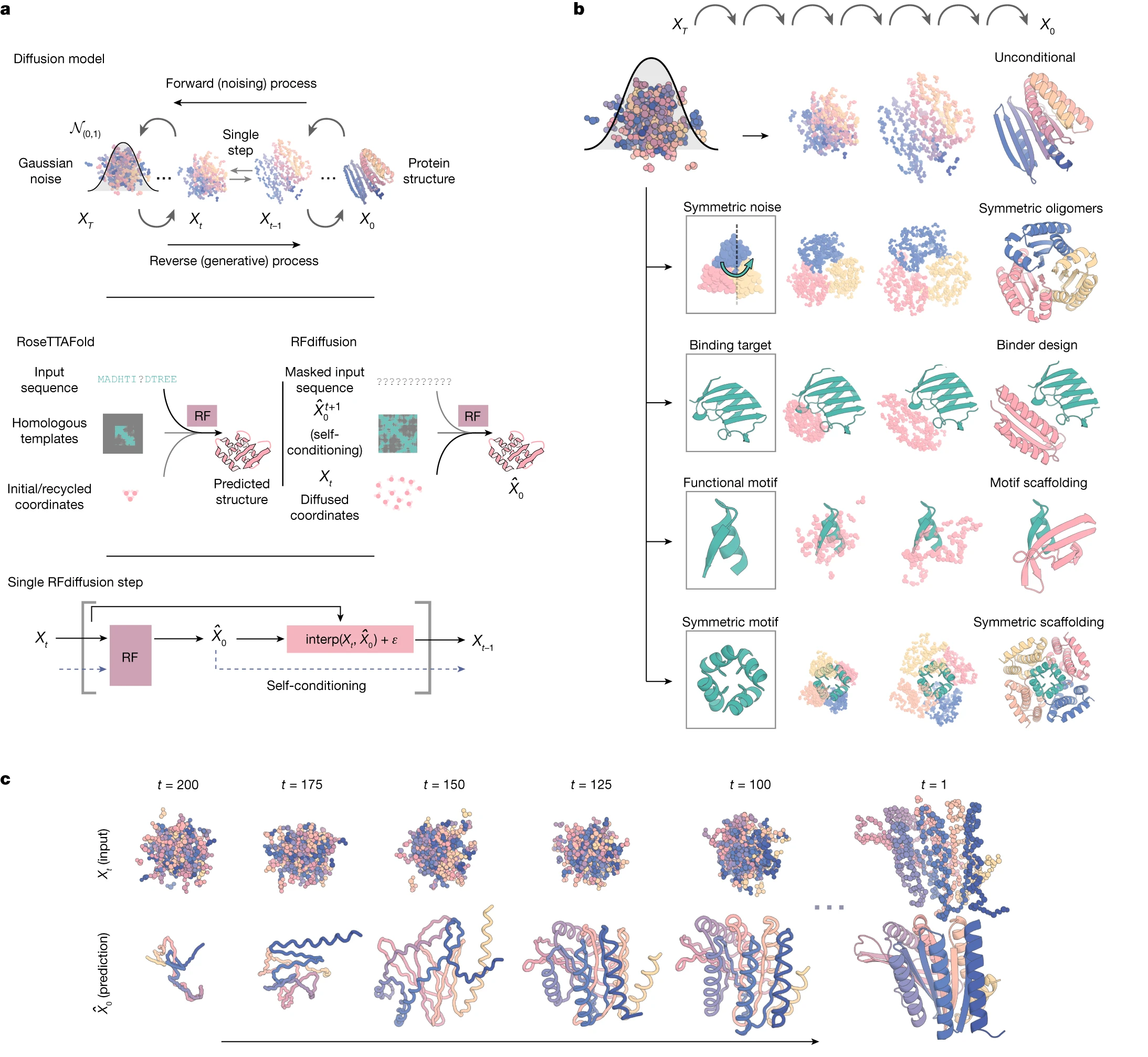
This workflow has produced remarkable results: picomolar-affinity binders, symmetric oligomers spanning C3 to octahedral symmetries, functional enzyme active sites, and de novo antibodies. The RFdiffusion All-Atom extension partners with LigandMPNN for ligand-aware design, enabling small-molecule binding site creation. Alternative backbone generators like Chroma similarly feed into ProteinMPNN-based sequence design.
For complex or large designs, iterative refinement approaches like AF2cycler alternate between AlphaFold prediction and ProteinMPNN redesign over 10-20 cycles, progressively improving both structure and sequence. This reflects the field’s understanding that single-shot design rarely optimizes all properties—iteration remains essential.
The future of inverse folding continues to evolve
ProteinMPNN established that neural networks could dramatically outperform physics-based methods for sequence design while being orders of magnitude faster. Its descendants—LigandMPNN, SolubleMPNN, ThermoMPNN, DynamicMPNN—address specific limitations while maintaining the core architectural innovations. The field continues advancing: Frame2seq claims 2% higher recovery with 6x faster inference through structure-conditioned masked language modeling, while CarbonDesign adapts AlphaFold2’s Evoformer for design.
Yet challenges remain. The gap between computational predictions and experimental outcomes persists—high recovery or pLDDT scores don’t guarantee expression, solubility, or function. Multi-state design for allosteric mechanisms or conformational switches requires explicit ensemble modeling that most methods lack. Predicting which designs will succeed experimentally before costly synthesis remains an unsolved problem.
For practitioners, ProteinMPNN (via the unified LigandMPNN repository at github.com/dauparas/LigandMPNN) remains the default starting point. Using the 0.10-0.20 Å noise models, generating multiple sequences per backbone, and filtering through AlphaFold validation provides robust workflows. Selecting appropriate variants—LigandMPNN for binding sites, SolubleMPNN for membrane-like topologies—addresses known failure modes. The method has matured from research prototype to production tool, enabling protein designs that would have been impossible just years ago.
Conclusion
ProteinMPNN transformed protein sequence design from a computational bottleneck requiring expert intervention into a rapid, reliable step enabling large-scale design campaigns. Its core innovations—distance-based backbone encoding, dual node-edge updating, random-order autoregressive decoding, and coordinate noise augmentation—combined to achieve 59% improvement over Rosetta while running 200 times faster. Real-world impact spans therapeutic binders, vaccine nanoparticles, and stabilized enzymes, with experimental success rates far exceeding previous methods.
The limitations that prompted LigandMPNN, SolubleMPNN, and other variants reveal that no single model handles all design challenges. Ligand awareness, solubility optimization, multi-state design, and stability prediction each require specialized approaches. The integration of ProteinMPNN with backbone generators like RFdiffusion and validators like AlphaFold2 has created a powerful ecosystem where each component contributes essential capabilities.
Looking forward, the inverse folding problem exemplifies how machine learning can complement rather than replace physical understanding. ProteinMPNN doesn’t simulate protein physics—it learns sequence-structure relationships from evolutionary data embedded in the PDB. This pragmatic approach, focused on prediction accuracy rather than mechanistic insight, proved transformatively effective. For computational biologists designing proteins today, ProteinMPNN and its descendants are indispensable tools that have fundamentally changed what’s achievable in protein engineering.
References
-
Ingraham et al., Generative models for graph-based protein design, NeurIPS 2019
-
Hsu et al., Learning inverse folding from millions of predicted structures, (Preprint) BioRxiv 2022
-
Dauparas et al., Robust deep learning–based protein sequence design using ProteinMPNN, Science 2022
-
Dauparas et al., Atomic context-conditioned protein sequence design using LigandMPNN, Nature Methods 2025
-
Abrudan et al., Multi-state Protein Design with DynamicMPNN, (Preprint) Arxiv 2025
-
Høie et al., AntiFold: Improved antibody structure-based design using inverse folding (Preprint) Arxiv 2024.
-
Diechaus et al., Transfer learning to leverage larger datasets for improved prediction of protein stability changes
-
Watson et al., De novo design of protein structure and function with RFdiffusion, Nature 2023.
Enjoy Reading This Article?
Here are some more articles you might like to read next: