RFDiffusion Revolution
From Backbone to Atomic Precision
Diffusion models transformed protein design in 2023—now RFDiffusion3 generates complete proteins atom by atom, including the first computationally designed DNA-binding proteins. The Baker Lab’s RFDiffusion family represents the most experimentally validated generative approach to protein design, with success rates two orders of magnitude higher than previous methods. What began as backbone generation has evolved into a unified foundation model capable of designing proteins that bind DNA, small molecules, and other proteins with atomic precision. This technical deep-dive traces the architectural evolution from RFDiffusion v1 through All-Atom to RFDiffusion3, explaining the innovations that made each leap possible.
The core insight that changed protein design
Before RFDiffusion, computational protein design relied on either physics-based methods (Rosetta) or “hallucination” approaches that iteratively optimized sequences to maximize neural network confidence. Both had fundamental limitations: Rosetta required extensive sampling with low success rates, while hallucination was computationally expensive and struggled beyond ~100 residues.
RFDiffusion’s key innovation was recognizing that structure prediction networks already encode deep knowledge of protein physics. Rather than building diffusion models from scratch, Watson et al. fine-tuned RoseTTAFold—a pre-trained structure prediction network—to serve as the denoising function. This transfer learning approach proved critical: training from scratch completely fails, but fine-tuning converges in just 5 epochs (~3 days on 8 A100 GPUs).
The mathematical framework follows denoising diffusion probabilistic models (DDPMs). Given a protein structure $X_0$, the forward process progressively corrupts it:
\[q(X_t | X_{t-1}) = \mathcal{N}(X_t; \sqrt{1-\beta_t}X_{t-1}, \beta_t\mathbf{I})\]The model learns to reverse this corruption, predicting the clean structure $\hat X_0$ from any noised state $X_t$. Unlike standard DDPMs that predict noise $ε$, RFDiffusion directly predicts the final denoised structure—a choice that improves generation quality and enables powerful self-conditioning.
RFDiffusion v1: residue-level generation with pretrained foundations
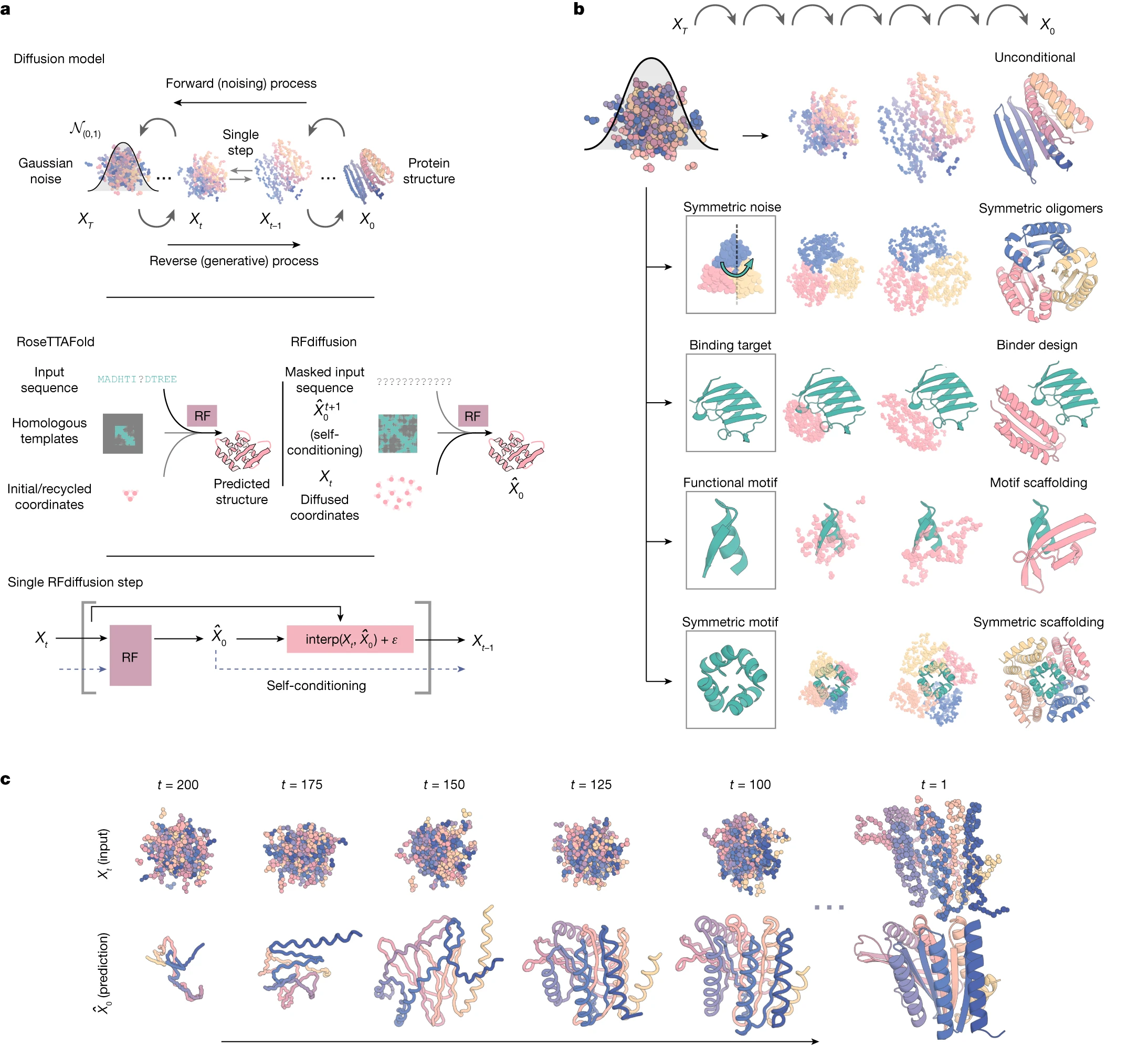
The original RFDiffusion (Watson et al., Nature 2023) operates at the residue level, representing each amino acid as a frame in SE(3)—the special Euclidean group capturing 3D rotations and translations. Each frame consists of:
- Translation $z$: $\mathrm{Cα}$ coordinate position in ℝ³
- Rotation $r$: 3×3 rotation matrix encoding N-$\mathrm{Cα}$-C backbone orientation
This frame representation elegantly captures backbone geometry while respecting the physical nature of protein structure. The diffusion process treats translations and rotations differently: translations receive standard 3D Gaussian noise, while rotations undergo Brownian motion on the SO(3) manifold using the isotropic Gaussian distribution (IGSO3).
The RoseTTAFold2 backbone provides a three-track architecture that simultaneously processes 1D sequence information, 2D pairwise features, and 3D coordinates through SE(3)-equivariant attention layers. For RFDiffusion, minimal modifications convert this predictor into a generator: the primary input changes from sequence to diffused residue frames, and template confidence features encode the denoising timestep instead.
1. Training strategy and self-conditioning
Training uses structures from the Protein Data Bank with a carefully balanced sampling ratio of 2:1:4:1 across monomers, homo-oligomers, hetero-oligomers, and AlphaFold2 models. The loss function uses mean squared error without alignment—a deliberate choice over the frame-aligned point error (FAPE) used in AlphaFold2. MSE promotes global coordinate continuity across timesteps, critical for generating coherent structures.
Self-conditioning, inspired by AlphaFold2’s recycling mechanism, dramatically improves performance. During training, the model alternates between receiving no prior prediction ($X_0 = 0$) and receiving its own previous prediction as additional input. At inference, this creates a feedback loop where each denoising step benefits from the model’s current best estimate of the final structure.
2. Capabilities that established the paradigm
RFDiffusion v1 demonstrated remarkable breadth:
- Unconditional generation: Produces diverse α, β, and mixed topologies up to 600 residues in 11 seconds (vs. 8.5 minutes for hallucination)
- Motif scaffolding: Holds functional motifs fixed while generating surrounding scaffold—solved 23/25 benchmark problems
- Binder design: Achieved ~19% experimental success rate, including a 28 nM influenza hemagglutinin binder validated by cryo-EM at 2.9 Å resolution
- Symmetric assemblies: Cyclic, dihedral, and icosahedral symmetries through explicit symmetrization at each denoising step
RFDiffusion All-Atom: conditioning on the molecular world
The original RFDiffusion had a critical limitation: it could not explicitly model small molecules, metals, or side-chain interactions. Binding pocket generation relied on heuristic contact potentials rather than actual molecular interactions. RFDiffusion All-Atom (Krishna et al., Science 2024) addressed this by building on RoseTTAFold All-Atom (RFAA)—a unified architecture for modeling proteins alongside arbitrary small molecules.
1. Dual representation architecture
RFAA introduces a hybrid representation system:
- Residue-based track: Extended from 20 amino acids to 28 tokens (adding 4 DNA and 4 RNA bases)
- Atom-bond graph track: Represents arbitrary small molecules as graphs with node features for atom types and edge features for bond types
Covalent bonds between proteins and small molecules receive special treatment through a bond adjacency matrix with dedicated tokens. This enables modeling of covalent modifications, covalent drugs, and enzyme-substrate complexes where chemistry crosses the protein-ligand boundary.
For RFDiffusionAA, the key change is that small molecule coordinates are held fixed while protein residue frames undergo diffusion. The network learns the conditional distribution of proteins given complete molecular context—not just shape complementarity, but actual atomic interactions.
2. Experimental validation of small molecule binders
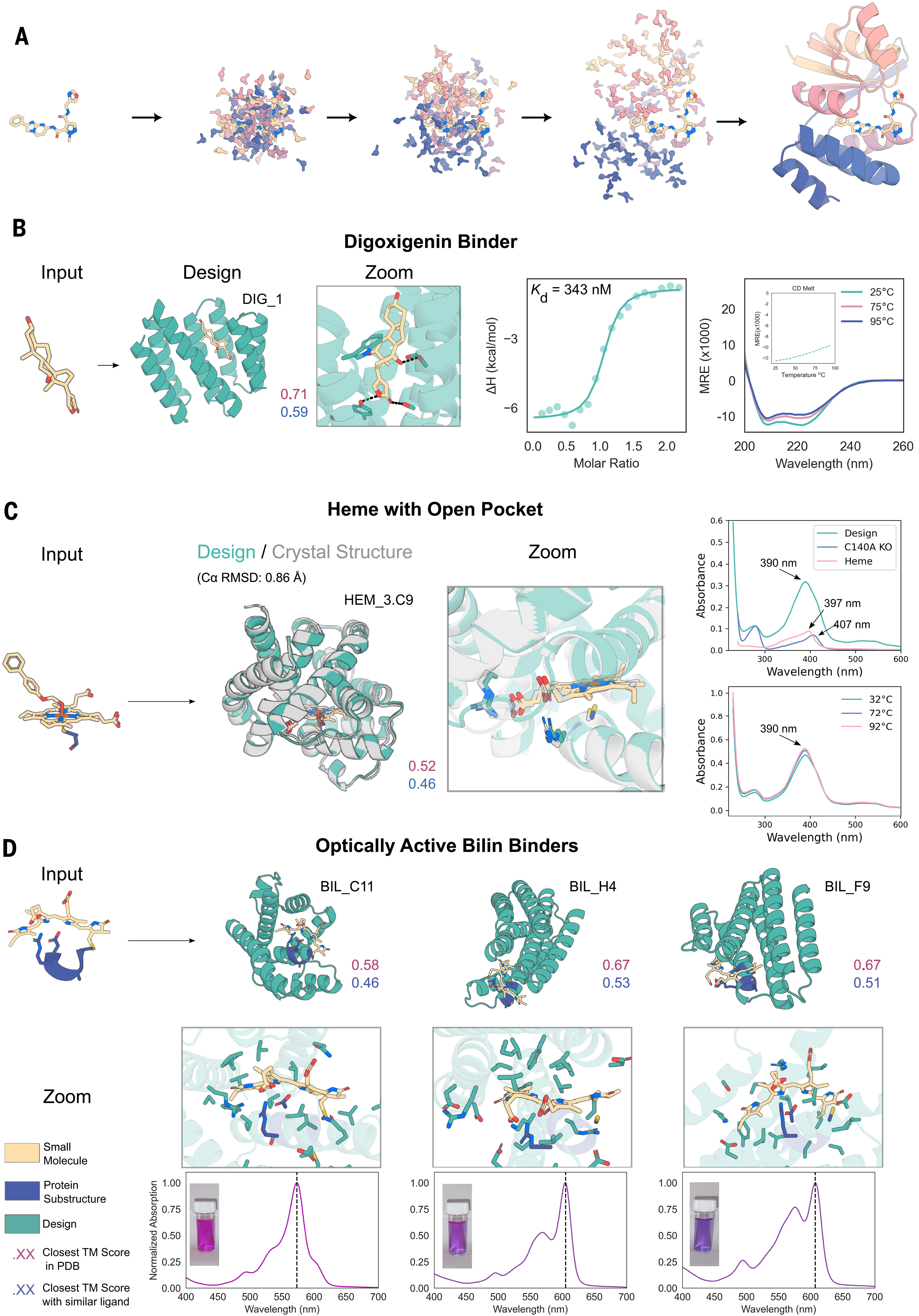
The approach produced striking results across three challenging targets:
| Target | Binding Affinity | Structural Validation | Novelty (TM to PDB) |
|---|---|---|---|
| Digoxigenin | Kd = 343 nM | CD, thermal stability | 0.59 |
| Heme | Tight binding | Crystal structure (0.86 Å RMSD) | <0.62 |
| Bilin | Fluorescence-active | CD, fluorescence | <0.52 |
The heme binder crystal structure deserves emphasis: 0.86 Å Cα RMSD between design and experiment represents near-perfect atomic accuracy for a de novo protein binding a complex cofactor. These designs show no sequence similarity to natural proteins, demonstrating genuine generalization beyond training examples.
RFDiffusion3: true all-atom co-diffusion
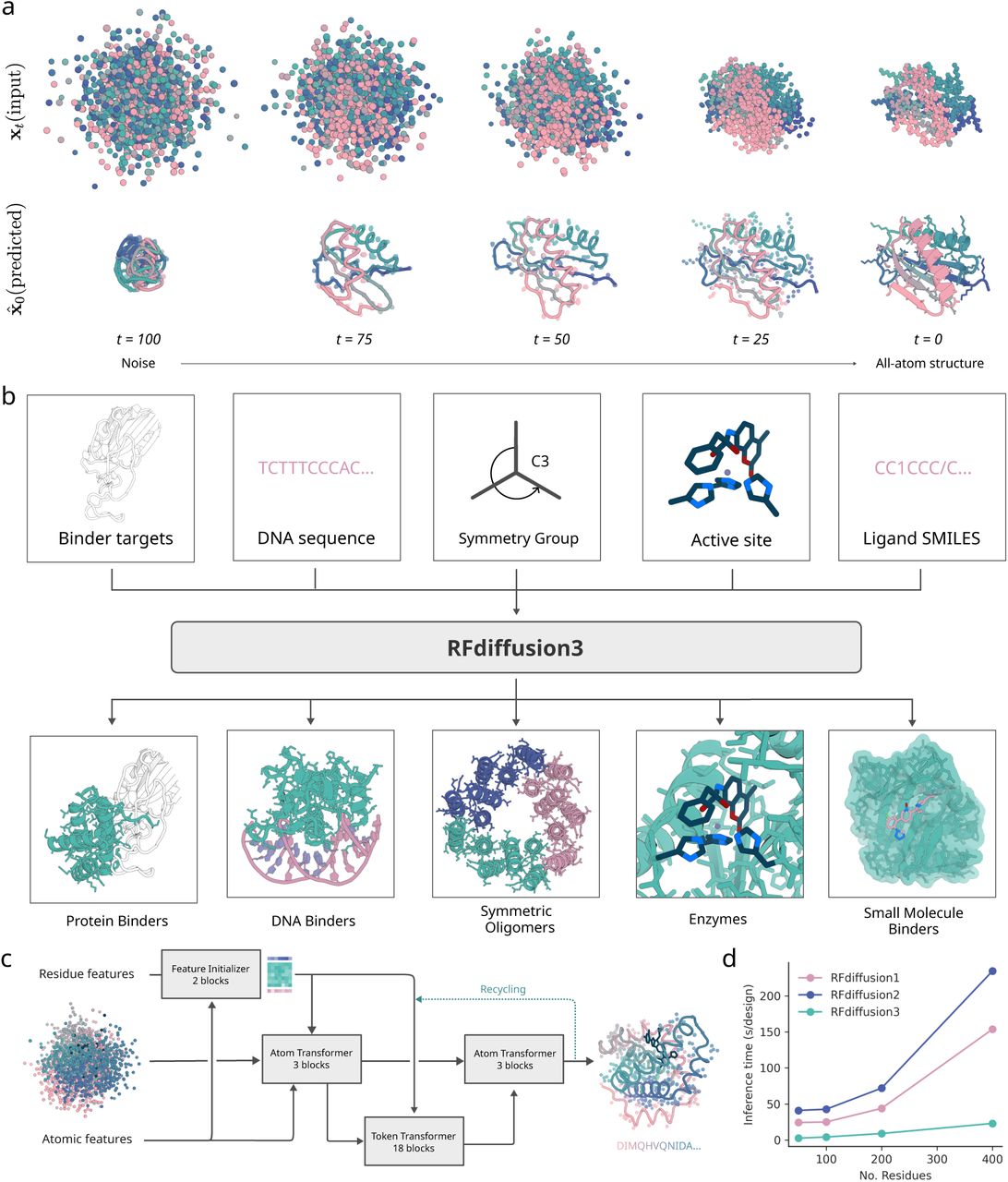
Released in December 2025, RFDiffusion3 represents the most ambitious leap yet. It abandons the residue-frame representation entirely, instead diffusing all atoms simultaneously—backbone, side chains, and binding partners together. The result is a unified foundation model achieving capabilities previously impossible.
1. Architectural transformation
RFDiffusion3 shares no code with its predecessors. Where previous versions adapted structure prediction networks, RFD3 inverts AlphaFold3’s architecture into a generative system:
- Transformer-based U-Net: Replaces AF3’s sequence-processing trunk (unnecessary for design)
- Sparse attention: Focuses computation on geometrically adjacent atoms rather than all-to-all attention
- Cross-attention pooling: Maintains coherence between sidechain and backbone representations
- ~168 million parameters: Approximately half the size of AlphaFold3 while achieving comparable accuracy
Each residue receives a uniform 14-atom representation (4 backbone + up to 10 sidechain), enabling the model to “see” and generate complete atomic detail. The diffusion process simultaneously refines protein and binding partner conformations through co-diffusion—both components adapt to each other throughout generation rather than one being fixed.
2. Atom-level conditioning enables unprecedented control
RFD3 introduces conditioning mechanisms unavailable to previous versions:
- Hydrogen bond specification: Define specific atoms as donors/acceptors
- Solvent accessibility (RASA): Control whether ligands are buried or surface-exposed
- Center of mass: Position the protein relative to targets
- Symmetry operations: Generate oligomers from initial noise
- Catalytic motifs: Place precise active site geometries
This fine-grained control proved essential for the breakthrough capability: DNA-binding protein design. Previous attempts required manually specifying every atomic position—RFD3 achieves ~9% success rate on arbitrary DNA sequences through learned hydrogen bonding patterns. One design achieved EC50 ≈ 5.9 μM against a randomly generated DNA sequence, opening the door to synthetic transcription factors.
3. Performance benchmarks demonstrate systematic improvements
| Task | RFD3 Performance | Comparison |
|---|---|---|
| Enzyme active sites | 90% success rate | 37/41 wins vs. RFD2 on complex sites |
| Protein binders | 4/5 wins vs. RFD1 | More diverse geometries |
| DNA binders | ~9% success | Previously near-impossible |
| Speed | 10× faster than RFD2 | Despite full all-atom modeling |
Enzyme design showcases the practical impact: 190 cysteine hydrolase designs yielded 35 active catalysts (~18% hit rate), with the best achieving kcat/Km = 3,557 M⁻¹s⁻¹—comparable to natural enzymes.
Technical innovations across the evolution
1. Frame representations and SE(3) equivariance
All versions leverage SE(3) equivariance—the property that outputs transform correctly under 3D rotations and translations. This isn’t merely computational convenience; it ensures physically realistic outputs regardless of arbitrary coordinate system choices. RoseTTAFold implements this through Invariant Point Attention (IPA), which operates on local reference frames while maintaining global consistency.
2. The critical role of pretraining
A consistent theme across versions: starting from pretrained structure prediction weights is essential. The billions of parameters in these networks encode evolutionary constraints, secondary structure preferences, and packing principles that would be impossible to learn from diffusion objectives alone. This transfer learning approach reduces training time from weeks to days while dramatically improving generation quality.
3. Loss function choices matter
RFDiffusion’s use of MSE loss (without alignment) instead of FAPE deserves attention. FAPE—the loss used in AlphaFold2—computes error in local frames and is invariant to global transformations. While ideal for structure prediction, it disrupts the global coordinate continuity needed for coherent diffusion trajectories. MSE anchors each prediction in a consistent coordinate system.
4. From backbone imagination to atomic reality
The evolution from v1 → All-Atom → v3 reflects increasing atomic fidelity:
| Version | Resolution | Sidechain Treatment | Ligand Treatment |
|---|---|---|---|
| v1 | Backbone frames | Implicit (ProteinMPNN adds later) | Contact potentials only |
| All-Atom | Backbone + ligand atoms | Still implicit | Explicit conditioning |
| v3 | Full all-atom | Explicit co-diffusion | Full co-diffusion |
This progression matters because protein function depends on atomic detail—enzyme catalysis, molecular recognition, and allosteric regulation all occur at the atomic level.
Applications transforming biology and medicine
1. Therapeutic protein design
RFDiffusion-designed proteins have reached clinical development:
- SKYCovione: First computationally designed protein vaccine (COVID-19), approved in South Korea
- Picomolar SARS-CoV-2 inhibitors: Miniproteins blocking viral entry
- MDM2 binders: 0.5-0.7 nM affinity, 1000× stronger than native p53 peptide
- pLYTACs and EndoTags: Targeted protein degradation and receptor modulation
The success rate improvement is quantifiable: RFDiffusion binder design achieves ~19% experimental success at 10 μM screening, compared to <0.1% for previous Rosetta-based approaches.
2. Enzyme engineering
RFDiffusion2 and RFDiffusion3 have advanced enzyme design substantially. Active site scaffolding—placing catalytic residues in precise geometries within stable protein scaffolds—previously required extensive manual intervention. Automated approaches now achieve:
- Metallohydrolases: kcat/Km up to 53,000 M⁻¹s⁻¹
- Cysteine hydrolases: ~18% hit rate for active designs
- Complex multi-residue sites: RFD3 outperforms on disconnected catalytic residues
3. Comparison with alternative approaches
| Method | Approach | Key Strength | Experimental Validation |
|---|---|---|---|
| RFDiffusion | Pretrained RF + DDPM | Broad applications, highest success rates | Extensive (hundreds of proteins) |
| Chroma | Custom GNN + diffusion | Text prompts, scalability | 310 proteins characterized |
| Genie 2 | Asymmetric diffusion | Multi-motif scaffolding | Limited |
| FrameFlow | SE(3) flow matching | No pretraining needed | Moderate |
RFDiffusion’s advantage lies in experimental validation depth. The standard workflow—RFDiffusion → ProteinMPNN → AlphaFold2 validation—has produced more structurally confirmed designs than any competing approach.
Looking forward: toward generalist protein models
RFDiffusion3’s release alongside RF3 (structure prediction), RFantibody (antibody design), and a unified Foundry platform signals a new phase: consolidated foundation models replacing task-specific tools. Jasper Butcher, RFD3’s lead developer, describes the goal as “generalist protein-design models capable of continuous iteration”—systems that design, validate, and refine proteins in automated loops.
Current limitations point toward future development:
- Post-translational modifications (glycosylation, phosphorylation) remain unsupported
- Non-canonical amino acids fall outside training distributions
- Correlation between in silico metrics and experimental success needs improvement
- Success rates for highly efficient enzymes remain ~1%, requiring extensive screening
The 2024 Nobel Prize to David Baker, alongside DeepMind’s Demis Hassabis and John Jumper, recognized how fundamentally these methods have transformed structural biology. RFDiffusion exemplifies this transformation: what once required years of manual design iterations now completes in seconds, with success rates that make previously impossible targets tractable.
Conclusion
The RFDiffusion family’s evolution from backbone frames to true all-atom generation mirrors the field’s maturation from proof-of-concept to practical tool. Three architectural themes emerge as central to this progress:
- Transfer learning from structure prediction: Pretrained networks provide essential physical knowledge that cannot be learned from diffusion objectives alone
- Increasing atomic fidelity: Function depends on atoms, and explicit atomic modeling yields better functional designs
- Unified representations: Single models handling proteins, nucleic acids, and small molecules reduce the fragmentation that plagued earlier approaches
For computational biologists entering this field, the practical takeaway is clear: RFDiffusion represents the current state-of-the-art for experimentally validated protein design, with RFD3 extending capabilities to DNA binding and complex enzyme sites. The code is freely available (github.com/RosettaCommons/foundry), and the standard design pipeline (diffusion → inverse folding → structure prediction validation) provides a reproducible framework for generating novel proteins with functions never seen in nature.
References
-
Watson et al., “De novo design of protein structure and function with RFdiffusion”, Nature 2023. https://doi.org/10.1038/s41586-023-06415-8
-
Krishna et al., “Generalized biomolecular modeling and design with RoseTTAFold All-Atom”, Science 2024. https://doi.org/10.1126/science.adl2528
-
Butcher et al., “De novo Design of All-atom Biomolecular Interactions with RFdiffusion3”, (Preprint) BioRxiv 2025. https://doi.org/10.1101/2025.09.18.676967
Enjoy Reading This Article?
Here are some more articles you might like to read next: