Transformers Revolutionize Protein Structure Prediction and Design
Attention mechanisms have fundamentally transformed computational structural biology, enabling tools like AlphaFold, ESMFold, and RFdiffusion to achieve atomic-level accuracy in predicting and designing protein structures. This technical deep-dive examines how the transformer architecture—originally developed for natural language processing—has been adapted to solve some of biology’s most challenging problems, from predicting protein folds to designing novel therapeutic molecules.
The breakthrough began in 2020 when AlphaFold2 achieved median backbone RMSD of 0.96 Å at CASP14, essentially solving the protein folding problem for single domains. Since then, transformers have become the dominant architecture across structure prediction, inverse folding, and generative design. This post provides the mathematical foundations and architectural details that graduate students and researchers need to understand and contribute to this rapidly evolving field.
Mathematical foundations of attention mechanisms
The transformer architecture, introduced by Vaswani et al. in the seminal 2017 paper “Attention Is All You Need”, relies entirely on attention mechanisms without recurrence or convolution. Understanding these mechanisms is essential for grasping how modern protein tools work.
Scaled dot-product attention
The core attention operation computes weighted sums of value vectors, where weights depend on query-key similarities:
\[\text{Attention}(Q, K, V) = \text{softmax}\left(\frac{QK^T}{\sqrt{d_k}}\right)V\]Here, $Q \in \mathbb{R}^{n \times d_k}$ represents queries, $K \in \mathbb{R}^{n \times d_k}$ represents keys, and $V \in \mathbb{R}^{n \times d_v}$ represents values. The scaling factor $\sqrt{d_k}$ prevents dot products from growing large enough to push softmax into regions with extremely small gradients. For input embeddings $x$, these matrices are computed via learned linear projections: $Q = xW^Q$, $K = xW^K$, $V = xW^V$.
Multi-head attention enables diverse representations
Rather than performing single attention, the transformer uses multiple parallel attention heads that learn different relationship patterns:
\[\mathrm{MultiHead(Q, K, V) = Concat(head_1, ..., head_h)W^O}\]where $\mathrm{head_i = Attention(QW_i^Q, KW_i^K, VW_i^V)}$. The original transformer used 8 heads with $d_k = d_v = 64$ and $d_\text{model} = 512$. This enables the model to jointly attend to information from different representation subspaces—crucial for proteins where residue relationships involve both local structural and long-range evolutionary patterns.
Cross-attention bridges different representations
Cross-attention differs from self-attention by sourcing queries from one representation and keys/values from another:
\[\text{CrossAttention}(Q_{target}, K_{source}, V_{source}) = \text{softmax}\left(\frac{Q_{target}K_{source}^T}{\sqrt{d_k}}\right)V_{source}\]This mechanism proves essential in protein modeling for integrating multiple information tracks—for example, connecting MSA representations with pair representations in AlphaFold, or linking protein embeddings with structure coordinates in design models.
Positional encoding preserves sequence information
Since attention is permutation-invariant, positional encodings must inject sequence order. The original transformer used sinusoidal functions:
\[PE_{(pos, 2i)} = \sin\left(\frac{pos}{10000^{2i/d_{model}}}\right), \quad PE_{(pos, 2i+1)} = \cos\left(\frac{pos}{10000^{2i/d_{model}}}\right)\]For proteins, various schemes have emerged including relative positional encodings (capturing residue distances) and rotary position embeddings (RoPE) used in ESM-2, which enable better generalization to varying sequence lengths.
The evolution from CNNs to transformers in structural biology
Before transformers, protein structure prediction relied on convolutional neural networks and recurrent architectures. The shift occurred because transformers address fundamental limitations of earlier approaches.
CNNs suffer from limited receptive fields—capturing long-range residue contacts requires deep stacking, making optimization difficult. A protein’s function often depends on interactions between residues separated by hundreds of positions in sequence. RNNs and LSTMs process sequences sequentially, preventing parallelization and exhibiting exponentially decaying attention to distant positions. Protein folding, however, involves global cooperative effects where distant residues influence each other simultaneously.
Transformers solve both problems: attention provides $\text{O(1)}$ path length between any two positions (versus $\text{O(n)}$ for RNNs), and the architecture enables complete parallelization during training. Self-attention allows each residue to directly attend to all others, naturally capturing the co-evolutionary signals that encode structural constraints.
The transition happened rapidly. CASP13 (2018) saw AlphaFold using primarily CNN-based architectures. By CASP14 (2020), AlphaFold2’s transformer-based Evoformer achieved experimental-level accuracy. Today, every major structure prediction and design tool uses transformers as their core architecture.
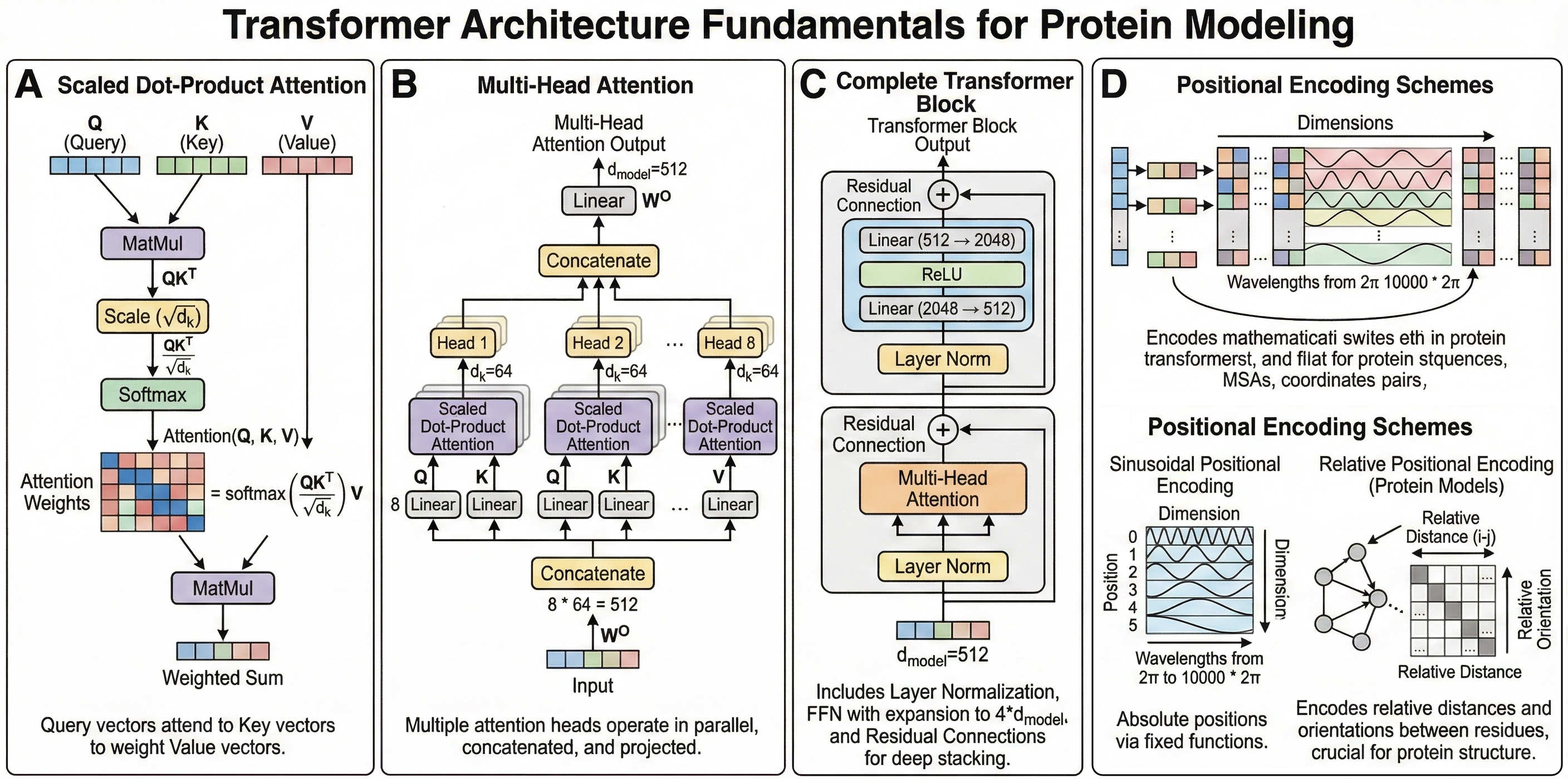
Figure 1: Transformer architecture fundamentals for protein modeling. (A) shows the scaled dot-product attention mechanism with the mathematical formulation overlaid, depicting how query vectors attend to key vectors to weight value vectors. (B) illustrates multi-head attention, showing how multiple attention heads operate in parallel and are concatenated before the output projection—include specific dimensions ($h=8$ heads, $d_k=64$) from the original transformer. C demonstrates the complete transformer block with layer normalization, feed-forward networks (showing the expansion to $d_\text{ff}=2048$), and residual connections that enable deep stacking. (D) compares sinusoidal positional encoding (wavelengths from $2\pi$ to $10000 \cdot 2\pi$) with relative positional schemes used in protein models. The figure should emphasize that the same mathematical operations underlie all protein transformers, with tool-specific innovations in how attention is applied (axial, triangular, invariant point) and what representations are processed (sequences, MSAs, coordinates, pairs).
AlphaFold2’s Evoformer revolutionizes structure prediction
AlphaFold2 (Jumper et al., Nature 2021) introduced the Evoformer, a novel transformer module that jointly processes multiple sequence alignments (MSA) and pairwise residue representations through 48 repeated blocks. This architecture achieved median GDT-TS of 92.4 at CASP14, producing structures with backbone accuracy approaching 1 Å.
MSA attention captures co-evolution
The Evoformer processes MSA representations through two complementary attention patterns. Row-wise gated self-attention applies attention independently to each sequence in the MSA, with the pair representation providing bias terms to attention logits: $\text{Attention}(Q, K, V) = \text{softmax}(QK^T/\sqrt{d} + z_{ij}) \times V$. This enables each sequence to refine its representation based on pairwise structural relationships. Column-wise attention operates across sequences at each residue position, capturing co-variation patterns that encode evolutionary constraints on structure.
This axial factorization—separating attention into row-wise and column-wise components—reduces computational complexity from $\mathrm{O(L^3)}$ to $\mathrm{O(L^2)}$ while maintaining the ability to capture both within-sequence and across-sequence relationships.
Triangle updates enforce geometric consistency
The Evoformer’s most distinctive innovation is triangle multiplicative updates for pair representations. Inspired by the triangle inequality constraint on distances, these operations update edge $ij$ using information from edges $ik$ and $kj$:
\[z_{ij} = z_{ij} + \sum_k (g_{ik} \odot \text{Linear}(z_{ik})) \odot (g_{jk} \odot \text{Linear}(z_{jk}))\]where $g$ represents learned gates. This ensures that pairwise representations maintain geometric self-consistency—if residues $i$ and $k$ are close, and $k$ and $j$ are close, this constrains the relationship between $i$ and $j$.
Invariant Point Attention for $\text{SE(3)}$-equivariant structure refinement
The Structure Module converts Evoformer outputs into 3D coordinates using Invariant Point Attention (IPA), an $\text{SE(3)}$-equivariant attention mechanism. Each residue is represented as a rigid body frame (rotation $\mathrm{R \in SO(3)}$ plus translation $t \in \mathbb{R}^3$) representing the $\mathrm{N-Cα-C}$ backbone geometry.
IPA attention weights combine three components:
\[w_{ij} = \text{softmax}\left(\frac{1}{\sqrt{c}}\left[q_i^T k_j + b_{ij} + w_c \sum_p \lVert T_i^{-1}(q_p^i) - T_j^{-1}(k_p^j) \rVert^2\right]\right)\]The third term—squared Euclidean distances between 3D query/key points projected through backbone frames—provides spatial awareness. Outputs are projected back to local frames, ensuring the entire operation is invariant to global rotations and translations. This property is essential for structure prediction, where the absolute orientation of a protein is arbitrary.
AlphaFold3 introduces diffusion-based architecture
AlphaFold3 (Abramson et al., Nature 2024) represents a fundamental architectural shift, replacing the deterministic Structure Module with a diffusion-based generative model and extending predictions to all biomolecular types.
Pairformer simplifies MSA processing
The new Pairformer architecture (48 blocks) separates MSA processing from the main trunk. A smaller MSA Module preprocesses multiple sequence alignments, extracting a single representation (first row of MSA) that feeds into the Pairformer. This simplification reduces computational overhead while maintaining accuracy, as the single representation carries sufficient evolutionary information from the language-model-like MSA processing.
Diffusion replaces deterministic structure prediction
AlphaFold3’s most significant innovation is its diffusion module, which generates structures by iteratively denoising from random atom positions. Starting from Gaussian noise (a “cloud of atoms”), the model applies ~200 denoising steps:
\[x_{update} = x_{noised} + \sigma(t) \times f_\theta(x_{noised}, z_{pair}, s_{single}, t)\]The denoising network uses a two-level atom/token architecture: local atom attention within 32-atom neighborhoods, full self-attention between token representatives, and cross-attention bridging levels. This enables efficient processing while maintaining atomic-level accuracy.
Critically, AlphaFold3 removes explicit $\text{SE(3)}$ equivariance constraints, instead learning rotational and translational invariance from data. This simplification, combined with diffusion’s generative nature, enables prediction of all biomolecular types—proteins, DNA, RNA, small molecule ligands, ions, and covalent modifications—within a unified framework.
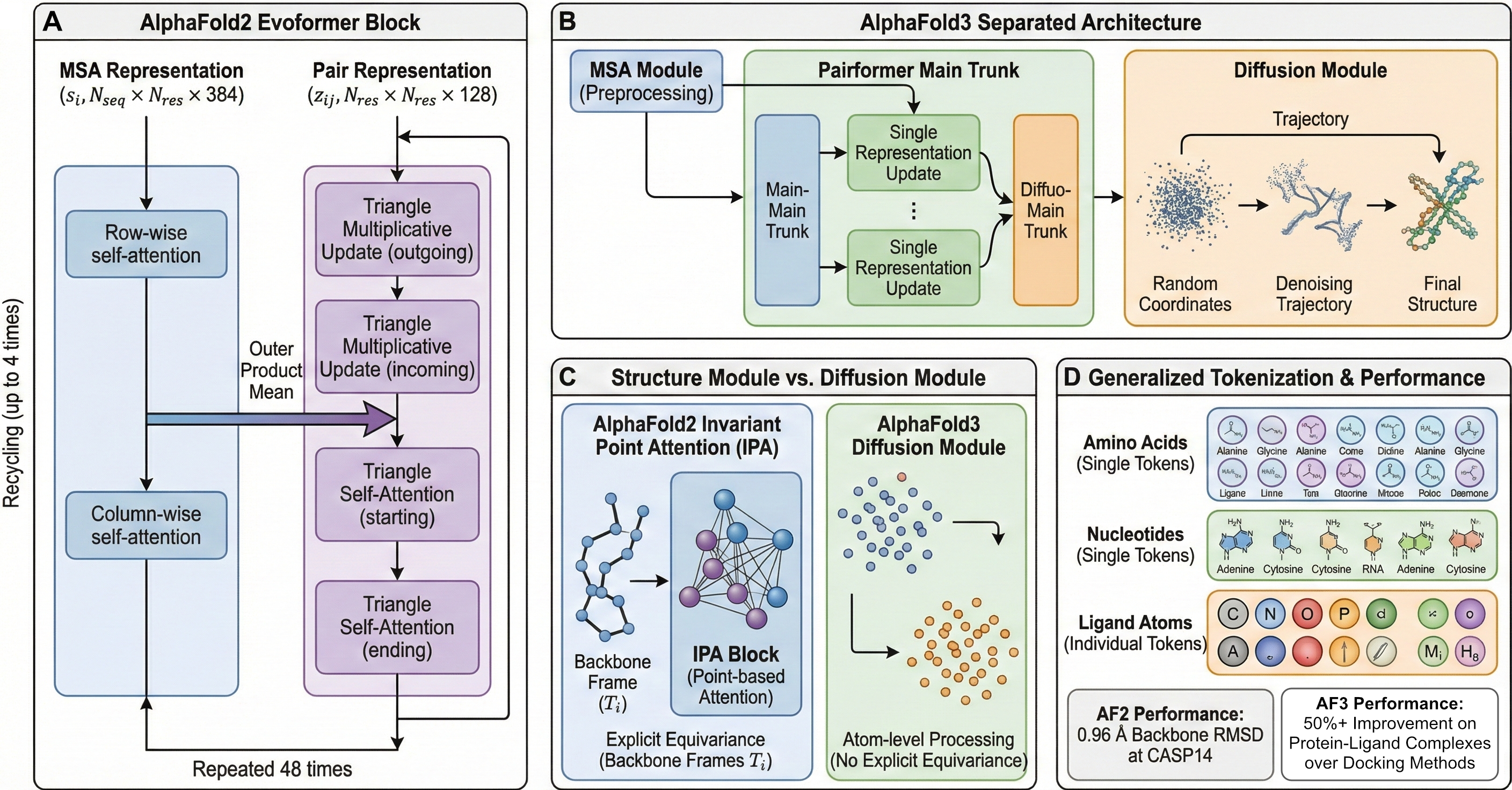
Figure 2: AlphaFold2 Evoformer and AlphaFold3 architectural comparison. (A) shows the AlphaFold2 Evoformer block with its two-track architecture: MSA representation ($N_{seq} \times N_{res} \times c$) undergoing row-wise and column-wise attention, pair representation ($N_{res} \times N_{res} \times c$) with triangle multiplicative updates and triangle self-attention, and the outer product mean connecting MSA to pair tracks. Include the 48-block repetition and recycling arrows. (B) depicts AlphaFold3's separated architecture: the MSA Module (smaller, preprocessing step), the Pairformer main trunk with simplified single representation updates, and the diffusion module showing the noising/denoising trajectory from random coordinates to final structure. (C) compares the Structure Module's Invariant Point Attention (with backbone frames $T_i$ and point-based attention) against AlphaFold3's diffusion module (atom-level processing, no explicit equivariance). (D) should illustrate AlphaFold3's generalized tokenization—amino acids as single tokens, nucleotides as single tokens, and ligand atoms as individual tokens—enabling the unified biomolecular prediction capability. Include performance metrics: AF2 achieving 0.96 Å backbone RMSD at CASP14, AF3 showing 50%+ improvement on protein-ligand complexes over docking methods.
ESMFold demonstrates language models encode structure
ESMFold (Lin et al., Science 2023) takes a radically different approach: rather than explicitly using MSAs, it relies on a 15-billion-parameter protein language model (ESM-2) that implicitly captures evolutionary information through masked language modeling on 250 million protein sequences.
Protein language models learn evolutionary constraints
ESM-2 uses BERT-style training: 15% of amino acids are randomly masked, and the model learns to predict them from context. Through this objective, attention heads learn pairwise residue correlations analogous to direct coupling analysis from covariance methods. The model captures co-evolutionary statistics without explicit MSA construction:
\[\mathcal{L}_{MLM} = -\sum_{i \in M} \log P(x_i | x_{-M}; \theta)\]The largest ESM-2 variant (15B parameters) has 48 layers with 40 attention heads and 5,120-dimensional embeddings. Remarkably, structure prediction accuracy scales with model size—the 15B model achieves 54.5% contact prediction accuracy versus 15.9% for the 8M variant.
Folding trunk adapts Evoformer for single sequences
ESMFold’s folding trunk is a simplified single-sequence version of the Evoformer, processing sequence and pairwise representations through 48 blocks without MSA attention. The sequence representation ($1 \times N_{res} \times 1024$) replaces the full MSA, dramatically reducing computation. The structure module inherits AlphaFold2’s IPA design for 3D coordinate prediction.
ESMFold achieves TM-score 0.83 on CAMEO (versus 0.88 for AlphaFold2 with MSAs) while running 6-60× faster—14 seconds for a 384-residue protein versus 85 seconds for AlphaFold2 (excluding MSA search time). This speed enables massive-scale applications: the ESM Metagenomic Atlas predicts structures for 617 million proteins from metagenomic samples.
ProteinMPNN pioneers deep learning inverse folding
ProteinMPNN (Dauparas et al., Science 2022) addresses the inverse problem: given a protein backbone structure, design an amino acid sequence that folds into that structure. Its message passing neural network with encoder-decoder architecture achieves 52.4% native sequence recovery—dramatically outperforming Rosetta’s 32.9%.
Distance-based encoding captures backbone geometry
ProteinMPNN encodes backbone structures using 25 pairwise distances between backbone atoms ($\mathrm{N, Cα, C, O}$, and virtual $\mathrm{Cβ}$) for residue pairs within a k-nearest neighbor graph ($k = 32$-$48$ based on Cα distances). These distances are transformed via radial basis functions into fixed-size features. The model has just 1.66 million parameters and runs in ~1 second per 100 residues on a CPU.
Critically, the architecture performs both node and edge updates during message passing:
\(h_i^{(l+1)} = \text{MLP}(h_i^{(l)}, \text{aggregate}(h_j^{(l)}, e_{ij}^{(l)}))\) \(e_{ij}^{(l+1)} = \text{MLP}(e_{ij}^{(l)}, h_i^{(l)}, h_j^{(l)})\)
This joint updating improved sequence recovery from 43.1% to 50.5% compared to node-only updates.
Autoregressive decoding with order-agnostic training
Unlike traditional $\text{N→C}$ terminal decoding, ProteinMPNN uses random permutation sampling during training, enabling design with arbitrary fixed residue constraints. The decoder autoregressively predicts amino acid logits given encoded structure features and previously sampled residues:
\[p(s|x) = \prod_i p(s_i | x, s_{\lt i})\]This flexibility enables symmetric and multi-state design: for homo-oligomers, logits from tied positions across chains are averaged; for conformational ensembles, logits from different states can be combined with positive or negative weights.
RFdiffusion generates novel protein structures
RFdiffusion (Watson et al., Nature 2023) applies denoising diffusion to protein backbone generation by fine-tuning the RoseTTAFold structure prediction network as a denoising model. This approach achieved 23/25 solved motif scaffolding problems (versus 15/25 for hallucination methods) and increased binder design success rates by ~100-fold.
Frame-based $\text{SE(3)}$-equivariant diffusion
Each residue is represented as a rigid frame in $\text{SE(3)}$—a 3D position (Cα coordinate) plus orientation ($\mathrm{N-Cα-C}$ rotation matrix). The forward diffusion process adds noise over 200 timesteps: Gaussian noise for translations and Brownian motion on $\text{SO(3)}$ for rotations.
RFdiffusion’s denoising network inherits RoseTTAFold’s three-track architecture: 1D sequence processing, 2D pairwise distance maps with biaxial attention, and 3D coordinates via $\text{SE(3)}$-equivariant transformers. The $\text{SE(3)}$-equivariance ensures outputs transform consistently with input rotations and translations—essential for physically meaningful structure generation.
Self-conditioning improves generation quality
RFdiffusion introduces self-conditioning, where the model receives its previous timestep prediction as template input:
At timestep $t$: input = $(X_t, \hat{X}_0^{(t+1)})$ → output = $\hat{X}_0^{(t)}$
This mechanism, analogous to AlphaFold2’s recycling, significantly improves both conditional (motif scaffolding) and unconditional design. The model also uses MSE loss instead of FAPE, promoting continuity of the global coordinate frame between timesteps—critical for diffusion’s iterative refinement.
Diverse conditioning enables targeted design
RFdiffusion supports multiple conditioning modes:
- Motif scaffolding: Fixed functional motifs (coordinates + sequence) with scaffold generation around them
- Binder design: Generation in context of target proteins with interface hotspot guidance
- Fold conditioning: Secondary structure or topology constraints for generating specific architectures
- Symmetric design: Equivariance enables cyclic, dihedral, and icosahedral assemblies
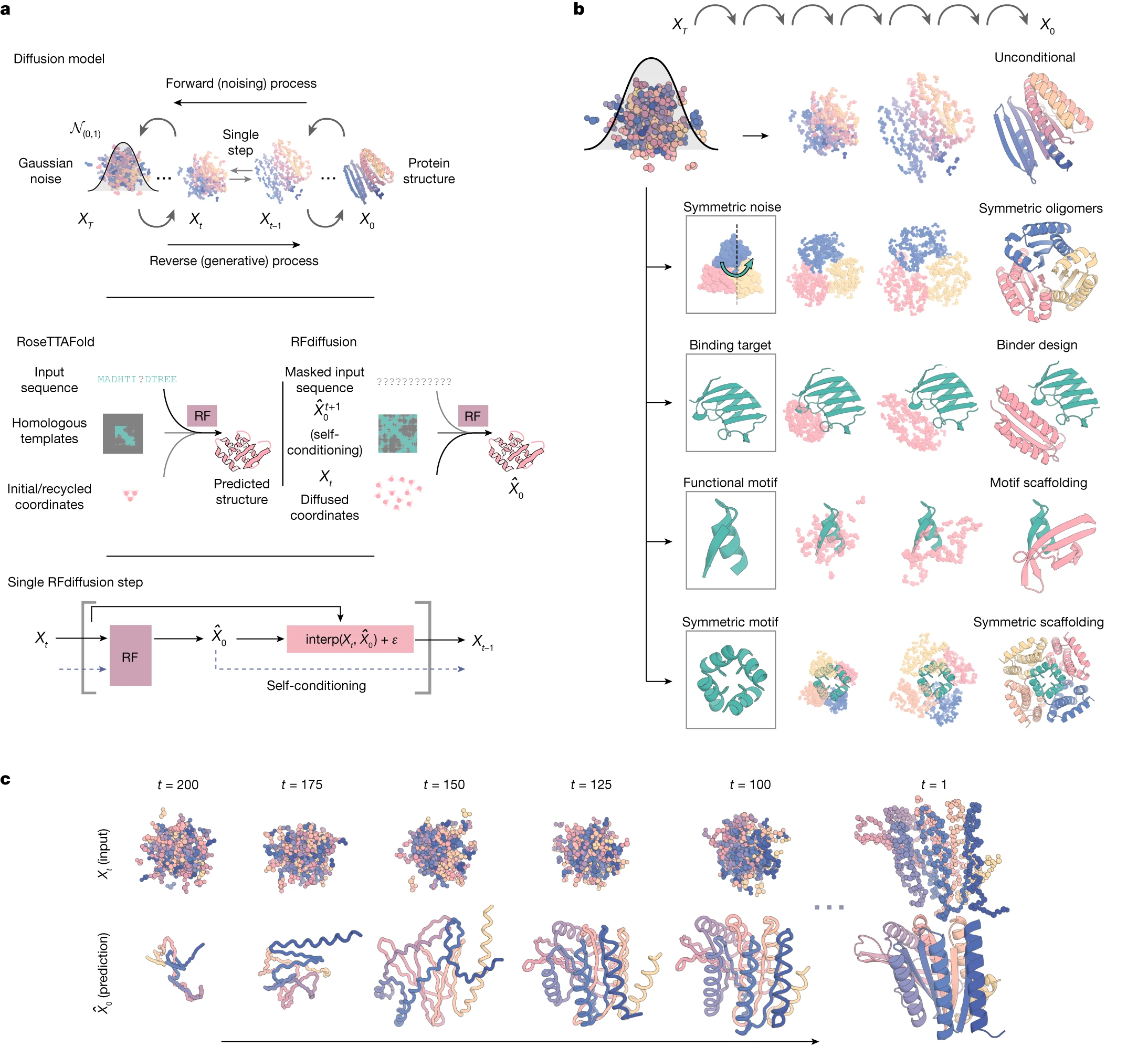
Figure 3: RFdiffusion diffusion process and conditioning mechanisms. (A) depicts the forward and reverse diffusion processes: starting from a PDB structure, showing progressive noising over 200 timesteps to random frames (translation + rotation), then reverse denoising from noise to designed structure. Include mathematical notation for Gaussian noise on translations and Brownian motion on SO(3) for rotations. (B) shows the RoseTTAFold three-track architecture adapted for denoising: 1D track processing sequence patterns, 2D track with biaxial attention for pairwise features, 3D track with SE(3)-transformer for coordinate updates, and arrows showing information flow between tracks. (C) illustrates self-conditioning: the previous timestep prediction $\hat{X}_0^{(t+1)}$ feeding back as template input alongside current noised structure $X_t$.
Boltz2 achieves open-source breakthrough in affinity prediction
The Boltz family (MIT Jameel Clinic / Recursion) represents the most accessible alternative to AlphaFold3. Boltz-1 achieved AlphaFold3-level accuracy under an MIT license enabling commercial use. Boltz-2 adds groundbreaking binding affinity prediction approaching Free Energy Perturbation accuracy—1000× faster than physics-based simulations.
Architectural innovations for efficiency and accuracy
Boltz-1 follows AlphaFold3’s general framework but introduces key innovations: a dense MSA pairing algorithm balancing signal quality with efficiency, Kabsch diffusion interpolation addressing limitations in AlphaFold3’s approach, and a redesigned confidence model leveraging pretrained trunk weights. Prediction time is 30-60 seconds on modern GPUs.
Boltz-2 adds an Affinity Module—a PairFormer that refines protein-ligand interactions and predicts both binding likelihood ($[0,1]$ probability) and continuous affinity (log μM scale). Training expanded to include molecular dynamics simulations, enabling physics-aware predictions. Mixed-precision computation (bfloat16) and trifast triangle attention reduce memory and runtime by ~50%.
The Boltz-2x steering mechanism applies physics-based potentials during inference to reduce steric clashes, while controllability features enable conditioning on experimental method (X-ray, NMR, MD-style), templates, and distance constraints.
Specialized tools address specific applications
Several transformer-based tools target specialized domains where general models underperform.
IgFold specializes in antibody structure prediction, using a language model pre-trained on 558 million antibody sequences. It achieves superior CDR H3 loop prediction (2.12 Å RMSD versus 2.98 Å for AlphaFold2) in under 25 seconds—critical for therapeutic antibody development.
OmegaFold pioneered MSA-free high-resolution prediction using a protein language model with geometry-inspired transformer. It excels on orphan proteins lacking evolutionary information (TM-score 0.73 versus 0.68 for AlphaFold2).
ColabFold integrates MMseqs2 for 40-60× faster homology search, enabling ~1,000 structure predictions per day on a single GPU. Its free Google Colab interface has made structure prediction accessible to laboratories without computational infrastructure.
OpenFold provides a trainable open-source AlphaFold2 implementation under Apache 2.0 license, along with OpenProteinSet—the largest public MSA database. It demonstrated that high accuracy can be achieved with as few as 1,000 training proteins, enabling efficient fine-tuning for specialized applications.
CASP benchmarks chart the field’s progress
The Critical Assessment of protein Structure Prediction (CASP) competition provides objective benchmarks for the field.
CASP14 (2020) marked AlphaFold2’s breakthrough: ~90 GDT-TS on moderately difficult targets, with predictions approaching experimental accuracy for the first time. The assessment concluded that the protein structure prediction problem was largely solved for single domains.
CASP15 (2022) saw continued AlphaFold2 dominance across all categories. Tertiary structure prediction no longer distinguished easy from hard targets—all became tractable. Protein complex prediction improved dramatically: 40% high-quality models versus 8% at CASP14, driven by AlphaFold-Multimer.
CASP16 (2024) confirmed that monomer prediction is a “largely solved problem”—no protein domain folds were missed by top methods. Focus shifted to remaining challenges: antibody-antigen complexes (lacking co-evolutionary signal), RNA tertiary structure, conformational dynamics, and ligand binding. On protein-ligand targets, Boltz-1 achieved 65% LDDT-PLI versus 40% for Chai-1.
| Competition | Top Method | GDT-TS / TM-score | Key Advancement |
|---|---|---|---|
| CASP13 (2018) | AlphaFold v1 | ~60 | CNN-based contact prediction |
| CASP14 (2020) | AlphaFold2 | ~92 | Transformer-based Evoformer |
| CASP15 (2022) | AF2 derivatives | ~92 | Complex prediction improvements |
| CASP16 (2024) | AF3/Boltz | TM >0.85 | Ligand and affinity prediction |
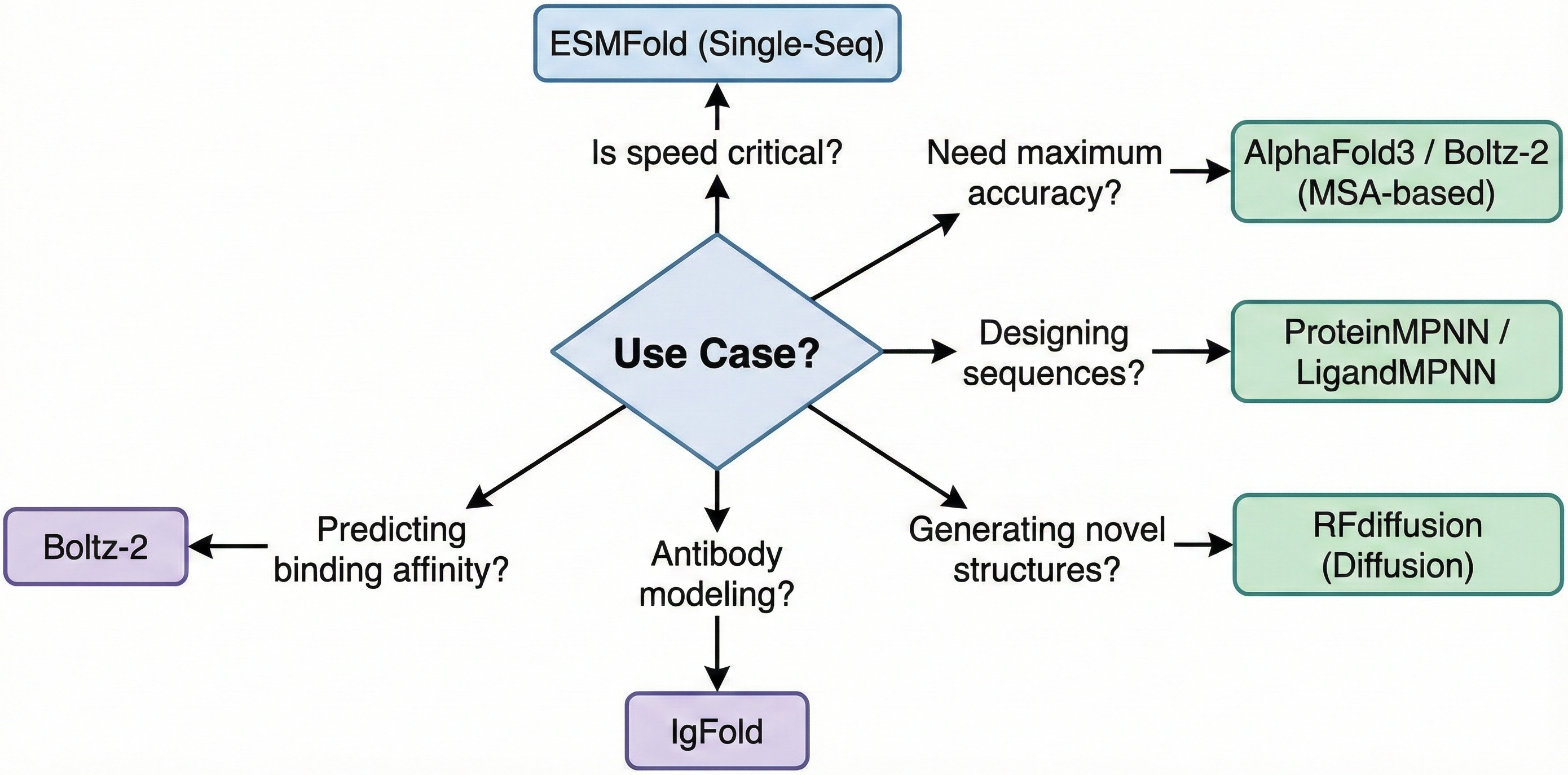
Synthesis: architectural principles across protein transformers
Examining these tools reveals common architectural principles that explain their success:
Attention captures long-range dependencies essential for folding. Unlike CNNs with limited receptive fields or RNNs with decaying memory, attention provides direct paths between distant residues. This matches protein physics: a residue’s environment depends on contacts that may be hundreds of positions away in sequence.
Multi-track architectures separate different information types. AlphaFold’s MSA/pair tracks, RoseTTAFold’s three-track design, and ESMFold’s sequence/pair representations all process distinct information sources before integration. This modularity enables specialized attention patterns for each representation type.
Geometric constraints appear through specialized attention mechanisms. Triangle attention/updates enforce distance geometry in pair representations. Invariant Point Attention and SE(3)-transformers maintain physical consistency in 3D. These inductive biases encode domain knowledge about protein structure.
Scale matters for language model approaches. ESM-2’s scaling from 8M to 15B parameters improved contact prediction from 15.9% to 54.5%. Large-scale pre-training on evolutionary data—250 million sequences for ESM-2, training on AlphaFold2 predictions for ESM-IF—provides rich initialization for structure-aware tasks.
Diffusion enables generative design. AlphaFold3, RFdiffusion, and Boltz all use diffusion processes for structure generation. This approach naturally handles multi-modal distributions (proteins can have multiple stable conformations) and enables conditioning through guidance mechanisms.
For researchers entering this field, mastering the mathematical foundations—scaled dot-product attention, multi-head attention, positional encodings—provides the vocabulary to understand any tool’s architecture. The field continues advancing rapidly: Boltz-2’s affinity prediction approaches physics-based simulation accuracy, ESM-3 generates novel functional proteins, and open-source implementations democratize access to state-of-the-art capabilities.
The transformer’s journey from language modeling to protein structure represents one of the most successful examples of architectural transfer in machine learning. As these tools mature, the focus shifts from predicting known structures to designing novel proteins with specified functions—a capability that will transform medicine, materials science, and our understanding of biology itself.
Key papers and DOIs
| Tool/Paper | DOI | Year |
|---|---|---|
| Attention Is All You Need | 10.48550/arXiv.1706.03762 | 2017 |
| AlphaFold2 | 10.1038/s41586-021-03819-2 | 2021 |
| AlphaFold3 | 10.1038/s41586-024-07487-w | 2024 |
| ESMFold/ESM-2 | 10.1126/science.ade2574 | 2023 |
| ESM-3 | 10.1126/science.ads0018 | 2025 |
| ProteinMPNN | 10.1126/science.add2187 | 2022 |
| LigandMPNN | 10.1038/s41592-025-02626-1 | 2025 |
| RFdiffusion | 10.1038/s41586-023-06415-8 | 2023 |
| RoseTTAFold | 10.1126/science.abj8754 | 2021 |
| Boltz-1 | 10.1101/2024.11.19.624167 | 2024 |
| Boltz-2 | 10.1101/2025.06.14.659707 | 2025 |
| OpenFold | 10.1038/s41592-024-02272-z | 2024 |
| ColabFold | 10.1038/s41592-022-01488-1 | 2022 |
| IgFold | 10.1038/s41467-023-38063-x | 2023 |
| OmegaFold | 10.1101/2022.07.21.500999 | 2022 |
| Chroma | 10.1038/s41586-023-06728-8 | 2023 |
| FrameDiff | 10.48550/arXiv.2302.02277 | 2023 |
Conclusion
The transformer architecture has become the universal backbone for computational structural biology, powering tools that predict protein structures with experimental accuracy, design novel sequences for target backbones, and generate entirely new protein architectures. The mathematical foundations—attention mechanisms, multi-head projections, positional encodings—remain consistent across applications, while domain-specific innovations like triangle updates, invariant point attention, and SE(3)-equivariant diffusion encode the geometric constraints unique to molecular biology.
Three key insights emerge for researchers entering this field: First, understanding the base transformer architecture transfers directly to comprehending any protein tool—the innovations are in how attention is applied and what representations are processed. Second, the choice between MSA-based (AlphaFold, Boltz) and language-model-based (ESMFold, OmegaFold) approaches reflects a fundamental tradeoff between accuracy and speed that depends on application requirements. Third, the field has shifted from prediction to design: RFdiffusion, ProteinMPNN, and generative language models like ESM-3 now enable creation of proteins that have never existed in nature.
The remaining frontiers—conformational dynamics, binding affinity prediction, antibody-antigen complexes, and RNA tertiary structure—represent the next wave of challenges. With Boltz-2 demonstrating FEP-level affinity prediction and diffusion models enabling atomic-level design accuracy, the transformer-powered revolution in structural biology continues to accelerate.
Enjoy Reading This Article?
Here are some more articles you might like to read next: